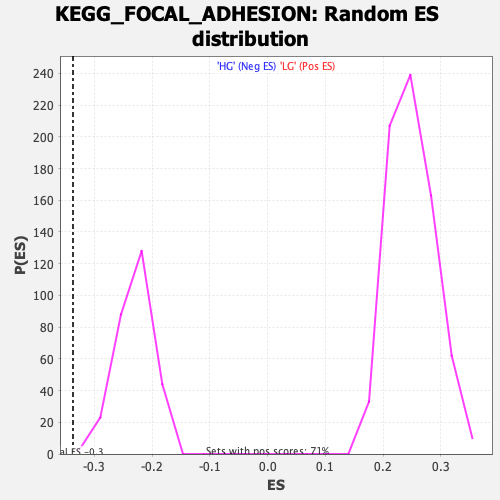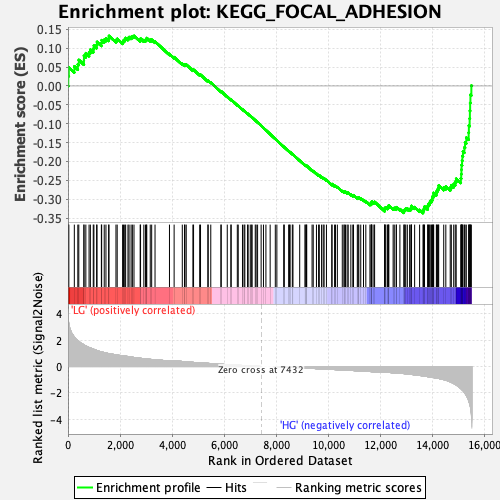
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_FOCAL_ADHESION |
| Enrichment Score (ES) | -0.3368867 |
| Normalized Enrichment Score (NES) | -1.4577802 |
| Nominal p-value | 0.0034965035 |
| FDR q-value | 0.25269252 |
| FWER p-Value | 0.897 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VWF | Vwf | 11 | 3.767 | 0.0267 | No | ||
| 2 | KDR | Kdr | 32 | 3.271 | 0.0492 | No | ||
| 3 | FLT4 | Flt4 | 243 | 2.244 | 0.0518 | No | ||
| 4 | CHAD | Chad | 378 | 1.964 | 0.0574 | No | ||
| 5 | COL11A2 | Col11a2 | 415 | 1.909 | 0.0690 | No | ||
| 6 | COL4A4 | Col4a4 | 612 | 1.644 | 0.0681 | No | ||
| 7 | ITGA9 | Itga9 | 614 | 1.642 | 0.0800 | No | ||
| 8 | PARVG | Parvg | 686 | 1.562 | 0.0868 | No | ||
| 9 | ITGB5 | Itgb5 | 810 | 1.446 | 0.0893 | No | ||
| 10 | LAMA1 | Lama1 | 860 | 1.405 | 0.0963 | No | ||
| 11 | LAMA3 | Lama3 | 971 | 1.318 | 0.0987 | No | ||
| 12 | PRKCG | Prkcg | 996 | 1.303 | 0.1066 | No | ||
| 13 | VEGFB | Vegfb | 1101 | 1.223 | 0.1088 | No | ||
| 14 | VEGFC | Vegfc | 1115 | 1.215 | 0.1168 | No | ||
| 15 | PAK6 | Pak6 | 1283 | 1.107 | 0.1139 | No | ||
| 16 | LAMB3 | Lamb3 | 1294 | 1.102 | 0.1213 | No | ||
| 17 | ITGB7 | Itgb7 | 1393 | 1.056 | 0.1226 | No | ||
| 18 | MYLK2 | Mylk2 | 1459 | 1.027 | 0.1258 | No | ||
| 19 | MYL10 | Myl10 | 1563 | 0.980 | 0.1262 | No | ||
| 20 | FYN | Fyn | 1575 | 0.976 | 0.1326 | No | ||
| 21 | PAK3 | Pak3 | 1844 | 0.880 | 0.1216 | No | ||
| 22 | RELN | Reln | 1891 | 0.866 | 0.1249 | No | ||
| 23 | ITGA10 | Gm42957 | 2099 | 0.810 | 0.1173 | No | ||
| 24 | PDGFB | Pdgfb | 2135 | 0.802 | 0.1208 | No | ||
| 25 | HRAS | Hras | 2180 | 0.791 | 0.1237 | No | ||
| 26 | ITGB6 | Itgb6 | 2216 | 0.782 | 0.1271 | No | ||
| 27 | RAC3 | Rac3 | 2306 | 0.753 | 0.1268 | No | ||
| 28 | THBS3 | Thbs3 | 2347 | 0.742 | 0.1296 | No | ||
| 29 | MYL9 | Myl9 | 2425 | 0.716 | 0.1298 | No | ||
| 30 | ERBB2 | Erbb2 | 2475 | 0.702 | 0.1317 | No | ||
| 31 | BCL2 | Bcl2 | 2537 | 0.682 | 0.1327 | No | ||
| 32 | VAV2 | Vav2 | 2775 | 0.625 | 0.1218 | No | ||
| 33 | CCND3 | Ccnd3 | 2789 | 0.623 | 0.1255 | No | ||
| 34 | MYL12B | Myl12b | 2916 | 0.593 | 0.1216 | No | ||
| 35 | SHC1 | Shc1 | 2978 | 0.579 | 0.1218 | No | ||
| 36 | ELK1 | Elk1 | 3014 | 0.569 | 0.1237 | No | ||
| 37 | PDGFA | Pdgfa | 3028 | 0.566 | 0.1269 | No | ||
| 38 | PIK3CG | Pik3cg | 3162 | 0.541 | 0.1222 | No | ||
| 39 | BAD | Bad | 3217 | 0.532 | 0.1225 | No | ||
| 40 | VTN | Vtn | 3346 | 0.506 | 0.1179 | No | ||
| 41 | LAMC3 | Lamc3 | 3901 | 0.428 | 0.0849 | No | ||
| 42 | PIK3R5 | Pik3r5 | 4080 | 0.414 | 0.0763 | No | ||
| 43 | COL2A1 | Col2a1 | 4394 | 0.380 | 0.0586 | No | ||
| 44 | COL5A2 | Col5a2 | 4478 | 0.365 | 0.0559 | No | ||
| 45 | PTEN | Pten | 4496 | 0.362 | 0.0574 | No | ||
| 46 | MYLK3 | Mylk3 | 4545 | 0.354 | 0.0568 | No | ||
| 47 | ITGB4 | Itgb4 | 4802 | 0.312 | 0.0424 | No | ||
| 48 | RAC2 | Rac2 | 4827 | 0.309 | 0.0431 | No | ||
| 49 | COL5A3 | Col5a3 | 5063 | 0.278 | 0.0298 | No | ||
| 50 | PIK3CD | Pik3cd | 5095 | 0.274 | 0.0298 | No | ||
| 51 | CAPN2 | Capn2 | 5378 | 0.237 | 0.0131 | No | ||
| 52 | LAMB2 | Lamb2 | 5389 | 0.236 | 0.0142 | No | ||
| 53 | MAPK9 | Mapk9 | 5489 | 0.221 | 0.0093 | No | ||
| 54 | GSK3B | Gsk3b | 5879 | 0.171 | -0.0148 | No | ||
| 55 | FLNC | Flnc | 5890 | 0.169 | -0.0142 | No | ||
| 56 | PARVB | Parvb | 6123 | 0.143 | -0.0283 | No | ||
| 57 | COL4A6 | Col4a6 | 6263 | 0.129 | -0.0364 | No | ||
| 58 | FN1 | Fn1 | 6272 | 0.127 | -0.0360 | No | ||
| 59 | TNXB | Tnxb | 6509 | 0.099 | -0.0507 | No | ||
| 60 | CTNNB1 | Ctnnb1 | 6545 | 0.096 | -0.0523 | No | ||
| 61 | MYL12A | Myl12a | 6708 | 0.077 | -0.0623 | No | ||
| 62 | ITGA11 | Itga11 | 6767 | 0.070 | -0.0656 | No | ||
| 63 | VASP | Vasp | 6798 | 0.066 | -0.0670 | No | ||
| 64 | RHOA | Rhoa | 6904 | 0.056 | -0.0735 | No | ||
| 65 | CRK | Crk | 6925 | 0.054 | -0.0744 | No | ||
| 66 | PIK3CB | Pik3cb | 7007 | 0.045 | -0.0793 | No | ||
| 67 | AKT2 | Akt2 | 7049 | 0.039 | -0.0817 | No | ||
| 68 | RAP1B | Rap1b | 7080 | 0.036 | -0.0834 | No | ||
| 69 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.0909 | No | ||
| 70 | DIAPH1 | Diaph1 | 7233 | 0.020 | -0.0930 | No | ||
| 71 | FLNA | Flna | 7289 | 0.015 | -0.0964 | No | ||
| 72 | BCAR1 | Bcar1 | 7425 | 0.001 | -0.1052 | No | ||
| 73 | VAV1 | Vav1 | 7515 | 0.000 | -0.1110 | No | ||
| 74 | IBSP | Ibsp | 7609 | 0.000 | -0.1171 | No | ||
| 75 | PAK5 | Pak7 | 7772 | 0.000 | -0.1277 | No | ||
| 76 | ITGB8 | Itgb8 | 7970 | -0.006 | -0.1405 | No | ||
| 77 | MAPK10 | Mapk10 | 8030 | -0.011 | -0.1442 | No | ||
| 78 | AKT1 | Akt1 | 8293 | -0.035 | -0.1611 | No | ||
| 79 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.1622 | No | ||
| 80 | ITGA8 | Itga8 | 8483 | -0.052 | -0.1727 | No | ||
| 81 | GRB2 | Grb2 | 8518 | -0.055 | -0.1745 | No | ||
| 82 | RAF1 | Raf1 | 8539 | -0.058 | -0.1754 | No | ||
| 83 | PIP5K1C | Pip5k1c | 8624 | -0.067 | -0.1804 | No | ||
| 84 | PDPK1 | Pdpk1 | 8649 | -0.070 | -0.1814 | No | ||
| 85 | ACTN1 | Actn1 | 8908 | -0.092 | -0.1976 | No | ||
| 86 | ITGA7 | Itga7 | 9113 | -0.117 | -0.2100 | No | ||
| 87 | ITGA2B | Itga2b | 9122 | -0.117 | -0.2097 | No | ||
| 88 | SOS2 | Sos2 | 9170 | -0.122 | -0.2119 | No | ||
| 89 | BRAF | Braf | 9178 | -0.123 | -0.2114 | No | ||
| 90 | CDC42 | Cdc42 | 9384 | -0.142 | -0.2238 | No | ||
| 91 | SOS1 | Sos1 | 9428 | -0.146 | -0.2255 | No | ||
| 92 | MAPK8 | Mapk8 | 9552 | -0.159 | -0.2324 | No | ||
| 93 | ARHGAP35 | Arhgap35 | 9635 | -0.167 | -0.2365 | No | ||
| 94 | THBS1 | Thbs1 | 9661 | -0.170 | -0.2369 | No | ||
| 95 | PAK2 | Pak2 | 9749 | -0.178 | -0.2413 | No | ||
| 96 | ROCK1 | Rock1 | 9810 | -0.184 | -0.2438 | No | ||
| 97 | PPP1CA | Ppp1ca | 9846 | -0.188 | -0.2448 | No | ||
| 98 | PAK4 | Pak4 | 9936 | -0.197 | -0.2491 | No | ||
| 99 | CAV2 | Cav2 | 10143 | -0.218 | -0.2610 | No | ||
| 100 | CRKL | Crkl | 10150 | -0.218 | -0.2598 | No | ||
| 101 | LAMA2 | Lama2 | 10249 | -0.228 | -0.2645 | No | ||
| 102 | LAMC2 | Lamc2 | 10283 | -0.232 | -0.2650 | No | ||
| 103 | TLN2 | Tln2 | 10355 | -0.240 | -0.2679 | No | ||
| 104 | VCL | Vcl | 10545 | -0.259 | -0.2783 | No | ||
| 105 | ITGA5 | Itga5 | 10612 | -0.266 | -0.2807 | No | ||
| 106 | RAP1A | Rap1a | 10620 | -0.267 | -0.2792 | No | ||
| 107 | ACTG1 | Actg1 | 10670 | -0.272 | -0.2804 | No | ||
| 108 | PPP1CB | Ppp1cb | 10731 | -0.279 | -0.2823 | No | ||
| 109 | ITGA3 | Itga3 | 10773 | -0.284 | -0.2829 | No | ||
| 110 | ACTN2 | Actn2 | 10874 | -0.293 | -0.2873 | No | ||
| 111 | CCND1 | Ccnd1 | 10950 | -0.301 | -0.2900 | No | ||
| 112 | ITGAV | Itgav | 10977 | -0.305 | -0.2894 | No | ||
| 113 | ROCK2 | Rock2 | 11123 | -0.321 | -0.2966 | No | ||
| 114 | VEGFD | Vegfd | 11156 | -0.326 | -0.2963 | No | ||
| 115 | CAV1 | Cav1 | 11181 | -0.328 | -0.2955 | No | ||
| 116 | PDGFC | Pdgfc | 11255 | -0.336 | -0.2978 | No | ||
| 117 | ITGB1 | Itgb1 | 11361 | -0.348 | -0.3021 | No | ||
| 118 | RAC1 | Rac1 | 11450 | -0.360 | -0.3052 | No | ||
| 119 | PXN | Pxn | 11609 | -0.377 | -0.3128 | No | ||
| 120 | ACTB | Actb | 11649 | -0.382 | -0.3125 | No | ||
| 121 | RAPGEF1 | Rapgef1 | 11670 | -0.385 | -0.3110 | No | ||
| 122 | FLNB | Flnb | 11676 | -0.386 | -0.3085 | No | ||
| 123 | PPP1R12A | Ppp1r12a | 11682 | -0.388 | -0.3060 | No | ||
| 124 | ACTN4 | Actn4 | 11761 | -0.397 | -0.3082 | No | ||
| 125 | IGF1R | Igf1r | 11788 | -0.400 | -0.3070 | No | ||
| 126 | PPP1CC | Ppp1cc | 12176 | -0.431 | -0.3291 | No | ||
| 127 | PTK2 | Ptk2 | 12178 | -0.432 | -0.3260 | No | ||
| 128 | THBS4 | Thbs4 | 12181 | -0.432 | -0.3230 | No | ||
| 129 | PDGFD | Pdgfd | 12217 | -0.437 | -0.3221 | No | ||
| 130 | MYL2 | Myl2 | 12280 | -0.440 | -0.3230 | No | ||
| 131 | TLN1 | Tln1 | 12307 | -0.442 | -0.3214 | No | ||
| 132 | LAMB1 | Lamb1 | 12329 | -0.446 | -0.3196 | No | ||
| 133 | SRC | Src | 12330 | -0.446 | -0.3163 | No | ||
| 134 | MYLK | Mylk | 12500 | -0.474 | -0.3239 | No | ||
| 135 | COL6A6 | Col6a6 | 12543 | -0.483 | -0.3231 | No | ||
| 136 | AKT3 | Akt3 | 12618 | -0.490 | -0.3244 | No | ||
| 137 | COL4A1 | Col4a1 | 12625 | -0.491 | -0.3212 | No | ||
| 138 | PIK3R1 | Pik3r1 | 12760 | -0.509 | -0.3262 | No | ||
| 139 | COL1A2 | Col1a2 | 12914 | -0.534 | -0.3323 | Yes | ||
| 140 | ITGA2 | Itga2 | 12923 | -0.536 | -0.3289 | Yes | ||
| 141 | VEGFA | Vegfa | 12953 | -0.540 | -0.3269 | Yes | ||
| 142 | COL11A1 | Col11a1 | 12986 | -0.548 | -0.3250 | Yes | ||
| 143 | COL4A2 | Col4a2 | 13046 | -0.562 | -0.3247 | Yes | ||
| 144 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.3262 | Yes | ||
| 145 | XIAP | Xiap | 13180 | -0.594 | -0.3248 | Yes | ||
| 146 | PAK1 | Pak1 | 13205 | -0.599 | -0.3220 | Yes | ||
| 147 | LAMC1 | Lamc1 | 13207 | -0.600 | -0.3177 | Yes | ||
| 148 | PIK3CA | Pik3ca | 13327 | -0.625 | -0.3209 | Yes | ||
| 149 | JUN | Jun | 13518 | -0.673 | -0.3284 | Yes | ||
| 150 | ITGA4 | Itga4 | 13649 | -0.711 | -0.3317 | Yes | ||
| 151 | BIRC3 | Birc3 | 13670 | -0.718 | -0.3278 | Yes | ||
| 152 | COL1A1 | Col1a1 | 13688 | -0.723 | -0.3236 | Yes | ||
| 153 | DOCK1 | Dock1 | 13707 | -0.729 | -0.3195 | Yes | ||
| 154 | PGF | Pgf | 13830 | -0.765 | -0.3219 | Yes | ||
| 155 | ARHGAP5 | Arhgap5 | 13836 | -0.767 | -0.3166 | Yes | ||
| 156 | MET | Met | 13861 | -0.774 | -0.3126 | Yes | ||
| 157 | PARVA | Parva | 13906 | -0.791 | -0.3097 | Yes | ||
| 158 | MYL7 | Myl7 | 13922 | -0.793 | -0.3049 | Yes | ||
| 159 | ITGA6 | Itga6 | 13973 | -0.809 | -0.3023 | Yes | ||
| 160 | LAMA5 | Lama5 | 14003 | -0.818 | -0.2982 | Yes | ||
| 161 | SHC3 | Shc3 | 14013 | -0.825 | -0.2928 | Yes | ||
| 162 | FLT1 | Flt1 | 14053 | -0.837 | -0.2892 | Yes | ||
| 163 | MYLPF | Mylpf | 14058 | -0.839 | -0.2834 | Yes | ||
| 164 | BIRC2 | Birc2 | 14162 | -0.865 | -0.2838 | Yes | ||
| 165 | EGFR | Egfr | 14176 | -0.870 | -0.2783 | Yes | ||
| 166 | COMP | Comp | 14208 | -0.884 | -0.2739 | Yes | ||
| 167 | COL5A1 | Col5a1 | 14238 | -0.898 | -0.2693 | Yes | ||
| 168 | ILK | Ilk | 14252 | -0.902 | -0.2635 | Yes | ||
| 169 | ZYX | Zyx | 14445 | -0.991 | -0.2688 | Yes | ||
| 170 | ITGA1 | Itga1 | 14526 | -1.039 | -0.2665 | Yes | ||
| 171 | SHC4 | Shc4 | 14695 | -1.160 | -0.2690 | Yes | ||
| 172 | PRKCA | Prkca | 14737 | -1.199 | -0.2630 | Yes | ||
| 173 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.2595 | Yes | ||
| 174 | VAV3 | Vav3 | 14888 | -1.356 | -0.2534 | Yes | ||
| 175 | EGF | Egf | 14924 | -1.396 | -0.2455 | Yes | ||
| 176 | CAV3 | Cav3 | 15102 | -1.666 | -0.2450 | Yes | ||
| 177 | LAMA4 | Lama4 | 15118 | -1.696 | -0.2336 | Yes | ||
| 178 | ACTN3 | Actn3 | 15126 | -1.708 | -0.2216 | Yes | ||
| 179 | SPP1 | Spp1 | 15131 | -1.718 | -0.2094 | Yes | ||
| 180 | COL6A3 | Col6a3 | 15151 | -1.768 | -0.1978 | Yes | ||
| 181 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.1855 | Yes | ||
| 182 | COL6A1 | Col6a1 | 15185 | -1.824 | -0.1735 | Yes | ||
| 183 | COL6A2 | Col6a2 | 15240 | -1.962 | -0.1628 | Yes | ||
| 184 | TNR | Tnr | 15265 | -2.028 | -0.1496 | Yes | ||
| 185 | COL3A1 | Col3a1 | 15313 | -2.188 | -0.1367 | Yes | ||
| 186 | THBS2 | Thbs2 | 15404 | -2.585 | -0.1238 | Yes | ||
| 187 | IGF1 | Igf1 | 15412 | -2.616 | -0.1052 | Yes | ||
| 188 | RASGRF1 | Rasgrf1 | 15442 | -2.844 | -0.0864 | Yes | ||
| 189 | TNC | Tnc | 15451 | -2.893 | -0.0658 | Yes | ||
| 190 | PRKCB | Prkcb | 15461 | -2.954 | -0.0449 | Yes | ||
| 191 | HGF | Hgf | 15472 | -3.068 | -0.0233 | Yes | ||
| 192 | TNN | Tnn | 15508 | -3.660 | 0.0011 | Yes |