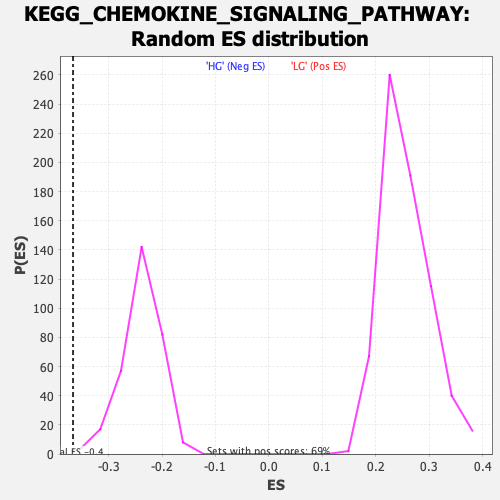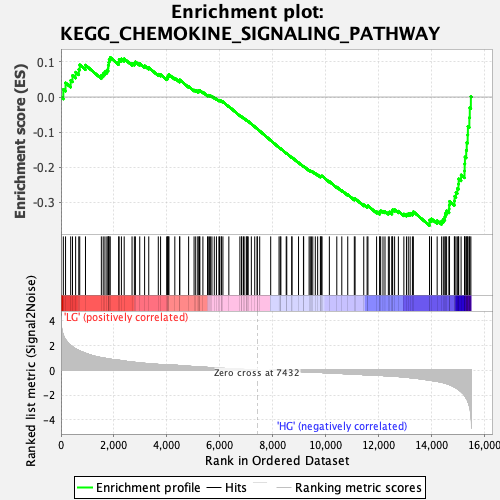
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_CHEMOKINE_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.36645043 |
| Normalized Enrichment Score (NES) | -1.5362176 |
| Nominal p-value | 0.003236246 |
| FDR q-value | 0.19729522 |
| FWER p-Value | 0.704 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GNG4 | Gng4 | 88 | 2.780 | 0.0218 | No | ||
| 2 | ITK | Itk | 172 | 2.428 | 0.0404 | No | ||
| 3 | ADCY1 | Adcy1 | 366 | 1.982 | 0.0475 | No | ||
| 4 | RASGRP2 | Rasgrp2 | 435 | 1.866 | 0.0616 | No | ||
| 5 | CXCL12 | Cxcl12 | 553 | 1.714 | 0.0709 | No | ||
| 6 | LYN | Lyn | 673 | 1.577 | 0.0788 | No | ||
| 7 | ELMO1 | Elmo1 | 709 | 1.541 | 0.0918 | No | ||
| 8 | GNG3 | Gng3 | 926 | 1.348 | 0.0911 | No | ||
| 9 | GNG8 | Gng8 | 1526 | 0.996 | 0.0620 | No | ||
| 10 | PLCB4 | Plcb4 | 1596 | 0.969 | 0.0671 | No | ||
| 11 | PREX1 | Prex1 | 1662 | 0.948 | 0.0722 | No | ||
| 12 | GNAI1 | Gnai1 | 1736 | 0.921 | 0.0766 | No | ||
| 13 | GNG10 | Gng10 | 1783 | 0.900 | 0.0825 | No | ||
| 14 | GNG13 | Gng13 | 1785 | 0.900 | 0.0914 | No | ||
| 15 | PRKCD | Prkcd | 1807 | 0.893 | 0.0988 | No | ||
| 16 | STAT1 | Stat1 | 1819 | 0.890 | 0.1069 | No | ||
| 17 | CXCR4 | Cxcr4 | 1862 | 0.875 | 0.1129 | No | ||
| 18 | HRAS | Hras | 2180 | 0.791 | 0.1001 | No | ||
| 19 | CCR2 | Ccr2 | 2197 | 0.789 | 0.1069 | No | ||
| 20 | STAT2 | Stat2 | 2285 | 0.762 | 0.1088 | No | ||
| 21 | ADCY9 | Adcy9 | 2390 | 0.729 | 0.1092 | No | ||
| 22 | ADCY3 | Adcy3 | 2687 | 0.646 | 0.0963 | No | ||
| 23 | VAV2 | Vav2 | 2775 | 0.625 | 0.0969 | No | ||
| 24 | IKBKG | Ikbkg | 2814 | 0.616 | 0.1005 | No | ||
| 25 | SHC1 | Shc1 | 2978 | 0.579 | 0.0956 | No | ||
| 26 | PIK3CG | Pik3cg | 3162 | 0.541 | 0.0891 | No | ||
| 27 | STAT5B | Stat5b | 3319 | 0.511 | 0.0840 | No | ||
| 28 | NRAS | Nras | 3680 | 0.455 | 0.0651 | No | ||
| 29 | NFKBIB | Nfkbib | 3766 | 0.441 | 0.0639 | No | ||
| 30 | CCL27 | Ccl27a | 3997 | 0.421 | 0.0531 | No | ||
| 31 | CCR6 | Ccr6 | 4033 | 0.421 | 0.0550 | No | ||
| 32 | CCR3 | Ccr3 | 4034 | 0.421 | 0.0592 | No | ||
| 33 | GRK5 | Grk5 | 4072 | 0.415 | 0.0609 | No | ||
| 34 | PIK3R5 | Pik3r5 | 4080 | 0.414 | 0.0645 | No | ||
| 35 | PARD3 | Pard3 | 4305 | 0.394 | 0.0539 | No | ||
| 36 | GNB5 | Gnb5 | 4486 | 0.364 | 0.0458 | No | ||
| 37 | CXCL16 | Cxcl16 | 4492 | 0.362 | 0.0490 | No | ||
| 38 | RAC2 | Rac2 | 4827 | 0.309 | 0.0304 | No | ||
| 39 | IKBKB | Ikbkb | 5033 | 0.283 | 0.0198 | No | ||
| 40 | PIK3CD | Pik3cd | 5095 | 0.274 | 0.0186 | No | ||
| 41 | KRAS | Kras | 5181 | 0.266 | 0.0157 | No | ||
| 42 | WASL | Wasl | 5185 | 0.265 | 0.0181 | No | ||
| 43 | PRKCZ | Prkcz | 5239 | 0.256 | 0.0172 | No | ||
| 44 | GRK4 | Grk4 | 5252 | 0.255 | 0.0189 | No | ||
| 45 | CCR7 | Ccr7 | 5367 | 0.239 | 0.0139 | No | ||
| 46 | GNB2 | Gnb2 | 5542 | 0.215 | 0.0047 | No | ||
| 47 | STAT3 | Stat3 | 5568 | 0.212 | 0.0052 | No | ||
| 48 | ADCY6 | Adcy6 | 5613 | 0.207 | 0.0044 | No | ||
| 49 | TIAM1 | Tiam1 | 5659 | 0.199 | 0.0034 | No | ||
| 50 | CCR10 | Ccr10 | 5709 | 0.194 | 0.0021 | No | ||
| 51 | GNG11 | Gng11 | 5798 | 0.181 | -0.0018 | No | ||
| 52 | GSK3B | Gsk3b | 5879 | 0.171 | -0.0053 | No | ||
| 53 | GNB1 | Gnb1 | 5973 | 0.160 | -0.0098 | No | ||
| 54 | GNG7 | Gng7 | 5994 | 0.157 | -0.0095 | No | ||
| 55 | CCR4 | Ccr4 | 6059 | 0.149 | -0.0122 | No | ||
| 56 | CSK | Csk | 6066 | 0.149 | -0.0111 | No | ||
| 57 | GNG12 | Gng12 | 6124 | 0.143 | -0.0134 | No | ||
| 58 | PLCB2 | Plcb2 | 6348 | 0.118 | -0.0267 | No | ||
| 59 | ARRB2 | Arrb2 | 6766 | 0.070 | -0.0532 | No | ||
| 60 | ADCY2 | Adcy2 | 6832 | 0.063 | -0.0568 | No | ||
| 61 | ADCY4 | Adcy4 | 6834 | 0.063 | -0.0562 | No | ||
| 62 | RHOA | Rhoa | 6904 | 0.056 | -0.0601 | No | ||
| 63 | CRK | Crk | 6925 | 0.054 | -0.0609 | No | ||
| 64 | PIK3CB | Pik3cb | 7007 | 0.045 | -0.0657 | No | ||
| 65 | GNAI3 | Gnai3 | 7030 | 0.042 | -0.0667 | No | ||
| 66 | AKT2 | Akt2 | 7049 | 0.039 | -0.0675 | No | ||
| 67 | RAP1B | Rap1b | 7080 | 0.036 | -0.0691 | No | ||
| 68 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.0766 | No | ||
| 69 | RELA | Rela | 7327 | 0.011 | -0.0847 | No | ||
| 70 | GNG5 | Gm4356 | 7417 | 0.002 | -0.0905 | No | ||
| 71 | BCAR1 | Bcar1 | 7425 | 0.001 | -0.0909 | No | ||
| 72 | VAV1 | Vav1 | 7515 | 0.000 | -0.0967 | No | ||
| 73 | GRK6 | Grk6 | 7931 | -0.003 | -0.1237 | No | ||
| 74 | ADCY5 | Adcy5 | 8247 | -0.030 | -0.1439 | No | ||
| 75 | AKT1 | Akt1 | 8293 | -0.035 | -0.1464 | No | ||
| 76 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.1475 | No | ||
| 77 | GRB2 | Grb2 | 8518 | -0.055 | -0.1600 | No | ||
| 78 | RAF1 | Raf1 | 8539 | -0.058 | -0.1608 | No | ||
| 79 | GNAI2 | Gnai2 | 8727 | -0.077 | -0.1722 | No | ||
| 80 | PPBP | Ppbp | 8737 | -0.078 | -0.1720 | No | ||
| 81 | JAK2 | Jak2 | 8984 | -0.102 | -0.1870 | No | ||
| 82 | SOS2 | Sos2 | 9170 | -0.122 | -0.1978 | No | ||
| 83 | BRAF | Braf | 9178 | -0.123 | -0.1970 | No | ||
| 84 | CDC42 | Cdc42 | 9384 | -0.142 | -0.2090 | No | ||
| 85 | SOS1 | Sos1 | 9428 | -0.146 | -0.2103 | No | ||
| 86 | GNB4 | Gnb4 | 9471 | -0.150 | -0.2116 | No | ||
| 87 | GSK3A | Gsk3a | 9523 | -0.156 | -0.2133 | No | ||
| 88 | PLCB3 | Plcb3 | 9618 | -0.166 | -0.2178 | No | ||
| 89 | NFKBIA | Nfkbia | 9707 | -0.174 | -0.2218 | No | ||
| 90 | ROCK1 | Rock1 | 9810 | -0.184 | -0.2266 | No | ||
| 91 | CCL20 | Ccl20 | 9827 | -0.186 | -0.2258 | No | ||
| 92 | GRK2 | Grk2 | 9845 | -0.188 | -0.2250 | No | ||
| 93 | JAK3 | Jak3 | 9873 | -0.191 | -0.2249 | No | ||
| 94 | CRKL | Crkl | 10150 | -0.218 | -0.2407 | No | ||
| 95 | PRKACB | Prkacb | 10431 | -0.249 | -0.2565 | No | ||
| 96 | RAP1A | Rap1a | 10620 | -0.267 | -0.2660 | No | ||
| 97 | CHUK | Chuk | 10845 | -0.290 | -0.2777 | No | ||
| 98 | GRK1 | Grk1 | 11094 | -0.319 | -0.2907 | No | ||
| 99 | ROCK2 | Rock2 | 11123 | -0.321 | -0.2894 | No | ||
| 100 | RAC1 | Rac1 | 11450 | -0.360 | -0.3070 | No | ||
| 101 | CX3CR1 | Cx3cr1 | 11578 | -0.373 | -0.3116 | No | ||
| 102 | PXN | Pxn | 11609 | -0.377 | -0.3098 | No | ||
| 103 | GNGT2 | Gngt2 | 11936 | -0.418 | -0.3268 | No | ||
| 104 | PRKACA | Prkaca | 12054 | -0.424 | -0.3302 | No | ||
| 105 | GRK3 | Grk3 | 12058 | -0.424 | -0.3262 | No | ||
| 106 | CXCR5 | Cxcr5 | 12091 | -0.424 | -0.3241 | No | ||
| 107 | PTK2 | Ptk2 | 12178 | -0.432 | -0.3254 | No | ||
| 108 | CXCL9 | Cxcl9 | 12254 | -0.440 | -0.3260 | No | ||
| 109 | GNB3 | Gnb3 | 12381 | -0.453 | -0.3297 | No | ||
| 110 | GNG2 | Gng2 | 12418 | -0.460 | -0.3275 | No | ||
| 111 | FGR | Fgr | 12518 | -0.479 | -0.3292 | No | ||
| 112 | CCL22 | Ccl22 | 12519 | -0.479 | -0.3244 | No | ||
| 113 | PTK2B | Ptk2b | 12544 | -0.483 | -0.3212 | No | ||
| 114 | AKT3 | Akt3 | 12618 | -0.490 | -0.3211 | No | ||
| 115 | PIK3R1 | Pik3r1 | 12760 | -0.509 | -0.3252 | No | ||
| 116 | TIAM2 | Tiam2 | 12968 | -0.543 | -0.3333 | No | ||
| 117 | CCR9 | Ccr9 | 13066 | -0.568 | -0.3340 | No | ||
| 118 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.3326 | No | ||
| 119 | PAK1 | Pak1 | 13205 | -0.599 | -0.3312 | No | ||
| 120 | FOXO3 | Foxo3 | 13292 | -0.619 | -0.3307 | No | ||
| 121 | PIK3CA | Pik3ca | 13327 | -0.625 | -0.3267 | No | ||
| 122 | NCF1 | Ncf1 | 13939 | -0.799 | -0.3585 | Yes | ||
| 123 | CXCL6 | Cxcl5 | 13940 | -0.800 | -0.3506 | Yes | ||
| 124 | SHC3 | Shc3 | 14013 | -0.825 | -0.3471 | Yes | ||
| 125 | DOCK2 | Dock2 | 14227 | -0.892 | -0.3522 | Yes | ||
| 126 | WAS | Was | 14396 | -0.970 | -0.3535 | Yes | ||
| 127 | CXCR2 | Cxcr2 | 14460 | -1.001 | -0.3477 | Yes | ||
| 128 | CXCR6 | Cxcr6 | 14519 | -1.034 | -0.3412 | Yes | ||
| 129 | CCL28 | Ccl28 | 14534 | -1.046 | -0.3318 | Yes | ||
| 130 | XCR1 | Xcr1 | 14584 | -1.073 | -0.3243 | Yes | ||
| 131 | NFKB1 | Nfkb1 | 14677 | -1.142 | -0.3190 | Yes | ||
| 132 | ADCY7 | Adcy7 | 14683 | -1.147 | -0.3080 | Yes | ||
| 133 | SHC4 | Shc4 | 14695 | -1.160 | -0.2972 | Yes | ||
| 134 | CX3CL1 | Cx3cl1 | 14878 | -1.343 | -0.2957 | Yes | ||
| 135 | VAV3 | Vav3 | 14888 | -1.356 | -0.2829 | Yes | ||
| 136 | CXCL11 | Cxcl11 | 14938 | -1.408 | -0.2721 | Yes | ||
| 137 | ARRB1 | Arrb1 | 14990 | -1.490 | -0.2607 | Yes | ||
| 138 | PLCB1 | Plcb1 | 15032 | -1.546 | -0.2481 | Yes | ||
| 139 | CCR1 | Ccr1 | 15042 | -1.555 | -0.2333 | Yes | ||
| 140 | CCL7 | Ccl7 | 15132 | -1.719 | -0.2220 | Yes | ||
| 141 | HCK | Hck | 15262 | -2.026 | -0.2103 | Yes | ||
| 142 | CCL2 | Ccl2 | 15268 | -2.037 | -0.1905 | Yes | ||
| 143 | CCL5 | Ccl5 | 15278 | -2.073 | -0.1706 | Yes | ||
| 144 | CCL8 | Ccl12 | 15326 | -2.242 | -0.1514 | Yes | ||
| 145 | CCL17 | Ccl17 | 15346 | -2.308 | -0.1298 | Yes | ||
| 146 | ADCY8 | Adcy8 | 15381 | -2.433 | -0.1079 | Yes | ||
| 147 | CXCL10 | Cxcl10 | 15390 | -2.511 | -0.0836 | Yes | ||
| 148 | CCL11 | Ccl11 | 15445 | -2.856 | -0.0588 | Yes | ||
| 149 | PRKCB | Prkcb | 15461 | -2.954 | -0.0306 | Yes | ||
| 150 | CXCL14 | Cxcl14 | 15502 | -3.500 | 0.0015 | Yes |