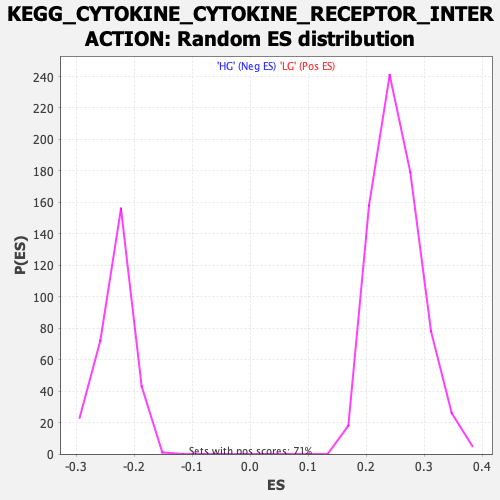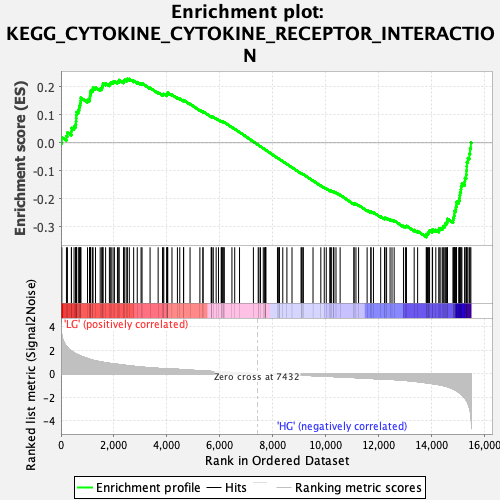
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_CYTOKINE_CYTOKINE_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | -0.33701554 |
| Normalized Enrichment Score (NES) | -1.4561291 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.21344237 |
| FWER p-Value | 0.902 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDR | Kdr | 32 | 3.271 | 0.0198 | No | ||
| 2 | GDF5 | Gdf5 | 206 | 2.345 | 0.0243 | No | ||
| 3 | FLT4 | Flt4 | 243 | 2.244 | 0.0369 | No | ||
| 4 | TNFRSF25 | Tnfrsf25 | 392 | 1.944 | 0.0403 | No | ||
| 5 | TNFRSF21 | Tnfrsf21 | 396 | 1.940 | 0.0531 | No | ||
| 6 | IL7 | Il7 | 494 | 1.777 | 0.0587 | No | ||
| 7 | CXCL12 | Cxcl12 | 553 | 1.714 | 0.0664 | No | ||
| 8 | TNFRSF19 | Tnfrsf19 | 566 | 1.700 | 0.0770 | No | ||
| 9 | IFNLR1 | Ifnlr1 | 574 | 1.690 | 0.0879 | No | ||
| 10 | FAS | Fas | 579 | 1.684 | 0.0989 | No | ||
| 11 | IL6 | Il6 | 585 | 1.675 | 0.1098 | No | ||
| 12 | FLT3LG | Flt3l | 652 | 1.598 | 0.1162 | No | ||
| 13 | IL17RA | Il17ra | 685 | 1.563 | 0.1246 | No | ||
| 14 | PRL | Prl | 704 | 1.545 | 0.1337 | No | ||
| 15 | CRLF2 | Crlf2 | 726 | 1.522 | 0.1426 | No | ||
| 16 | CD40 | Cd40 | 745 | 1.502 | 0.1514 | No | ||
| 17 | FLT3 | Flt3 | 750 | 1.498 | 0.1612 | No | ||
| 18 | TNFRSF11A | Tnfrsf11a | 1003 | 1.295 | 0.1535 | No | ||
| 19 | TGFB2 | Tgfb2 | 1078 | 1.240 | 0.1569 | No | ||
| 20 | TNFSF13 | Tnfsf13 | 1093 | 1.225 | 0.1642 | No | ||
| 21 | VEGFB | Vegfb | 1101 | 1.223 | 0.1720 | No | ||
| 22 | VEGFC | Vegfc | 1115 | 1.215 | 0.1793 | No | ||
| 23 | IL18R1 | Il18r1 | 1126 | 1.207 | 0.1867 | No | ||
| 24 | IL15RA | Il15ra | 1198 | 1.152 | 0.1898 | No | ||
| 25 | AMHR2 | Amhr2 | 1208 | 1.146 | 0.1969 | No | ||
| 26 | IL15 | Il15 | 1304 | 1.096 | 0.1980 | No | ||
| 27 | IL12RB1 | Il12rb1 | 1482 | 1.015 | 0.1933 | No | ||
| 28 | TNFRSF1B | Tnfrsf1b | 1543 | 0.989 | 0.1960 | No | ||
| 29 | ACVR2A | Acvr2a | 1556 | 0.984 | 0.2018 | No | ||
| 30 | KITLG | Kitl | 1580 | 0.974 | 0.2068 | No | ||
| 31 | AMH | Amh | 1590 | 0.971 | 0.2127 | No | ||
| 32 | INHBB | Inhbb | 1687 | 0.938 | 0.2128 | No | ||
| 33 | TNFRSF14 | Tnfrsf14 | 1830 | 0.887 | 0.2095 | No | ||
| 34 | CXCR4 | Cxcr4 | 1862 | 0.875 | 0.2133 | No | ||
| 35 | IL2RA | Il2ra | 1899 | 0.863 | 0.2167 | No | ||
| 36 | CSF2RA | Csf2ra | 1968 | 0.843 | 0.2179 | No | ||
| 37 | CNTFR | Cntfr | 2020 | 0.830 | 0.2202 | No | ||
| 38 | PDGFB | Pdgfb | 2135 | 0.802 | 0.2181 | No | ||
| 39 | TNFSF15 | Tnfsf15 | 2186 | 0.791 | 0.2202 | No | ||
| 40 | CCR2 | Ccr2 | 2197 | 0.789 | 0.2248 | No | ||
| 41 | IL17RB | Il17rb | 2363 | 0.736 | 0.2190 | No | ||
| 42 | TGFBR1 | Tgfbr1 | 2382 | 0.730 | 0.2227 | No | ||
| 43 | IL9R | Il9r | 2415 | 0.721 | 0.2254 | No | ||
| 44 | RELT | Relt | 2491 | 0.697 | 0.2252 | No | ||
| 45 | CSF3 | Csf3 | 2509 | 0.690 | 0.2287 | No | ||
| 46 | TNFRSF1A | Tnfrsf1a | 2584 | 0.672 | 0.2284 | No | ||
| 47 | LTBR | Ltbr | 2748 | 0.631 | 0.2220 | No | ||
| 48 | IL10RB | Il10rb | 2884 | 0.600 | 0.2172 | No | ||
| 49 | PDGFA | Pdgfa | 3028 | 0.566 | 0.2117 | No | ||
| 50 | IL4 | Il4 | 3065 | 0.559 | 0.2131 | No | ||
| 51 | IL1R2 | Il1r2 | 3369 | 0.503 | 0.1967 | No | ||
| 52 | TNFRSF12A | Tnfrsf12a | 3674 | 0.457 | 0.1800 | No | ||
| 53 | MPL | Mpl | 3841 | 0.432 | 0.1721 | No | ||
| 54 | IL1B | Il1b | 3859 | 0.432 | 0.1738 | No | ||
| 55 | IL12RB2 | Il12rb2 | 3888 | 0.431 | 0.1749 | No | ||
| 56 | CCL27 | Ccl27a | 3997 | 0.421 | 0.1707 | No | ||
| 57 | TNFSF8 | Tnfsf8 | 4013 | 0.421 | 0.1725 | No | ||
| 58 | TNFRSF17 | Tnfrsf17 | 4020 | 0.421 | 0.1749 | No | ||
| 59 | CCR6 | Ccr6 | 4033 | 0.421 | 0.1770 | No | ||
| 60 | CCR3 | Ccr3 | 4034 | 0.421 | 0.1798 | No | ||
| 61 | EDAR | Edar | 4197 | 0.407 | 0.1720 | No | ||
| 62 | CD27 | Cd27 | 4403 | 0.379 | 0.1611 | No | ||
| 63 | CXCL16 | Cxcl16 | 4492 | 0.362 | 0.1578 | No | ||
| 64 | TNFSF12 | Tnfsf12 | 4634 | 0.338 | 0.1509 | No | ||
| 65 | TNFRSF4 | Tnfrsf4 | 4639 | 0.337 | 0.1529 | No | ||
| 66 | LTA | Lta | 4881 | 0.300 | 0.1392 | No | ||
| 67 | IFNAR1 | Ifnar1 | 5253 | 0.254 | 0.1167 | No | ||
| 68 | CCR7 | Ccr7 | 5367 | 0.239 | 0.1110 | No | ||
| 69 | EDA | Eda | 5372 | 0.238 | 0.1123 | No | ||
| 70 | ACVR2B | Acvr2b | 5684 | 0.197 | 0.0933 | No | ||
| 71 | IL1A | Il1a | 5686 | 0.197 | 0.0946 | No | ||
| 72 | CCR10 | Ccr10 | 5709 | 0.194 | 0.0945 | No | ||
| 73 | EDA2R | Eda2r | 5760 | 0.186 | 0.0925 | No | ||
| 74 | IFNB1 | Ifnb1 | 5868 | 0.171 | 0.0866 | No | ||
| 75 | CSF1R | Csf1r | 5957 | 0.162 | 0.0820 | No | ||
| 76 | CCR4 | Ccr4 | 6059 | 0.149 | 0.0764 | No | ||
| 77 | TNFSF10 | Tnfsf10 | 6075 | 0.148 | 0.0764 | No | ||
| 78 | TNFRSF8 | Tnfrsf8 | 6097 | 0.145 | 0.0760 | No | ||
| 79 | IL2 | Il2 | 6130 | 0.142 | 0.0749 | No | ||
| 80 | CSF1 | Csf1 | 6131 | 0.142 | 0.0758 | No | ||
| 81 | IL6ST | Il6st | 6177 | 0.137 | 0.0738 | No | ||
| 82 | TGFB1 | Tgfb1 | 6465 | 0.104 | 0.0558 | No | ||
| 83 | OSMR | Osmr | 6570 | 0.092 | 0.0497 | No | ||
| 84 | TGFBR2 | Tgfbr2 | 6752 | 0.071 | 0.0383 | No | ||
| 85 | ACVR1B | Acvr1b | 7277 | 0.016 | 0.0043 | No | ||
| 86 | IL21R | Il21r | 7459 | 0.000 | -0.0075 | No | ||
| 87 | EPO | Epo | 7487 | 0.000 | -0.0092 | No | ||
| 88 | CSF2RB | Csf2rb | 7546 | 0.000 | -0.0130 | No | ||
| 89 | IL20RA | Il20ra | 7650 | 0.000 | -0.0197 | No | ||
| 90 | TNF | Tnf | 7693 | 0.000 | -0.0225 | No | ||
| 91 | IL20 | Il20 | 7736 | 0.000 | -0.0252 | No | ||
| 92 | IL25 | Il25 | 7737 | 0.000 | -0.0252 | No | ||
| 93 | IL13 | Il13 | 7738 | 0.000 | -0.0252 | No | ||
| 94 | IL7R | Il7r | 7747 | 0.000 | -0.0257 | No | ||
| 95 | IFNGR1 | Ifngr1 | 8187 | -0.027 | -0.0542 | No | ||
| 96 | IL12A | Il12a | 8218 | -0.028 | -0.0559 | No | ||
| 97 | LTB | Ltb | 8253 | -0.031 | -0.0579 | No | ||
| 98 | CLCF1 | Clcf1 | 8261 | -0.032 | -0.0582 | No | ||
| 99 | IL17B | Il17b | 8388 | -0.043 | -0.0661 | No | ||
| 100 | IFNGR2 | Ifngr2 | 8544 | -0.058 | -0.0758 | No | ||
| 101 | PPBP | Ppbp | 8737 | -0.078 | -0.0878 | No | ||
| 102 | KIT | Kit | 9079 | -0.113 | -0.1093 | No | ||
| 103 | ACVRL1 | Acvrl1 | 9102 | -0.116 | -0.1099 | No | ||
| 104 | IL20RB | Il20rb | 9146 | -0.120 | -0.1119 | No | ||
| 105 | BMPR1B | Bmpr1b | 9166 | -0.122 | -0.1123 | No | ||
| 106 | TNFRSF13C | Tnfrsf13c | 9532 | -0.157 | -0.1351 | No | ||
| 107 | CCL20 | Ccl20 | 9827 | -0.186 | -0.1530 | No | ||
| 108 | TGFB3 | Tgfb3 | 9957 | -0.199 | -0.1601 | No | ||
| 109 | IL2RB | Il2rb | 10033 | -0.207 | -0.1636 | No | ||
| 110 | IFNE | Ifne | 10167 | -0.220 | -0.1708 | No | ||
| 111 | TNFSF4 | Tnfsf4 | 10190 | -0.222 | -0.1707 | No | ||
| 112 | FASLG | Fasl | 10236 | -0.227 | -0.1721 | No | ||
| 113 | IL5 | Il5 | 10308 | -0.234 | -0.1752 | No | ||
| 114 | PRLR | Prlr | 10323 | -0.236 | -0.1745 | No | ||
| 115 | IL24 | Il24 | 10401 | -0.245 | -0.1779 | No | ||
| 116 | TNFSF13B | Tnfsf13b | 10557 | -0.261 | -0.1862 | No | ||
| 117 | IL4R | Il4ra | 11081 | -0.318 | -0.2182 | No | ||
| 118 | IL19 | Il19 | 11082 | -0.318 | -0.2161 | No | ||
| 119 | VEGFD | Vegfd | 11156 | -0.326 | -0.2186 | No | ||
| 120 | PDGFC | Pdgfc | 11255 | -0.336 | -0.2228 | No | ||
| 121 | CX3CR1 | Cx3cr1 | 11578 | -0.373 | -0.2413 | No | ||
| 122 | CNTF | Cntf | 11716 | -0.391 | -0.2476 | No | ||
| 123 | BMPR2 | Bmpr2 | 11728 | -0.392 | -0.2457 | No | ||
| 124 | BMPR1A | Bmpr1a | 11818 | -0.405 | -0.2488 | No | ||
| 125 | CXCR5 | Cxcr5 | 12091 | -0.424 | -0.2636 | No | ||
| 126 | IL23R | Il23r | 12236 | -0.440 | -0.2701 | No | ||
| 127 | CXCL9 | Cxcl9 | 12254 | -0.440 | -0.2682 | No | ||
| 128 | INHBE | Inhbe | 12314 | -0.444 | -0.2691 | No | ||
| 129 | LEPR | Lepr | 12446 | -0.465 | -0.2745 | No | ||
| 130 | CCL22 | Ccl22 | 12519 | -0.479 | -0.2760 | No | ||
| 131 | CTF1 | Ctf1 | 12601 | -0.489 | -0.2780 | No | ||
| 132 | VEGFA | Vegfa | 12953 | -0.540 | -0.2973 | No | ||
| 133 | TNFSF9 | Tnfsf9 | 13027 | -0.558 | -0.2983 | No | ||
| 134 | CCR9 | Ccr9 | 13066 | -0.568 | -0.2970 | No | ||
| 135 | TNFSF18 | Tnfsf18 | 13358 | -0.633 | -0.3117 | No | ||
| 136 | TNFRSF11B | Tnfrsf11b | 13488 | -0.665 | -0.3156 | No | ||
| 137 | TSLP | Tslp | 13817 | -0.762 | -0.3319 | Yes | ||
| 138 | GHR | Ghr | 13819 | -0.762 | -0.3269 | Yes | ||
| 139 | MET | Met | 13861 | -0.774 | -0.3244 | Yes | ||
| 140 | IL6R | Il6ra | 13893 | -0.785 | -0.3211 | Yes | ||
| 141 | IL10 | Il10 | 13916 | -0.793 | -0.3173 | Yes | ||
| 142 | CXCL6 | Cxcl5 | 13940 | -0.800 | -0.3134 | Yes | ||
| 143 | BMP7 | Bmp7 | 14043 | -0.834 | -0.3145 | Yes | ||
| 144 | FLT1 | Flt1 | 14053 | -0.837 | -0.3094 | Yes | ||
| 145 | EGFR | Egfr | 14176 | -0.870 | -0.3116 | Yes | ||
| 146 | IL18RAP | Il18rap | 14295 | -0.924 | -0.3131 | Yes | ||
| 147 | ACVR1 | Acvr1 | 14299 | -0.926 | -0.3071 | Yes | ||
| 148 | TNFSF11 | Tnfsf11 | 14366 | -0.956 | -0.3049 | Yes | ||
| 149 | EPOR | Epor | 14436 | -0.988 | -0.3028 | Yes | ||
| 150 | CXCR2 | Cxcr2 | 14460 | -1.001 | -0.2976 | Yes | ||
| 151 | CXCR6 | Cxcr6 | 14519 | -1.034 | -0.2945 | Yes | ||
| 152 | CCL28 | Ccl28 | 14534 | -1.046 | -0.2884 | Yes | ||
| 153 | XCR1 | Xcr1 | 14584 | -1.073 | -0.2844 | Yes | ||
| 154 | NGFR | Ngfr | 14612 | -1.094 | -0.2788 | Yes | ||
| 155 | LIF | Lif | 14618 | -1.096 | -0.2718 | Yes | ||
| 156 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.2769 | Yes | ||
| 157 | IL18 | Il18 | 14836 | -1.294 | -0.2686 | Yes | ||
| 158 | TNFRSF9 | Tnfrsf9 | 14864 | -1.331 | -0.2614 | Yes | ||
| 159 | CX3CL1 | Cx3cl1 | 14878 | -1.343 | -0.2533 | Yes | ||
| 160 | LIFR | Lifr | 14879 | -1.343 | -0.2443 | Yes | ||
| 161 | EGF | Egf | 14924 | -1.396 | -0.2378 | Yes | ||
| 162 | CXCL11 | Cxcl11 | 14938 | -1.408 | -0.2292 | Yes | ||
| 163 | IL1R1 | Il1r1 | 14956 | -1.437 | -0.2207 | Yes | ||
| 164 | IL11 | Il11 | 14960 | -1.447 | -0.2112 | Yes | ||
| 165 | CCR1 | Ccr1 | 15042 | -1.555 | -0.2061 | Yes | ||
| 166 | IL1RAP | Il1rap | 15074 | -1.621 | -0.1973 | Yes | ||
| 167 | IL23A | Il23a | 15082 | -1.635 | -0.1868 | Yes | ||
| 168 | BMP2 | Bmp2 | 15094 | -1.657 | -0.1764 | Yes | ||
| 169 | CSF2 | Csf2 | 15123 | -1.702 | -0.1668 | Yes | ||
| 170 | CCL7 | Ccl7 | 15132 | -1.719 | -0.1558 | Yes | ||
| 171 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.1458 | Yes | ||
| 172 | CCL2 | Ccl2 | 15268 | -2.037 | -0.1389 | Yes | ||
| 173 | CCL5 | Ccl5 | 15278 | -2.073 | -0.1256 | Yes | ||
| 174 | CCL8 | Ccl12 | 15326 | -2.242 | -0.1137 | Yes | ||
| 175 | INHBA | Inhba | 15335 | -2.275 | -0.0990 | Yes | ||
| 176 | CCL17 | Ccl17 | 15346 | -2.308 | -0.0842 | Yes | ||
| 177 | IL13RA1 | Il13ra1 | 15351 | -2.324 | -0.0689 | Yes | ||
| 178 | CXCL10 | Cxcl10 | 15390 | -2.511 | -0.0545 | Yes | ||
| 179 | CCL11 | Ccl11 | 15445 | -2.856 | -0.0389 | Yes | ||
| 180 | HGF | Hgf | 15472 | -3.068 | -0.0201 | Yes | ||
| 181 | CXCL14 | Cxcl14 | 15502 | -3.500 | 0.0015 | Yes |