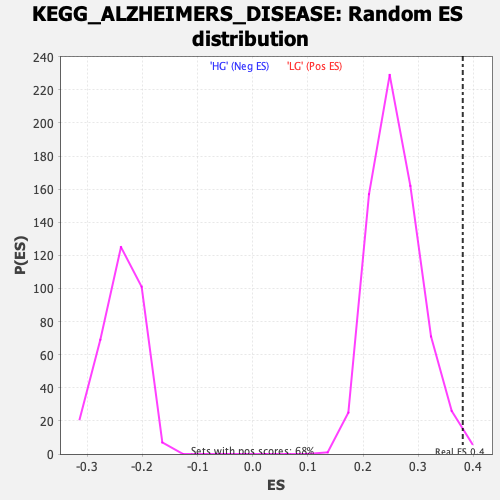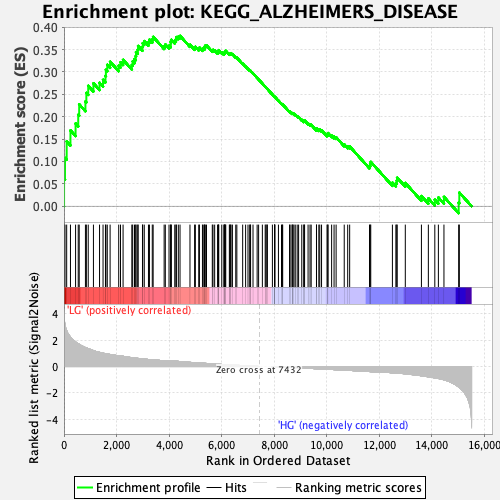
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_ALZHEIMERS_DISEASE |
| Enrichment Score (ES) | 0.38074696 |
| Normalized Enrichment Score (NES) | 1.4726322 |
| Nominal p-value | 0.0088626295 |
| FDR q-value | 0.22336806 |
| FWER p-Value | 0.953 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COX6A2 | Cox6a2 | 3 | 4.063 | 0.0613 | Yes | ||
| 2 | RYR3 | Ryr3 | 38 | 3.221 | 0.1079 | Yes | ||
| 3 | GRIN1 | Grin1 | 102 | 2.719 | 0.1450 | Yes | ||
| 4 | MME | Mme | 246 | 2.231 | 0.1694 | Yes | ||
| 5 | PSEN2 | Psen2 | 441 | 1.857 | 0.1849 | Yes | ||
| 6 | BID | Bid | 542 | 1.724 | 0.2045 | Yes | ||
| 7 | FAS | Fas | 579 | 1.684 | 0.2277 | Yes | ||
| 8 | SNCA | Snca | 816 | 1.442 | 0.2342 | Yes | ||
| 9 | APOE | Apoe | 852 | 1.410 | 0.2533 | Yes | ||
| 10 | ATP2A3 | Atp2a3 | 923 | 1.350 | 0.2692 | Yes | ||
| 11 | NDUFA4L2 | Ndufa4l2 | 1121 | 1.210 | 0.2747 | Yes | ||
| 12 | ATP2A1 | Atp2a1 | 1354 | 1.073 | 0.2758 | Yes | ||
| 13 | CASP7 | Casp7 | 1487 | 1.011 | 0.2826 | Yes | ||
| 14 | ITPR2 | Itpr2 | 1579 | 0.974 | 0.2914 | Yes | ||
| 15 | PLCB4 | Plcb4 | 1596 | 0.969 | 0.3050 | Yes | ||
| 16 | UQCRQ | Uqcrq | 1648 | 0.955 | 0.3162 | Yes | ||
| 17 | LPL | Lpl | 1756 | 0.911 | 0.3230 | Yes | ||
| 18 | CASP9 | Casp9 | 2084 | 0.814 | 0.3141 | Yes | ||
| 19 | COX5B | Cox5b | 2152 | 0.799 | 0.3218 | Yes | ||
| 20 | MT-CYB | mt-Cytb | 2250 | 0.773 | 0.3272 | Yes | ||
| 21 | TNFRSF1A | Tnfrsf1a | 2584 | 0.672 | 0.3157 | Yes | ||
| 22 | APP | App | 2616 | 0.663 | 0.3237 | Yes | ||
| 23 | COX8C | Cox8b | 2685 | 0.646 | 0.3291 | Yes | ||
| 24 | GRIN2D | Grin2d | 2733 | 0.635 | 0.3357 | Yes | ||
| 25 | CDK5R1 | Cdk5r1 | 2744 | 0.632 | 0.3446 | Yes | ||
| 26 | IDE | Ide | 2809 | 0.617 | 0.3498 | Yes | ||
| 27 | MAPT | Mapt | 2823 | 0.613 | 0.3582 | Yes | ||
| 28 | CDK5 | Cdk5 | 2989 | 0.575 | 0.3562 | Yes | ||
| 29 | NDUFB9 | Ndufb9 | 2998 | 0.573 | 0.3643 | Yes | ||
| 30 | COX7A1 | Cox7a1 | 3057 | 0.561 | 0.3691 | Yes | ||
| 31 | BAD | Bad | 3217 | 0.532 | 0.3668 | Yes | ||
| 32 | GAPDH | Gapdh | 3254 | 0.525 | 0.3724 | Yes | ||
| 33 | NDUFS7 | Ndufs7 | 3362 | 0.504 | 0.3731 | Yes | ||
| 34 | NDUFA1 | Ndufa1 | 3395 | 0.498 | 0.3785 | Yes | ||
| 35 | NDUFB6 | Ndufb6 | 3814 | 0.432 | 0.3579 | Yes | ||
| 36 | IL1B | Il1b | 3859 | 0.432 | 0.3615 | Yes | ||
| 37 | COX7B2 | Cox7b2 | 4000 | 0.421 | 0.3588 | Yes | ||
| 38 | NDUFS8 | Ndufs8 | 4063 | 0.417 | 0.3611 | Yes | ||
| 39 | MT-CO1 | mt-Co1 | 4064 | 0.417 | 0.3674 | Yes | ||
| 40 | PSEN1 | Psen1 | 4089 | 0.412 | 0.3721 | Yes | ||
| 41 | GRIN2C | Grin2c | 4219 | 0.407 | 0.3699 | Yes | ||
| 42 | FADD | Fadd | 4262 | 0.401 | 0.3732 | Yes | ||
| 43 | COX5A | Cox5a | 4281 | 0.397 | 0.3780 | Yes | ||
| 44 | CALM1 | Calm1 | 4354 | 0.386 | 0.3792 | Yes | ||
| 45 | ITPR3 | Itpr3 | 4419 | 0.376 | 0.3807 | Yes | ||
| 46 | APBB1 | Apbb1 | 4798 | 0.312 | 0.3609 | No | ||
| 47 | COX7A2L | Cox7a2l | 4974 | 0.289 | 0.3539 | No | ||
| 48 | CAPN1 | Capn1 | 5003 | 0.286 | 0.3564 | No | ||
| 49 | COX8A | Cox8a | 5132 | 0.271 | 0.3522 | No | ||
| 50 | NCSTN | Ncstn | 5158 | 0.268 | 0.3546 | No | ||
| 51 | NDUFA10 | Ndufa10 | 5273 | 0.251 | 0.3510 | No | ||
| 52 | COX4I1 | Cox4i1 | 5301 | 0.248 | 0.3530 | No | ||
| 53 | NDUFA5 | Ndufa5 | 5352 | 0.242 | 0.3534 | No | ||
| 54 | NDUFA2 | Ndufa2 | 5361 | 0.240 | 0.3565 | No | ||
| 55 | CAPN2 | Capn2 | 5378 | 0.237 | 0.3591 | No | ||
| 56 | CHP1 | Chp1 | 5427 | 0.231 | 0.3594 | No | ||
| 57 | NDUFV1 | Ndufv1 | 5648 | 0.201 | 0.3482 | No | ||
| 58 | MT-CO2 | mt-Co2 | 5674 | 0.198 | 0.3495 | No | ||
| 59 | APAF1 | Apaf1 | 5738 | 0.190 | 0.3483 | No | ||
| 60 | NDUFS6 | Ndufs6 | 5849 | 0.174 | 0.3438 | No | ||
| 61 | GSK3B | Gsk3b | 5879 | 0.171 | 0.3445 | No | ||
| 62 | NAE1 | Nae1 | 5882 | 0.170 | 0.3470 | No | ||
| 63 | NDUFB7 | Ndufb7 | 5894 | 0.168 | 0.3488 | No | ||
| 64 | NDUFB4 | Ndufb4 | 6009 | 0.155 | 0.3437 | No | ||
| 65 | NDUFS3 | Ndufs3 | 6090 | 0.146 | 0.3408 | No | ||
| 66 | NDUFA9 | Ndufa9 | 6103 | 0.145 | 0.3422 | No | ||
| 67 | CYC1 | Cyc1 | 6122 | 0.143 | 0.3432 | No | ||
| 68 | APH1A | Aph1a | 6126 | 0.143 | 0.3451 | No | ||
| 69 | UQCR10 | Uqcr10 | 6139 | 0.141 | 0.3465 | No | ||
| 70 | NDUFB5 | Ndufb5 | 6160 | 0.139 | 0.3473 | No | ||
| 71 | BACE1 | Bace1 | 6298 | 0.124 | 0.3403 | No | ||
| 72 | NDUFS5 | Ndufs5 | 6319 | 0.122 | 0.3408 | No | ||
| 73 | PLCB2 | Plcb2 | 6348 | 0.118 | 0.3408 | No | ||
| 74 | CASP8 | Casp8 | 6359 | 0.117 | 0.3419 | No | ||
| 75 | NDUFA6 | Ndufa6 | 6414 | 0.110 | 0.3401 | No | ||
| 76 | PPP3CC | Ppp3cc | 6542 | 0.096 | 0.3333 | No | ||
| 77 | CACNA1S | Cacna1s | 6583 | 0.091 | 0.3320 | No | ||
| 78 | COX7C | Cox7c | 6805 | 0.065 | 0.3186 | No | ||
| 79 | SDHB | Sdhb | 6917 | 0.055 | 0.3123 | No | ||
| 80 | NDUFB1 | Ndufb1-ps | 7008 | 0.045 | 0.3071 | No | ||
| 81 | COX6A1 | Cox6a1 | 7077 | 0.036 | 0.3032 | No | ||
| 82 | SDHC | Sdhc | 7086 | 0.036 | 0.3032 | No | ||
| 83 | CALML3 | Calml3 | 7106 | 0.034 | 0.3025 | No | ||
| 84 | MAPK3 | Mapk3 | 7199 | 0.024 | 0.2969 | No | ||
| 85 | NDUFB3 | Ndufb3 | 7356 | 0.007 | 0.2869 | No | ||
| 86 | NDUFA8 | Ndufa8 | 7412 | 0.002 | 0.2833 | No | ||
| 87 | MT-ATP6 | mt-Atp6 | 7555 | 0.000 | 0.2741 | No | ||
| 88 | PPP3R2 | Ppp3r2 | 7663 | 0.000 | 0.2671 | No | ||
| 89 | TNF | Tnf | 7693 | 0.000 | 0.2653 | No | ||
| 90 | MT-CO3 | mt-Co3 | 7729 | 0.000 | 0.2630 | No | ||
| 91 | CACNA1F | Cacna1f | 7745 | 0.000 | 0.2620 | No | ||
| 92 | NDUFC1 | Ndufc1 | 7937 | -0.003 | 0.2496 | No | ||
| 93 | NDUFS4 | Ndufs4 | 8020 | -0.010 | 0.2445 | No | ||
| 94 | ATP5F1E | Atp5e | 8023 | -0.010 | 0.2445 | No | ||
| 95 | UQCRB | Uqcrb | 8025 | -0.011 | 0.2446 | No | ||
| 96 | NDUFA4 | Ndufa4 | 8027 | -0.011 | 0.2447 | No | ||
| 97 | ATP5PD | Atp5h | 8167 | -0.025 | 0.2360 | No | ||
| 98 | HSD17B10 | Hsd17b10 | 8284 | -0.034 | 0.2290 | No | ||
| 99 | MAPK1 | Mapk1 | 8316 | -0.037 | 0.2275 | No | ||
| 100 | NDUFB10 | Ndufb10 | 8329 | -0.039 | 0.2273 | No | ||
| 101 | ERN1 | Ern1 | 8591 | -0.065 | 0.2114 | No | ||
| 102 | ADAM10 | Adam10 | 8611 | -0.066 | 0.2111 | No | ||
| 103 | NDUFS2 | Ndufs2 | 8687 | -0.073 | 0.2074 | No | ||
| 104 | ATP2A2 | Atp2a2 | 8693 | -0.074 | 0.2082 | No | ||
| 105 | NDUFS1 | Ndufs1 | 8719 | -0.076 | 0.2077 | No | ||
| 106 | NOS1 | Nos1 | 8757 | -0.080 | 0.2065 | No | ||
| 107 | COX4I2 | Cox4i2 | 8807 | -0.084 | 0.2046 | No | ||
| 108 | EIF2AK3 | Eif2ak3 | 8881 | -0.090 | 0.2012 | No | ||
| 109 | COX6C | Cox6c | 8935 | -0.096 | 0.1992 | No | ||
| 110 | LRP1 | Lrp1 | 9056 | -0.110 | 0.1931 | No | ||
| 111 | ATP5F1D | Atp5d | 9134 | -0.118 | 0.1898 | No | ||
| 112 | COX6B1 | Cox6b1 | 9141 | -0.119 | 0.1913 | No | ||
| 113 | ATF6 | Atf6 | 9163 | -0.122 | 0.1917 | No | ||
| 114 | ATP5PF | Atp5j | 9296 | -0.134 | 0.1852 | No | ||
| 115 | ATP5F1B | Atp5b | 9370 | -0.141 | 0.1826 | No | ||
| 116 | CYCS | Cycs | 9414 | -0.145 | 0.1820 | No | ||
| 117 | NDUFA3 | Ndufa3 | 9616 | -0.166 | 0.1714 | No | ||
| 118 | PLCB3 | Plcb3 | 9618 | -0.166 | 0.1739 | No | ||
| 119 | ATP5F1C | Atp5c1 | 9706 | -0.174 | 0.1708 | No | ||
| 120 | CALM3 | Calm3 | 9742 | -0.178 | 0.1713 | No | ||
| 121 | NDUFV3 | Ndufv3 | 9809 | -0.184 | 0.1698 | No | ||
| 122 | PPP3CA | Ppp3ca | 10018 | -0.206 | 0.1593 | No | ||
| 123 | CALM2 | Calm2 | 10048 | -0.209 | 0.1606 | No | ||
| 124 | ATP5MC3 | Atp5g3 | 10054 | -0.209 | 0.1635 | No | ||
| 125 | NDUFV2 | Ndufv2 | 10193 | -0.222 | 0.1579 | No | ||
| 126 | BACE2 | Bace2 | 10286 | -0.232 | 0.1554 | No | ||
| 127 | UQCRC2 | Uqcrc2 | 10364 | -0.240 | 0.1540 | No | ||
| 128 | COX7A2 | Cox7a2 | 10673 | -0.273 | 0.1381 | No | ||
| 129 | UQCRC1 | Uqcrc1 | 10805 | -0.287 | 0.1340 | No | ||
| 130 | NDUFB2 | Ndufb2 | 10880 | -0.294 | 0.1336 | No | ||
| 131 | ATP5F1A | Atp5a1 | 11636 | -0.380 | 0.0903 | No | ||
| 132 | PPP3CB | Ppp3cb | 11662 | -0.384 | 0.0945 | No | ||
| 133 | NDUFAB1 | Ndufab1 | 11683 | -0.388 | 0.0990 | No | ||
| 134 | UQCRFS1 | Uqcrfs1 | 12508 | -0.475 | 0.0527 | No | ||
| 135 | COX7B | Cox7b | 12640 | -0.494 | 0.0516 | No | ||
| 136 | CHP2 | Chp2 | 12674 | -0.499 | 0.0570 | No | ||
| 137 | GRIN2B | Grin2b | 12688 | -0.500 | 0.0638 | No | ||
| 138 | GRIN2A | Grin2a | 12999 | -0.553 | 0.0520 | No | ||
| 139 | ITPR1 | Itpr1 | 13613 | -0.701 | 0.0227 | No | ||
| 140 | CACNA1D | Cacna1d | 13877 | -0.781 | 0.0174 | No | ||
| 141 | COX6B2 | Cox6b2 | 14123 | -0.857 | 0.0145 | No | ||
| 142 | ADAM17 | Adam17 | 14259 | -0.905 | 0.0194 | No | ||
| 143 | CASP3 | Casp3 | 14471 | -1.004 | 0.0209 | No | ||
| 144 | PLCB1 | Plcb1 | 15032 | -1.546 | 0.0079 | No | ||
| 145 | CACNA1C | Cacna1c | 15057 | -1.590 | 0.0304 | No |