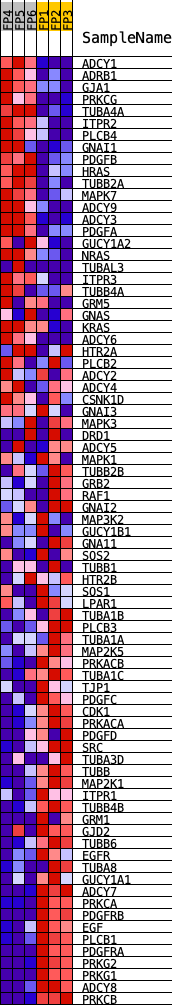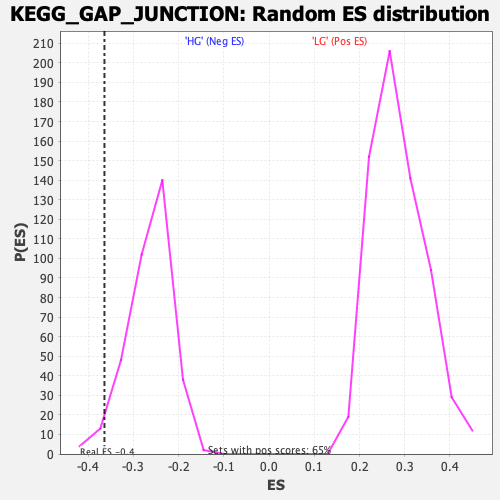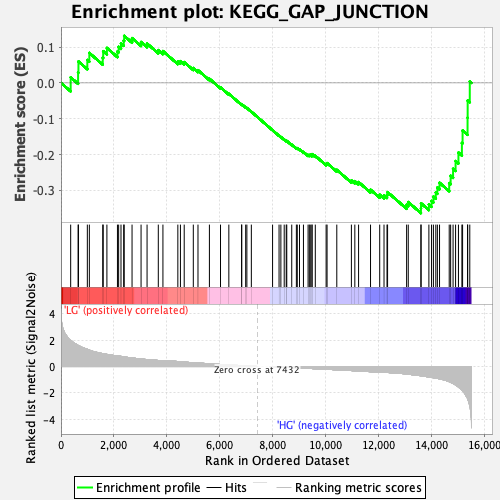
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_GAP_JUNCTION |
| Enrichment Score (ES) | -0.36386806 |
| Normalized Enrichment Score (NES) | -1.3827095 |
| Nominal p-value | 0.0259366 |
| FDR q-value | 0.27989718 |
| FWER p-Value | 0.981 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADCY1 | Adcy1 | 366 | 1.982 | 0.0156 | No | ||
| 2 | ADRB1 | Adrb1 | 647 | 1.600 | 0.0291 | No | ||
| 3 | GJA1 | Gja6 | 660 | 1.594 | 0.0599 | No | ||
| 4 | PRKCG | Prkcg | 996 | 1.303 | 0.0640 | No | ||
| 5 | TUBA4A | Tuba4a | 1072 | 1.243 | 0.0838 | No | ||
| 6 | ITPR2 | Itpr2 | 1579 | 0.974 | 0.0704 | No | ||
| 7 | PLCB4 | Plcb4 | 1596 | 0.969 | 0.0885 | No | ||
| 8 | GNAI1 | Gnai1 | 1736 | 0.921 | 0.0978 | No | ||
| 9 | PDGFB | Pdgfb | 2135 | 0.802 | 0.0879 | No | ||
| 10 | HRAS | Hras | 2180 | 0.791 | 0.1007 | No | ||
| 11 | TUBB2A | Tubb2a | 2269 | 0.766 | 0.1102 | No | ||
| 12 | MAPK7 | Mapk7 | 2371 | 0.734 | 0.1182 | No | ||
| 13 | ADCY9 | Adcy9 | 2390 | 0.729 | 0.1314 | No | ||
| 14 | ADCY3 | Adcy3 | 2687 | 0.646 | 0.1251 | No | ||
| 15 | PDGFA | Pdgfa | 3028 | 0.566 | 0.1143 | No | ||
| 16 | GUCY1A2 | Gucy1a2 | 3255 | 0.524 | 0.1100 | No | ||
| 17 | NRAS | Nras | 3680 | 0.455 | 0.0916 | No | ||
| 18 | TUBAL3 | Tubal3 | 3855 | 0.432 | 0.0889 | No | ||
| 19 | ITPR3 | Itpr3 | 4419 | 0.376 | 0.0599 | No | ||
| 20 | TUBB4A | Tubb4a | 4517 | 0.359 | 0.0607 | No | ||
| 21 | GRM5 | Grm5 | 4660 | 0.333 | 0.0581 | No | ||
| 22 | GNAS | Gnas | 5004 | 0.286 | 0.0416 | No | ||
| 23 | KRAS | Kras | 5181 | 0.266 | 0.0355 | No | ||
| 24 | ADCY6 | Adcy6 | 5613 | 0.207 | 0.0117 | No | ||
| 25 | HTR2A | Htr2a | 6033 | 0.152 | -0.0125 | No | ||
| 26 | PLCB2 | Plcb2 | 6348 | 0.118 | -0.0304 | No | ||
| 27 | ADCY2 | Adcy2 | 6832 | 0.063 | -0.0605 | No | ||
| 28 | ADCY4 | Adcy4 | 6834 | 0.063 | -0.0593 | No | ||
| 29 | CSNK1D | Csnk1d | 6985 | 0.048 | -0.0680 | No | ||
| 30 | GNAI3 | Gnai3 | 7030 | 0.042 | -0.0700 | No | ||
| 31 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.0805 | No | ||
| 32 | DRD1 | Drd1 | 8002 | -0.009 | -0.1322 | No | ||
| 33 | ADCY5 | Adcy5 | 8247 | -0.030 | -0.1474 | No | ||
| 34 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.1511 | No | ||
| 35 | TUBB2B | Tubb2b | 8442 | -0.048 | -0.1582 | No | ||
| 36 | GRB2 | Grb2 | 8518 | -0.055 | -0.1620 | No | ||
| 37 | RAF1 | Raf1 | 8539 | -0.058 | -0.1621 | No | ||
| 38 | GNAI2 | Gnai2 | 8727 | -0.077 | -0.1727 | No | ||
| 39 | MAP3K2 | Map3k2 | 8902 | -0.092 | -0.1821 | No | ||
| 40 | GUCY1B1 | Gucy1b1 | 8936 | -0.096 | -0.1824 | No | ||
| 41 | GNA11 | Gna11 | 9020 | -0.106 | -0.1856 | No | ||
| 42 | SOS2 | Sos2 | 9170 | -0.122 | -0.1929 | No | ||
| 43 | TUBB1 | Tubb1 | 9345 | -0.138 | -0.2014 | No | ||
| 44 | HTR2B | Htr2b | 9391 | -0.143 | -0.2015 | No | ||
| 45 | SOS1 | Sos1 | 9428 | -0.146 | -0.2009 | No | ||
| 46 | LPAR1 | Lpar1 | 9492 | -0.152 | -0.2020 | No | ||
| 47 | TUBA1B | Tuba1b | 9495 | -0.153 | -0.1991 | No | ||
| 48 | PLCB3 | Plcb3 | 9618 | -0.166 | -0.2037 | No | ||
| 49 | TUBA1A | Tuba1a | 10027 | -0.206 | -0.2260 | No | ||
| 50 | MAP2K5 | Map2k5 | 10063 | -0.211 | -0.2241 | No | ||
| 51 | PRKACB | Prkacb | 10431 | -0.249 | -0.2430 | No | ||
| 52 | TUBA1C | Tuba1c | 10983 | -0.305 | -0.2726 | No | ||
| 53 | TJP1 | Tjp1 | 11113 | -0.321 | -0.2746 | No | ||
| 54 | PDGFC | Pdgfc | 11255 | -0.336 | -0.2770 | No | ||
| 55 | CDK1 | Cdk1 | 11707 | -0.390 | -0.2985 | No | ||
| 56 | PRKACA | Prkaca | 12054 | -0.424 | -0.3125 | No | ||
| 57 | PDGFD | Pdgfd | 12217 | -0.437 | -0.3143 | No | ||
| 58 | SRC | Src | 12330 | -0.446 | -0.3128 | No | ||
| 59 | TUBA3D | Tuba3b | 12352 | -0.449 | -0.3052 | No | ||
| 60 | TUBB | Tubb5 | 13072 | -0.569 | -0.3405 | No | ||
| 61 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.3329 | No | ||
| 62 | ITPR1 | Itpr1 | 13613 | -0.701 | -0.3500 | Yes | ||
| 63 | TUBB4B | Tubb4b | 13618 | -0.702 | -0.3363 | Yes | ||
| 64 | GRM1 | Grm1 | 13914 | -0.793 | -0.3397 | Yes | ||
| 65 | GJD2 | Gjd2 | 14015 | -0.825 | -0.3299 | Yes | ||
| 66 | TUBB6 | Tubb6 | 14085 | -0.844 | -0.3176 | Yes | ||
| 67 | EGFR | Egfr | 14176 | -0.870 | -0.3062 | Yes | ||
| 68 | TUBA8 | Tuba8 | 14235 | -0.897 | -0.2922 | Yes | ||
| 69 | GUCY1A1 | Gucy1a1 | 14317 | -0.932 | -0.2790 | Yes | ||
| 70 | ADCY7 | Adcy7 | 14683 | -1.147 | -0.2799 | Yes | ||
| 71 | PRKCA | Prkca | 14737 | -1.199 | -0.2596 | Yes | ||
| 72 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.2399 | Yes | ||
| 73 | EGF | Egf | 14924 | -1.396 | -0.2184 | Yes | ||
| 74 | PLCB1 | Plcb1 | 15032 | -1.546 | -0.1947 | Yes | ||
| 75 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.1677 | Yes | ||
| 76 | PRKG2 | Prkg2 | 15189 | -1.835 | -0.1329 | Yes | ||
| 77 | PRKG1 | Prkg1 | 15377 | -2.412 | -0.0973 | Yes | ||
| 78 | ADCY8 | Adcy8 | 15381 | -2.433 | -0.0492 | Yes | ||
| 79 | PRKCB | Prkcb | 15461 | -2.954 | 0.0041 | Yes |