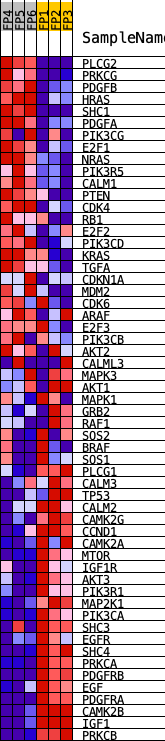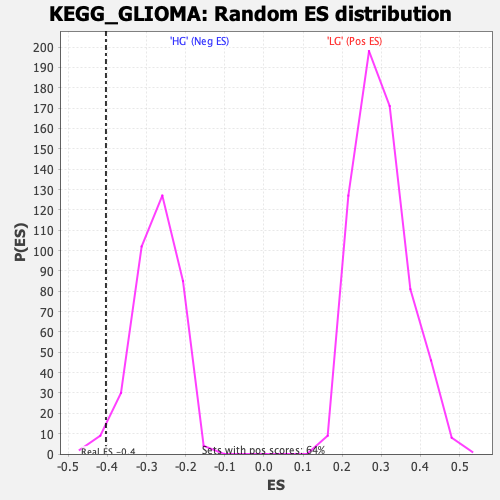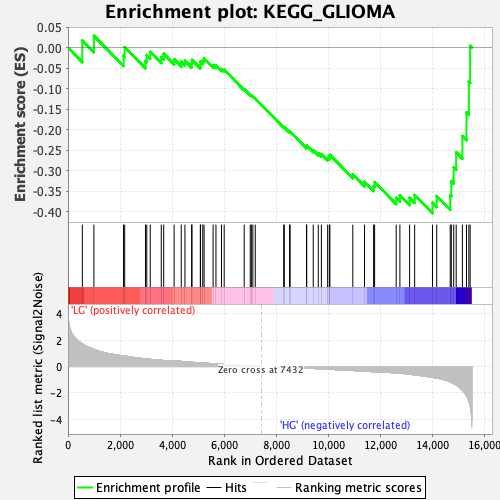
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_GLIOMA |
| Enrichment Score (ES) | -0.4039426 |
| Normalized Enrichment Score (NES) | -1.4569918 |
| Nominal p-value | 0.025069637 |
| FDR q-value | 0.23113047 |
| FWER p-Value | 0.9 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLCG2 | Plcg2 | 551 | 1.717 | 0.0172 | No | ||
| 2 | PRKCG | Prkcg | 996 | 1.303 | 0.0285 | No | ||
| 3 | PDGFB | Pdgfb | 2135 | 0.802 | -0.0204 | No | ||
| 4 | HRAS | Hras | 2180 | 0.791 | 0.0010 | No | ||
| 5 | SHC1 | Shc1 | 2978 | 0.579 | -0.0327 | No | ||
| 6 | PDGFA | Pdgfa | 3028 | 0.566 | -0.0185 | No | ||
| 7 | PIK3CG | Pik3cg | 3162 | 0.541 | -0.0104 | No | ||
| 8 | E2F1 | E2f1 | 3586 | 0.473 | -0.0232 | No | ||
| 9 | NRAS | Nras | 3680 | 0.455 | -0.0153 | No | ||
| 10 | PIK3R5 | Pik3r5 | 4080 | 0.414 | -0.0284 | No | ||
| 11 | CALM1 | Calm1 | 4354 | 0.386 | -0.0341 | No | ||
| 12 | PTEN | Pten | 4496 | 0.362 | -0.0321 | No | ||
| 13 | CDK4 | Cdk4 | 4753 | 0.318 | -0.0389 | No | ||
| 14 | RB1 | Rb1 | 4767 | 0.317 | -0.0300 | No | ||
| 15 | E2F2 | E2f2 | 5087 | 0.275 | -0.0422 | No | ||
| 16 | PIK3CD | Pik3cd | 5095 | 0.274 | -0.0342 | No | ||
| 17 | KRAS | Kras | 5181 | 0.266 | -0.0315 | No | ||
| 18 | TGFA | Tgfa | 5231 | 0.257 | -0.0268 | No | ||
| 19 | CDKN1A | Cdkn1a | 5579 | 0.211 | -0.0427 | No | ||
| 20 | MDM2 | Mdm2 | 5688 | 0.197 | -0.0437 | No | ||
| 21 | CDK6 | Cdk6 | 5902 | 0.167 | -0.0523 | No | ||
| 22 | ARAF | Araf | 6005 | 0.156 | -0.0541 | No | ||
| 23 | E2F3 | E2f3 | 6774 | 0.069 | -0.1016 | No | ||
| 24 | PIK3CB | Pik3cb | 7007 | 0.045 | -0.1152 | No | ||
| 25 | AKT2 | Akt2 | 7049 | 0.039 | -0.1167 | No | ||
| 26 | CALML3 | Calml3 | 7106 | 0.034 | -0.1192 | No | ||
| 27 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.1245 | No | ||
| 28 | AKT1 | Akt1 | 8293 | -0.035 | -0.1940 | No | ||
| 29 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.1943 | No | ||
| 30 | GRB2 | Grb2 | 8518 | -0.055 | -0.2056 | No | ||
| 31 | RAF1 | Raf1 | 8539 | -0.058 | -0.2052 | No | ||
| 32 | SOS2 | Sos2 | 9170 | -0.122 | -0.2421 | No | ||
| 33 | BRAF | Braf | 9178 | -0.123 | -0.2388 | No | ||
| 34 | SOS1 | Sos1 | 9428 | -0.146 | -0.2504 | No | ||
| 35 | PLCG1 | Plcg1 | 9622 | -0.167 | -0.2578 | No | ||
| 36 | CALM3 | Calm3 | 9742 | -0.178 | -0.2600 | No | ||
| 37 | TP53 | Trp53 | 9983 | -0.202 | -0.2693 | No | ||
| 38 | CALM2 | Calm2 | 10048 | -0.209 | -0.2670 | No | ||
| 39 | CAMK2G | Camk2g | 10068 | -0.211 | -0.2618 | No | ||
| 40 | CCND1 | Ccnd1 | 10950 | -0.301 | -0.3095 | No | ||
| 41 | CAMK2A | Camk2a | 11400 | -0.353 | -0.3277 | No | ||
| 42 | MTOR | Mtor | 11749 | -0.395 | -0.3380 | No | ||
| 43 | IGF1R | Igf1r | 11788 | -0.400 | -0.3282 | No | ||
| 44 | AKT3 | Akt3 | 12618 | -0.490 | -0.3667 | No | ||
| 45 | PIK3R1 | Pik3r1 | 12760 | -0.509 | -0.3602 | No | ||
| 46 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.3665 | No | ||
| 47 | PIK3CA | Pik3ca | 13327 | -0.625 | -0.3597 | No | ||
| 48 | SHC3 | Shc3 | 14013 | -0.825 | -0.3786 | Yes | ||
| 49 | EGFR | Egfr | 14176 | -0.870 | -0.3623 | Yes | ||
| 50 | SHC4 | Shc4 | 14695 | -1.160 | -0.3602 | Yes | ||
| 51 | PRKCA | Prkca | 14737 | -1.199 | -0.3260 | Yes | ||
| 52 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.2923 | Yes | ||
| 53 | EGF | Egf | 14924 | -1.396 | -0.2555 | Yes | ||
| 54 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.2159 | Yes | ||
| 55 | CAMK2B | Camk2b | 15320 | -2.213 | -0.1580 | Yes | ||
| 56 | IGF1 | Igf1 | 15412 | -2.616 | -0.0835 | Yes | ||
| 57 | PRKCB | Prkcb | 15461 | -2.954 | 0.0041 | Yes |