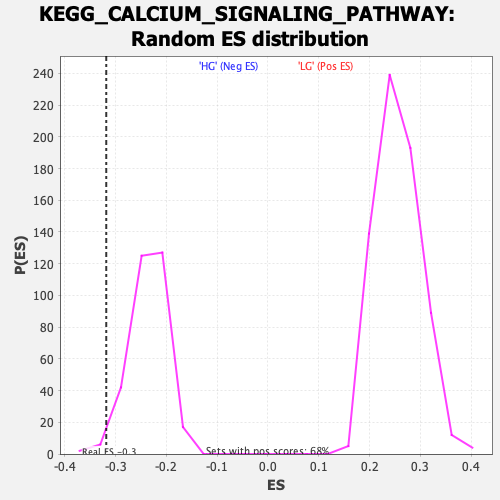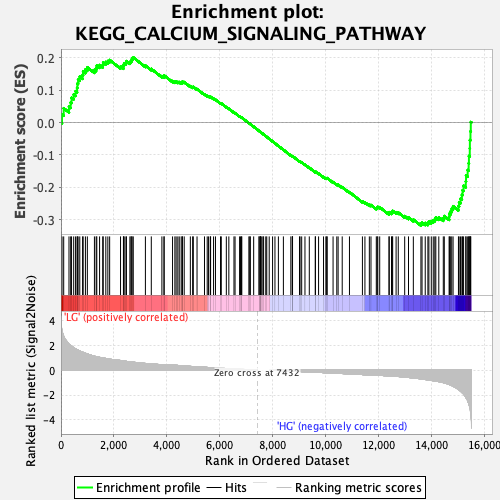
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_CALCIUM_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.31841573 |
| Normalized Enrichment Score (NES) | -1.3453138 |
| Nominal p-value | 0.021943573 |
| FDR q-value | 0.3336386 |
| FWER p-Value | 0.993 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RYR3 | Ryr3 | 38 | 3.221 | 0.0251 | No | ||
| 2 | GRIN1 | Grin1 | 102 | 2.719 | 0.0443 | No | ||
| 3 | SLC8A1 | Slc8a1 | 302 | 2.078 | 0.0492 | No | ||
| 4 | ADCY1 | Adcy1 | 366 | 1.982 | 0.0621 | No | ||
| 5 | ADRB2 | Adrb2 | 399 | 1.934 | 0.0766 | No | ||
| 6 | P2RX3 | P2rx3 | 480 | 1.802 | 0.0868 | No | ||
| 7 | PLCG2 | Plcg2 | 551 | 1.717 | 0.0970 | No | ||
| 8 | SLC8A2 | Slc8a2 | 608 | 1.650 | 0.1075 | No | ||
| 9 | SPHK1 | Sphk1 | 621 | 1.628 | 0.1207 | No | ||
| 10 | ADRB1 | Adrb1 | 647 | 1.600 | 0.1328 | No | ||
| 11 | PLCD1 | Plcd1 | 710 | 1.541 | 0.1419 | No | ||
| 12 | TACR3 | Tacr3 | 817 | 1.442 | 0.1474 | No | ||
| 13 | PLCE1 | Plce1 | 837 | 1.426 | 0.1584 | No | ||
| 14 | ATP2A3 | Atp2a3 | 923 | 1.350 | 0.1644 | No | ||
| 15 | PRKCG | Prkcg | 996 | 1.303 | 0.1709 | No | ||
| 16 | ITPKB | Itpkb | 1264 | 1.115 | 0.1631 | No | ||
| 17 | P2RX4 | P2rx4 | 1329 | 1.083 | 0.1682 | No | ||
| 18 | ATP2A1 | Atp2a1 | 1354 | 1.073 | 0.1759 | No | ||
| 19 | MYLK2 | Mylk2 | 1459 | 1.027 | 0.1779 | No | ||
| 20 | ITPR2 | Itpr2 | 1579 | 0.974 | 0.1785 | No | ||
| 21 | PLCB4 | Plcb4 | 1596 | 0.969 | 0.1858 | No | ||
| 22 | PTGER1 | Ptger1 | 1695 | 0.934 | 0.1874 | No | ||
| 23 | GNAL | Gnal | 1772 | 0.905 | 0.1902 | No | ||
| 24 | PLCD4 | Plcd4 | 1840 | 0.882 | 0.1934 | No | ||
| 25 | TNNC1 | Tnnc1 | 2252 | 0.773 | 0.1733 | No | ||
| 26 | CACNA1B | Cacna1b | 2361 | 0.738 | 0.1726 | No | ||
| 27 | PDE1C | Pde1c | 2365 | 0.735 | 0.1787 | No | ||
| 28 | ADCY9 | Adcy9 | 2390 | 0.729 | 0.1834 | No | ||
| 29 | CHRM1 | Chrm1 | 2456 | 0.707 | 0.1852 | No | ||
| 30 | ERBB2 | Erbb2 | 2475 | 0.702 | 0.1900 | No | ||
| 31 | HRH1 | Hrh1 | 2606 | 0.667 | 0.1873 | No | ||
| 32 | ADORA2B | Adora2b | 2645 | 0.656 | 0.1905 | No | ||
| 33 | P2RX2 | P2rx2 | 2676 | 0.649 | 0.1941 | No | ||
| 34 | ADCY3 | Adcy3 | 2687 | 0.646 | 0.1990 | No | ||
| 35 | GRIN2D | Grin2d | 2733 | 0.635 | 0.2015 | No | ||
| 36 | ERBB3 | Erbb3 | 3192 | 0.536 | 0.1763 | No | ||
| 37 | ADRA1A | Adra1a | 3415 | 0.494 | 0.1660 | No | ||
| 38 | HTR5A | Htr5a | 3816 | 0.432 | 0.1437 | No | ||
| 39 | CACNA1H | Cacna1h | 3896 | 0.430 | 0.1422 | No | ||
| 40 | PTAFR | Ptafr | 3900 | 0.429 | 0.1457 | No | ||
| 41 | GRIN2C | Grin2c | 4219 | 0.407 | 0.1285 | No | ||
| 42 | RYR2 | Ryr2 | 4307 | 0.394 | 0.1262 | No | ||
| 43 | CALM1 | Calm1 | 4354 | 0.386 | 0.1265 | No | ||
| 44 | ITPR3 | Itpr3 | 4419 | 0.376 | 0.1256 | No | ||
| 45 | TACR2 | Tacr2 | 4472 | 0.366 | 0.1254 | No | ||
| 46 | MYLK3 | Mylk3 | 4545 | 0.354 | 0.1237 | No | ||
| 47 | PHKG1 | Phkg1 | 4588 | 0.345 | 0.1239 | No | ||
| 48 | PHKA1 | Phka1 | 4590 | 0.345 | 0.1268 | No | ||
| 49 | GRM5 | Grm5 | 4660 | 0.333 | 0.1252 | No | ||
| 50 | TBXA2R | Tbxa2r | 4892 | 0.299 | 0.1127 | No | ||
| 51 | P2RX6 | P2rx6 | 4979 | 0.289 | 0.1096 | No | ||
| 52 | GNAS | Gnas | 5004 | 0.286 | 0.1105 | No | ||
| 53 | PTGER3 | Ptger3 | 5145 | 0.270 | 0.1037 | No | ||
| 54 | CHP1 | Chp1 | 5427 | 0.231 | 0.0874 | No | ||
| 55 | BDKRB2 | Bdkrb2 | 5530 | 0.216 | 0.0826 | No | ||
| 56 | F2R | F2r | 5581 | 0.211 | 0.0811 | No | ||
| 57 | PLCD3 | Plcd3 | 5646 | 0.201 | 0.0787 | No | ||
| 58 | SLC25A4 | Slc25a4 | 5658 | 0.199 | 0.0797 | No | ||
| 59 | HTR4 | Htr4 | 5770 | 0.184 | 0.0741 | No | ||
| 60 | PHKG2 | Phkg2 | 5842 | 0.176 | 0.0709 | No | ||
| 61 | HTR2A | Htr2a | 6033 | 0.152 | 0.0599 | No | ||
| 62 | VDAC3 | Vdac3 | 6062 | 0.149 | 0.0593 | No | ||
| 63 | ITPKA | Itpka | 6253 | 0.130 | 0.0481 | No | ||
| 64 | PLCB2 | Plcb2 | 6348 | 0.118 | 0.0430 | No | ||
| 65 | PPP3CC | Ppp3cc | 6542 | 0.096 | 0.0312 | No | ||
| 66 | CACNA1S | Cacna1s | 6583 | 0.091 | 0.0294 | No | ||
| 67 | PHKB | Phkb | 6754 | 0.071 | 0.0190 | No | ||
| 68 | CAMK4 | Camk4 | 6790 | 0.068 | 0.0173 | No | ||
| 69 | ADCY2 | Adcy2 | 6832 | 0.063 | 0.0151 | No | ||
| 70 | ADCY4 | Adcy4 | 6834 | 0.063 | 0.0156 | No | ||
| 71 | CALML3 | Calml3 | 7106 | 0.034 | -0.0017 | No | ||
| 72 | ATP2B3 | Atp2b3 | 7139 | 0.032 | -0.0035 | No | ||
| 73 | LTB4R2 | Ltb4r2 | 7161 | 0.028 | -0.0047 | No | ||
| 74 | EDNRB | Ednrb | 7287 | 0.015 | -0.0127 | No | ||
| 75 | AVPR1B | Avpr1b | 7492 | 0.000 | -0.0260 | No | ||
| 76 | AVPR1A | Avpr1a | 7493 | 0.000 | -0.0260 | No | ||
| 77 | CHRM5 | Chrm5 | 7538 | 0.000 | -0.0288 | No | ||
| 78 | NTSR1 | Ntsr1 | 7543 | 0.000 | -0.0291 | No | ||
| 79 | ERBB4 | Erbb4 | 7547 | 0.000 | -0.0293 | No | ||
| 80 | ATP2B2 | Atp2b2 | 7591 | 0.000 | -0.0321 | No | ||
| 81 | PLN | Pln | 7638 | 0.000 | -0.0351 | No | ||
| 82 | PPP3R2 | Ppp3r2 | 7663 | 0.000 | -0.0366 | No | ||
| 83 | CACNA1F | Cacna1f | 7745 | 0.000 | -0.0419 | No | ||
| 84 | TNNC2 | Tnnc2 | 7782 | 0.000 | -0.0442 | No | ||
| 85 | HRH2 | Hrh2 | 7875 | 0.000 | -0.0502 | No | ||
| 86 | DRD1 | Drd1 | 8002 | -0.009 | -0.0583 | No | ||
| 87 | ADORA2A | Adora2a | 8092 | -0.018 | -0.0640 | No | ||
| 88 | SPHK2 | Sphk2 | 8222 | -0.028 | -0.0721 | No | ||
| 89 | RYR1 | Ryr1 | 8411 | -0.045 | -0.0840 | No | ||
| 90 | ATP2A2 | Atp2a2 | 8693 | -0.074 | -0.1016 | No | ||
| 91 | VDAC2 | Vdac2 | 8746 | -0.079 | -0.1043 | No | ||
| 92 | ADRA1D | Adra1d | 8756 | -0.080 | -0.1042 | No | ||
| 93 | NOS1 | Nos1 | 8757 | -0.080 | -0.1036 | No | ||
| 94 | GNA11 | Gna11 | 9020 | -0.106 | -0.1197 | No | ||
| 95 | CYSLTR1 | Cysltr1 | 9042 | -0.109 | -0.1201 | No | ||
| 96 | CCKAR | Cckar | 9103 | -0.116 | -0.1230 | No | ||
| 97 | P2RX1 | P2rx1 | 9227 | -0.127 | -0.1300 | No | ||
| 98 | HTR2B | Htr2b | 9391 | -0.143 | -0.1393 | No | ||
| 99 | PLCB3 | Plcb3 | 9618 | -0.166 | -0.1526 | No | ||
| 100 | PLCG1 | Plcg1 | 9622 | -0.167 | -0.1514 | No | ||
| 101 | CALM3 | Calm3 | 9742 | -0.178 | -0.1576 | No | ||
| 102 | PPID | Ppid | 9929 | -0.196 | -0.1680 | No | ||
| 103 | PPP3CA | Ppp3ca | 10018 | -0.206 | -0.1720 | No | ||
| 104 | CALM2 | Calm2 | 10048 | -0.209 | -0.1721 | No | ||
| 105 | CAMK2G | Camk2g | 10068 | -0.211 | -0.1715 | No | ||
| 106 | VDAC1 | Vdac1 | 10290 | -0.233 | -0.1839 | No | ||
| 107 | PRKACB | Prkacb | 10431 | -0.249 | -0.1909 | No | ||
| 108 | GNA14 | Gna14 | 10481 | -0.252 | -0.1919 | No | ||
| 109 | SLC25A5 | Slc25a5 | 10636 | -0.269 | -0.1996 | No | ||
| 110 | BST1 | Bst1 | 10910 | -0.296 | -0.2149 | No | ||
| 111 | CAMK2A | Camk2a | 11400 | -0.353 | -0.2437 | No | ||
| 112 | ADRA1B | Adra1b | 11495 | -0.365 | -0.2467 | No | ||
| 113 | PPP3CB | Ppp3cb | 11662 | -0.384 | -0.2542 | No | ||
| 114 | ATP2B1 | Atp2b1 | 11724 | -0.392 | -0.2548 | No | ||
| 115 | AGTR1 | Agtr1a | 11925 | -0.418 | -0.2642 | No | ||
| 116 | SLC25A31 | Slc25a31 | 11959 | -0.418 | -0.2628 | No | ||
| 117 | TACR1 | Tacr1 | 11982 | -0.418 | -0.2606 | No | ||
| 118 | PRKACA | Prkaca | 12054 | -0.424 | -0.2616 | No | ||
| 119 | HTR6 | Htr6 | 12406 | -0.458 | -0.2805 | No | ||
| 120 | CACNA1E | Cacna1e | 12413 | -0.459 | -0.2770 | No | ||
| 121 | MYLK | Mylk | 12500 | -0.474 | -0.2785 | No | ||
| 122 | PHKA2 | Phka2 | 12527 | -0.479 | -0.2761 | No | ||
| 123 | PTK2B | Ptk2b | 12544 | -0.483 | -0.2730 | No | ||
| 124 | CHP2 | Chp2 | 12674 | -0.499 | -0.2771 | No | ||
| 125 | HTR7 | Htr7 | 12756 | -0.508 | -0.2780 | No | ||
| 126 | GRIN2A | Grin2a | 12999 | -0.553 | -0.2890 | No | ||
| 127 | EDNRA | Ednra | 13143 | -0.585 | -0.2933 | No | ||
| 128 | OXTR | Oxtr | 13326 | -0.625 | -0.2998 | No | ||
| 129 | ITPR1 | Itpr1 | 13613 | -0.701 | -0.3124 | Yes | ||
| 130 | PDE1B | Pde1b | 13655 | -0.712 | -0.3090 | Yes | ||
| 131 | CHRM3 | Chrm3 | 13776 | -0.750 | -0.3103 | Yes | ||
| 132 | CACNA1D | Cacna1d | 13877 | -0.781 | -0.3102 | Yes | ||
| 133 | GRM1 | Grm1 | 13914 | -0.793 | -0.3057 | Yes | ||
| 134 | CACNA1A | Cacna1a | 14005 | -0.819 | -0.3045 | Yes | ||
| 135 | TRHR | Trhr | 14079 | -0.842 | -0.3021 | Yes | ||
| 136 | NOS2 | Nos2 | 14138 | -0.860 | -0.2985 | Yes | ||
| 137 | EGFR | Egfr | 14176 | -0.870 | -0.2934 | Yes | ||
| 138 | TRPC1 | Trpc1 | 14289 | -0.920 | -0.2928 | Yes | ||
| 139 | SLC8A3 | Slc8a3 | 14459 | -1.001 | -0.2952 | Yes | ||
| 140 | GNA15 | Gna15 | 14497 | -1.023 | -0.2889 | Yes | ||
| 141 | ADCY7 | Adcy7 | 14683 | -1.147 | -0.2911 | Yes | ||
| 142 | CD38 | Cd38 | 14688 | -1.151 | -0.2815 | Yes | ||
| 143 | PRKCA | Prkca | 14737 | -1.199 | -0.2743 | Yes | ||
| 144 | GRPR | Grpr | 14779 | -1.235 | -0.2664 | Yes | ||
| 145 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.2585 | Yes | ||
| 146 | PLCB1 | Plcb1 | 15032 | -1.546 | -0.2584 | Yes | ||
| 147 | CACNA1C | Cacna1c | 15057 | -1.590 | -0.2464 | Yes | ||
| 148 | CACNA1G | Cacna1g | 15111 | -1.688 | -0.2353 | Yes | ||
| 149 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.2233 | Yes | ||
| 150 | NOS3 | Nos3 | 15187 | -1.828 | -0.2092 | Yes | ||
| 151 | PTGFR | Ptgfr | 15226 | -1.922 | -0.1952 | Yes | ||
| 152 | ATP2B4 | Atp2b4 | 15310 | -2.170 | -0.1820 | Yes | ||
| 153 | CAMK2B | Camk2b | 15320 | -2.213 | -0.1636 | Yes | ||
| 154 | ADCY8 | Adcy8 | 15381 | -2.433 | -0.1466 | Yes | ||
| 155 | P2RX7 | P2rx7 | 15418 | -2.691 | -0.1259 | Yes | ||
| 156 | BDKRB1 | Bdkrb1 | 15429 | -2.734 | -0.1031 | Yes | ||
| 157 | PRKCB | Prkcb | 15461 | -2.954 | -0.0798 | Yes | ||
| 158 | CCKBR | Cckbr | 15465 | -3.027 | -0.0540 | Yes | ||
| 159 | CACNA1I | Cacna1i | 15487 | -3.296 | -0.0272 | Yes | ||
| 160 | PDE1A | Pde1a | 15499 | -3.449 | 0.0017 | Yes |