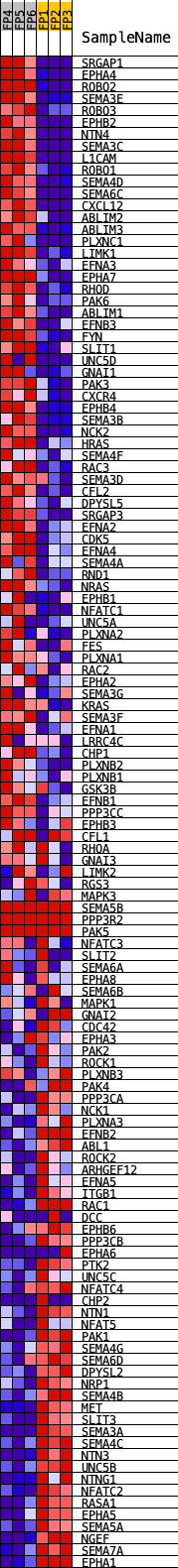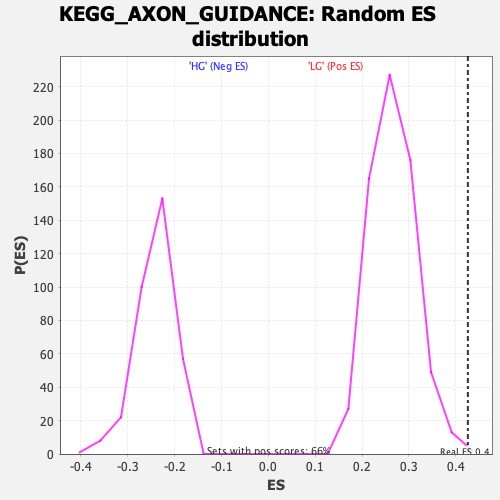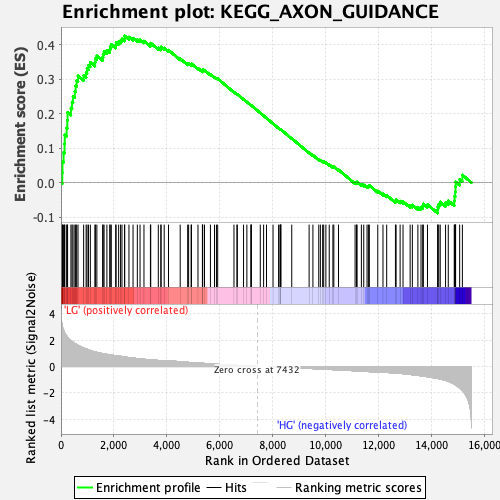
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_AXON_GUIDANCE |
| Enrichment Score (ES) | 0.42586383 |
| Normalized Enrichment Score (NES) | 1.6082749 |
| Nominal p-value | 0.0015174507 |
| FDR q-value | 0.122683264 |
| FWER p-Value | 0.6 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SRGAP1 | Srgap1 | 48 | 3.094 | 0.0296 | Yes | ||
| 2 | EPHA4 | Epha4 | 56 | 3.037 | 0.0613 | Yes | ||
| 3 | ROBO2 | Robo2 | 97 | 2.750 | 0.0878 | Yes | ||
| 4 | SEMA3E | Sema3e | 135 | 2.563 | 0.1125 | Yes | ||
| 5 | ROBO3 | Robo3 | 143 | 2.526 | 0.1387 | Yes | ||
| 6 | EPHB2 | Ephb2 | 214 | 2.318 | 0.1587 | Yes | ||
| 7 | NTN4 | Ntn4 | 239 | 2.256 | 0.1810 | Yes | ||
| 8 | SEMA3C | Sema3c | 249 | 2.214 | 0.2038 | Yes | ||
| 9 | L1CAM | L1cam | 375 | 1.967 | 0.2165 | Yes | ||
| 10 | ROBO1 | Robo1 | 418 | 1.897 | 0.2338 | Yes | ||
| 11 | SEMA4D | Sema4d | 456 | 1.837 | 0.2509 | Yes | ||
| 12 | SEMA6C | Sema6c | 524 | 1.736 | 0.2649 | Yes | ||
| 13 | CXCL12 | Cxcl12 | 553 | 1.714 | 0.2812 | Yes | ||
| 14 | ABLIM2 | Ablim2 | 592 | 1.660 | 0.2963 | Yes | ||
| 15 | ABLIM3 | Ablim3 | 643 | 1.602 | 0.3100 | Yes | ||
| 16 | PLXNC1 | Plxnc1 | 856 | 1.408 | 0.3111 | Yes | ||
| 17 | LIMK1 | Limk1 | 950 | 1.334 | 0.3192 | Yes | ||
| 18 | EFNA3 | Efna3 | 985 | 1.310 | 0.3308 | Yes | ||
| 19 | EPHA7 | Epha7 | 1041 | 1.269 | 0.3407 | Yes | ||
| 20 | RHOD | Rhod | 1110 | 1.217 | 0.3491 | Yes | ||
| 21 | PAK6 | Pak6 | 1283 | 1.107 | 0.3497 | Yes | ||
| 22 | ABLIM1 | Ablim1 | 1300 | 1.098 | 0.3602 | Yes | ||
| 23 | EFNB3 | Efnb3 | 1356 | 1.072 | 0.3680 | Yes | ||
| 24 | FYN | Fyn | 1575 | 0.976 | 0.3642 | Yes | ||
| 25 | SLIT1 | Slit1 | 1593 | 0.971 | 0.3733 | Yes | ||
| 26 | UNC5D | Unc5d | 1629 | 0.960 | 0.3812 | Yes | ||
| 27 | GNAI1 | Gnai1 | 1736 | 0.921 | 0.3841 | Yes | ||
| 28 | PAK3 | Pak3 | 1844 | 0.880 | 0.3864 | Yes | ||
| 29 | CXCR4 | Cxcr4 | 1862 | 0.875 | 0.3946 | Yes | ||
| 30 | EPHB4 | Ephb4 | 1902 | 0.862 | 0.4012 | Yes | ||
| 31 | SEMA3B | Sema3b | 2070 | 0.819 | 0.3990 | Yes | ||
| 32 | NCK2 | Nck2 | 2082 | 0.814 | 0.4069 | Yes | ||
| 33 | HRAS | Hras | 2180 | 0.791 | 0.4089 | Yes | ||
| 34 | SEMA4F | Sema4f | 2253 | 0.772 | 0.4124 | Yes | ||
| 35 | RAC3 | Rac3 | 2306 | 0.753 | 0.4170 | Yes | ||
| 36 | SEMA3D | Sema3d | 2405 | 0.726 | 0.4183 | Yes | ||
| 37 | CFL2 | Cfl2 | 2408 | 0.724 | 0.4259 | Yes | ||
| 38 | DPYSL5 | Dpysl5 | 2570 | 0.674 | 0.4225 | No | ||
| 39 | SRGAP3 | Srgap3 | 2725 | 0.636 | 0.4193 | No | ||
| 40 | EFNA2 | Efna2 | 2887 | 0.599 | 0.4152 | No | ||
| 41 | CDK5 | Cdk5 | 2989 | 0.575 | 0.4147 | No | ||
| 42 | EFNA4 | Efna4 | 3138 | 0.544 | 0.4108 | No | ||
| 43 | SEMA4A | Sema4a | 3382 | 0.500 | 0.4003 | No | ||
| 44 | RND1 | Rnd1 | 3394 | 0.498 | 0.4049 | No | ||
| 45 | NRAS | Nras | 3680 | 0.455 | 0.3912 | No | ||
| 46 | EPHB1 | Ephb1 | 3779 | 0.438 | 0.3895 | No | ||
| 47 | NFATC1 | Nfatc1 | 3783 | 0.437 | 0.3939 | No | ||
| 48 | UNC5A | Unc5a | 3903 | 0.427 | 0.3907 | No | ||
| 49 | PLXNA2 | Plxna2 | 4066 | 0.416 | 0.3846 | No | ||
| 50 | FES | Fes | 4508 | 0.360 | 0.3597 | No | ||
| 51 | PLXNA1 | Plxna1 | 4791 | 0.313 | 0.3447 | No | ||
| 52 | RAC2 | Rac2 | 4827 | 0.309 | 0.3457 | No | ||
| 53 | EPHA2 | Epha2 | 4921 | 0.295 | 0.3428 | No | ||
| 54 | SEMA3G | Sema3g | 4936 | 0.294 | 0.3450 | No | ||
| 55 | KRAS | Kras | 5181 | 0.266 | 0.3320 | No | ||
| 56 | SEMA3F | Sema3f | 5348 | 0.242 | 0.3238 | No | ||
| 57 | EFNA1 | Efna1 | 5353 | 0.241 | 0.3260 | No | ||
| 58 | LRRC4C | Lrrc4c | 5359 | 0.240 | 0.3283 | No | ||
| 59 | CHP1 | Chp1 | 5427 | 0.231 | 0.3263 | No | ||
| 60 | PLXNB2 | Plxnb2 | 5651 | 0.200 | 0.3140 | No | ||
| 61 | PLXNB1 | Plxnb1 | 5806 | 0.180 | 0.3059 | No | ||
| 62 | GSK3B | Gsk3b | 5879 | 0.171 | 0.3030 | No | ||
| 63 | EFNB1 | Efnb1 | 5926 | 0.166 | 0.3018 | No | ||
| 64 | PPP3CC | Ppp3cc | 6542 | 0.096 | 0.2629 | No | ||
| 65 | EPHB3 | Ephb3 | 6654 | 0.083 | 0.2565 | No | ||
| 66 | CFL1 | Cfl1 | 6669 | 0.082 | 0.2565 | No | ||
| 67 | RHOA | Rhoa | 6904 | 0.056 | 0.2419 | No | ||
| 68 | GNAI3 | Gnai3 | 7030 | 0.042 | 0.2342 | No | ||
| 69 | LIMK2 | Limk2 | 7184 | 0.025 | 0.2245 | No | ||
| 70 | RGS3 | Rgs3 | 7192 | 0.024 | 0.2243 | No | ||
| 71 | MAPK3 | Mapk3 | 7199 | 0.024 | 0.2242 | No | ||
| 72 | SEMA5B | Sema5b | 7537 | 0.000 | 0.2023 | No | ||
| 73 | PPP3R2 | Ppp3r2 | 7663 | 0.000 | 0.1942 | No | ||
| 74 | PAK5 | Pak7 | 7772 | 0.000 | 0.1872 | No | ||
| 75 | NFATC3 | Nfatc3 | 8019 | -0.010 | 0.1713 | No | ||
| 76 | SLIT2 | Slit2 | 8229 | -0.029 | 0.1581 | No | ||
| 77 | SEMA6A | Sema6a | 8280 | -0.034 | 0.1552 | No | ||
| 78 | EPHA8 | Epha8 | 8303 | -0.036 | 0.1541 | No | ||
| 79 | SEMA6B | Sema6b | 8309 | -0.036 | 0.1542 | No | ||
| 80 | MAPK1 | Mapk1 | 8316 | -0.037 | 0.1542 | No | ||
| 81 | GNAI2 | Gnai2 | 8727 | -0.077 | 0.1284 | No | ||
| 82 | CDC42 | Cdc42 | 9384 | -0.142 | 0.0873 | No | ||
| 83 | EPHA3 | Epha3 | 9528 | -0.157 | 0.0796 | No | ||
| 84 | PAK2 | Pak2 | 9749 | -0.178 | 0.0672 | No | ||
| 85 | ROCK1 | Rock1 | 9810 | -0.184 | 0.0653 | No | ||
| 86 | PLXNB3 | Plxnb3 | 9896 | -0.193 | 0.0618 | No | ||
| 87 | PAK4 | Pak4 | 9936 | -0.197 | 0.0614 | No | ||
| 88 | PPP3CA | Ppp3ca | 10018 | -0.206 | 0.0583 | No | ||
| 89 | NCK1 | Nck1 | 10145 | -0.218 | 0.0524 | No | ||
| 90 | PLXNA3 | Plxna3 | 10284 | -0.232 | 0.0459 | No | ||
| 91 | EFNB2 | Efnb2 | 10319 | -0.236 | 0.0462 | No | ||
| 92 | ABL1 | Abl1 | 10496 | -0.253 | 0.0374 | No | ||
| 93 | ROCK2 | Rock2 | 11123 | -0.321 | 0.0002 | No | ||
| 94 | ARHGEF12 | Arhgef12 | 11177 | -0.328 | 0.0002 | No | ||
| 95 | EFNA5 | Efna5 | 11199 | -0.330 | 0.0023 | No | ||
| 96 | ITGB1 | Itgb1 | 11361 | -0.348 | -0.0045 | No | ||
| 97 | RAC1 | Rac1 | 11450 | -0.360 | -0.0064 | No | ||
| 98 | DCC | Dcc | 11575 | -0.373 | -0.0105 | No | ||
| 99 | EPHB6 | Ephb6 | 11626 | -0.379 | -0.0097 | No | ||
| 100 | PPP3CB | Ppp3cb | 11662 | -0.384 | -0.0079 | No | ||
| 101 | EPHA6 | Epha6 | 11981 | -0.418 | -0.0241 | No | ||
| 102 | PTK2 | Ptk2 | 12178 | -0.432 | -0.0323 | No | ||
| 103 | UNC5C | Unc5c | 12318 | -0.445 | -0.0366 | No | ||
| 104 | NFATC4 | Nfatc4 | 12649 | -0.495 | -0.0528 | No | ||
| 105 | CHP2 | Chp2 | 12674 | -0.499 | -0.0491 | No | ||
| 106 | NTN1 | Ntn1 | 12826 | -0.521 | -0.0534 | No | ||
| 107 | NFAT5 | Nfat5 | 12936 | -0.538 | -0.0548 | No | ||
| 108 | PAK1 | Pak1 | 13205 | -0.599 | -0.0659 | No | ||
| 109 | SEMA4G | Sema4g | 13289 | -0.618 | -0.0647 | No | ||
| 110 | SEMA6D | Sema6d | 13498 | -0.667 | -0.0712 | No | ||
| 111 | DPYSL2 | Dpysl2 | 13616 | -0.702 | -0.0713 | No | ||
| 112 | NRP1 | Nrp1 | 13680 | -0.720 | -0.0678 | No | ||
| 113 | SEMA4B | Sema4b | 13704 | -0.729 | -0.0616 | No | ||
| 114 | MET | Met | 13861 | -0.774 | -0.0635 | No | ||
| 115 | SLIT3 | Slit3 | 14245 | -0.900 | -0.0789 | No | ||
| 116 | SEMA3A | Sema3a | 14254 | -0.903 | -0.0699 | No | ||
| 117 | SEMA4C | Sema4c | 14288 | -0.920 | -0.0623 | No | ||
| 118 | NTN3 | Ntn3 | 14343 | -0.942 | -0.0558 | No | ||
| 119 | UNC5B | Unc5b | 14545 | -1.051 | -0.0578 | No | ||
| 120 | NTNG1 | Ntng1 | 14647 | -1.121 | -0.0525 | No | ||
| 121 | NFATC2 | Nfatc2 | 14873 | -1.340 | -0.0529 | No | ||
| 122 | RASA1 | Rasa1 | 14886 | -1.350 | -0.0394 | No | ||
| 123 | EPHA5 | Epha5 | 14912 | -1.382 | -0.0264 | No | ||
| 124 | SEMA5A | Sema5a | 14920 | -1.391 | -0.0122 | No | ||
| 125 | NGEF | Ngef | 14925 | -1.396 | 0.0023 | No | ||
| 126 | SEMA7A | Sema7a | 15078 | -1.630 | 0.0097 | No | ||
| 127 | EPHA1 | Epha1 | 15181 | -1.820 | 0.0223 | No |