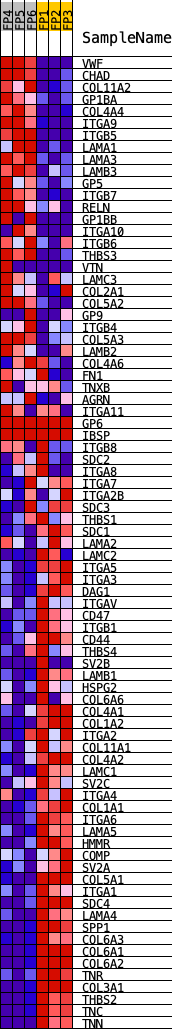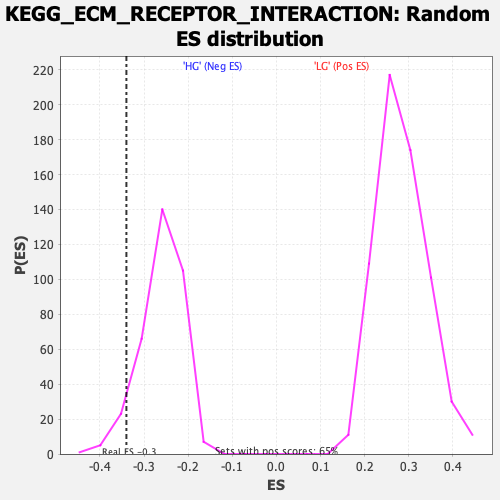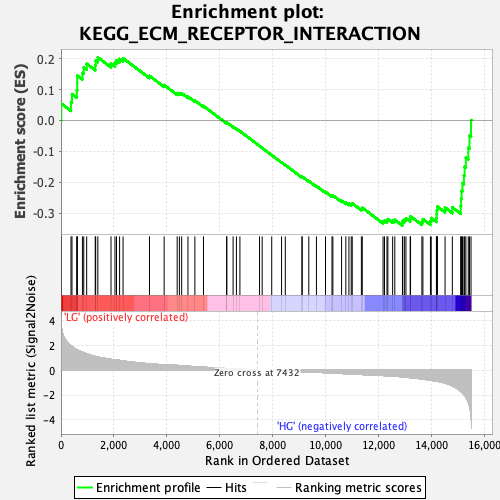
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_ECM_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | -0.33999917 |
| Normalized Enrichment Score (NES) | -1.3052408 |
| Nominal p-value | 0.06051873 |
| FDR q-value | 0.35848427 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VWF | Vwf | 11 | 3.767 | 0.0545 | No | ||
| 2 | CHAD | Chad | 378 | 1.964 | 0.0596 | No | ||
| 3 | COL11A2 | Col11a2 | 415 | 1.909 | 0.0853 | No | ||
| 4 | GP1BA | Gp1ba | 597 | 1.657 | 0.0978 | No | ||
| 5 | COL4A4 | Col4a4 | 612 | 1.644 | 0.1210 | No | ||
| 6 | ITGA9 | Itga9 | 614 | 1.642 | 0.1450 | No | ||
| 7 | ITGB5 | Itgb5 | 810 | 1.446 | 0.1536 | No | ||
| 8 | LAMA1 | Lama1 | 860 | 1.405 | 0.1710 | No | ||
| 9 | LAMA3 | Lama3 | 971 | 1.318 | 0.1832 | No | ||
| 10 | LAMB3 | Lamb3 | 1294 | 1.102 | 0.1785 | No | ||
| 11 | GP5 | Gp5 | 1311 | 1.093 | 0.1935 | No | ||
| 12 | ITGB7 | Itgb7 | 1393 | 1.056 | 0.2037 | No | ||
| 13 | RELN | Reln | 1891 | 0.866 | 0.1842 | No | ||
| 14 | GP1BB | Gp1bb | 2041 | 0.825 | 0.1867 | No | ||
| 15 | ITGA10 | Gm42957 | 2099 | 0.810 | 0.1949 | No | ||
| 16 | ITGB6 | Itgb6 | 2216 | 0.782 | 0.1988 | No | ||
| 17 | THBS3 | Thbs3 | 2347 | 0.742 | 0.2013 | No | ||
| 18 | VTN | Vtn | 3346 | 0.506 | 0.1441 | No | ||
| 19 | LAMC3 | Lamc3 | 3901 | 0.428 | 0.1145 | No | ||
| 20 | COL2A1 | Col2a1 | 4394 | 0.380 | 0.0882 | No | ||
| 21 | COL5A2 | Col5a2 | 4478 | 0.365 | 0.0882 | No | ||
| 22 | GP9 | Gp9 | 4564 | 0.350 | 0.0878 | No | ||
| 23 | ITGB4 | Itgb4 | 4802 | 0.312 | 0.0771 | No | ||
| 24 | COL5A3 | Col5a3 | 5063 | 0.278 | 0.0643 | No | ||
| 25 | LAMB2 | Lamb2 | 5389 | 0.236 | 0.0467 | No | ||
| 26 | COL4A6 | Col4a6 | 6263 | 0.129 | -0.0079 | No | ||
| 27 | FN1 | Fn1 | 6272 | 0.127 | -0.0066 | No | ||
| 28 | TNXB | Tnxb | 6509 | 0.099 | -0.0204 | No | ||
| 29 | AGRN | Agrn | 6635 | 0.085 | -0.0272 | No | ||
| 30 | ITGA11 | Itga11 | 6767 | 0.070 | -0.0347 | No | ||
| 31 | GP6 | Gp6 | 7509 | 0.000 | -0.0827 | No | ||
| 32 | IBSP | Ibsp | 7609 | 0.000 | -0.0891 | No | ||
| 33 | ITGB8 | Itgb8 | 7970 | -0.006 | -0.1123 | No | ||
| 34 | SDC2 | Sdc2 | 8342 | -0.040 | -0.1357 | No | ||
| 35 | ITGA8 | Itga8 | 8483 | -0.052 | -0.1441 | No | ||
| 36 | ITGA7 | Itga7 | 9113 | -0.117 | -0.1831 | No | ||
| 37 | ITGA2B | Itga2b | 9122 | -0.117 | -0.1819 | No | ||
| 38 | SDC3 | Sdc3 | 9372 | -0.141 | -0.1959 | No | ||
| 39 | THBS1 | Thbs1 | 9661 | -0.170 | -0.2121 | No | ||
| 40 | SDC1 | Sdc1 | 10005 | -0.204 | -0.2313 | No | ||
| 41 | LAMA2 | Lama2 | 10249 | -0.228 | -0.2437 | No | ||
| 42 | LAMC2 | Lamc2 | 10283 | -0.232 | -0.2424 | No | ||
| 43 | ITGA5 | Itga5 | 10612 | -0.266 | -0.2598 | No | ||
| 44 | ITGA3 | Itga3 | 10773 | -0.284 | -0.2660 | No | ||
| 45 | DAG1 | Dag1 | 10890 | -0.295 | -0.2691 | No | ||
| 46 | ITGAV | Itgav | 10977 | -0.305 | -0.2702 | No | ||
| 47 | CD47 | Cd47 | 11015 | -0.309 | -0.2681 | No | ||
| 48 | ITGB1 | Itgb1 | 11361 | -0.348 | -0.2853 | No | ||
| 49 | CD44 | Cd44 | 11395 | -0.352 | -0.2823 | No | ||
| 50 | THBS4 | Thbs4 | 12181 | -0.432 | -0.3268 | No | ||
| 51 | SV2B | Sv2b | 12235 | -0.440 | -0.3238 | No | ||
| 52 | LAMB1 | Lamb1 | 12329 | -0.446 | -0.3233 | No | ||
| 53 | HSPG2 | Hspg2 | 12363 | -0.451 | -0.3188 | No | ||
| 54 | COL6A6 | Col6a6 | 12543 | -0.483 | -0.3233 | No | ||
| 55 | COL4A1 | Col4a1 | 12625 | -0.491 | -0.3214 | No | ||
| 56 | COL1A2 | Col1a2 | 12914 | -0.534 | -0.3322 | Yes | ||
| 57 | ITGA2 | Itga2 | 12923 | -0.536 | -0.3248 | Yes | ||
| 58 | COL11A1 | Col11a1 | 12986 | -0.548 | -0.3208 | Yes | ||
| 59 | COL4A2 | Col4a2 | 13046 | -0.562 | -0.3164 | Yes | ||
| 60 | LAMC1 | Lamc1 | 13207 | -0.600 | -0.3180 | Yes | ||
| 61 | SV2C | Sv2c | 13220 | -0.602 | -0.3099 | Yes | ||
| 62 | ITGA4 | Itga4 | 13649 | -0.711 | -0.3272 | Yes | ||
| 63 | COL1A1 | Col1a1 | 13688 | -0.723 | -0.3191 | Yes | ||
| 64 | ITGA6 | Itga6 | 13973 | -0.809 | -0.3256 | Yes | ||
| 65 | LAMA5 | Lama5 | 14003 | -0.818 | -0.3155 | Yes | ||
| 66 | HMMR | Hmmr | 14199 | -0.883 | -0.3152 | Yes | ||
| 67 | COMP | Comp | 14208 | -0.884 | -0.3027 | Yes | ||
| 68 | SV2A | Sv2a | 14214 | -0.887 | -0.2900 | Yes | ||
| 69 | COL5A1 | Col5a1 | 14238 | -0.898 | -0.2784 | Yes | ||
| 70 | ITGA1 | Itga1 | 14526 | -1.039 | -0.2817 | Yes | ||
| 71 | SDC4 | Sdc4 | 14801 | -1.261 | -0.2810 | Yes | ||
| 72 | LAMA4 | Lama4 | 15118 | -1.696 | -0.2766 | Yes | ||
| 73 | SPP1 | Spp1 | 15131 | -1.718 | -0.2522 | Yes | ||
| 74 | COL6A3 | Col6a3 | 15151 | -1.768 | -0.2275 | Yes | ||
| 75 | COL6A1 | Col6a1 | 15185 | -1.824 | -0.2029 | Yes | ||
| 76 | COL6A2 | Col6a2 | 15240 | -1.962 | -0.1776 | Yes | ||
| 77 | TNR | Tnr | 15265 | -2.028 | -0.1494 | Yes | ||
| 78 | COL3A1 | Col3a1 | 15313 | -2.188 | -0.1204 | Yes | ||
| 79 | THBS2 | Thbs2 | 15404 | -2.585 | -0.0884 | Yes | ||
| 80 | TNC | Tnc | 15451 | -2.893 | -0.0489 | Yes | ||
| 81 | TNN | Tnn | 15508 | -3.660 | 0.0011 | Yes |