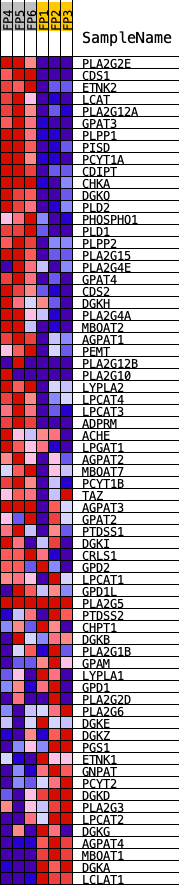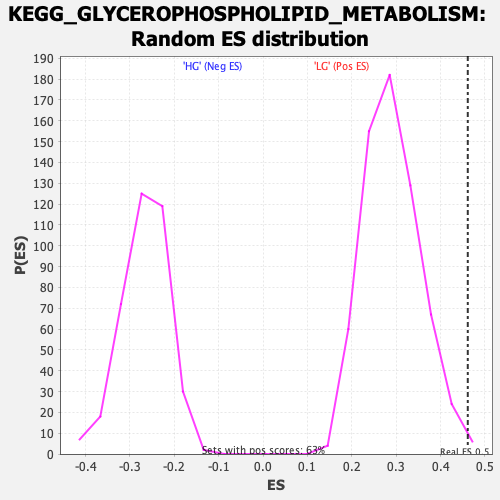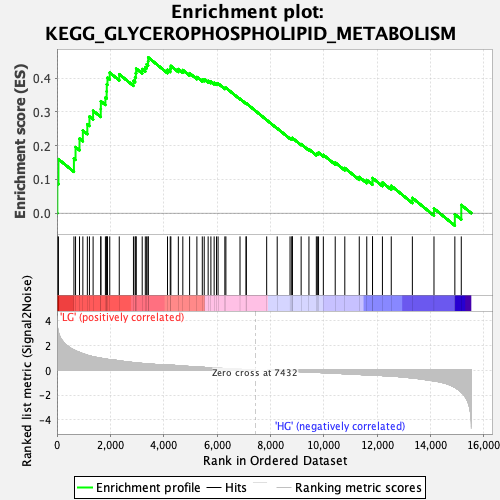
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_GLYCEROPHOSPHOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.46125993 |
| Normalized Enrichment Score (NES) | 1.5880698 |
| Nominal p-value | 0.0063795852 |
| FDR q-value | 0.117359854 |
| FWER p-Value | 0.666 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLA2G2E | Pla2g2e | 13 | 3.648 | 0.0881 | Yes | ||
| 2 | CDS1 | Cds1 | 54 | 3.051 | 0.1599 | Yes | ||
| 3 | ETNK2 | Etnk2 | 634 | 1.611 | 0.1617 | Yes | ||
| 4 | LCAT | Lcat | 695 | 1.553 | 0.1957 | Yes | ||
| 5 | PLA2G12A | Pla2g12a | 844 | 1.420 | 0.2208 | Yes | ||
| 6 | GPAT3 | Gpat3 | 969 | 1.319 | 0.2449 | Yes | ||
| 7 | PLPP1 | Plpp1 | 1137 | 1.198 | 0.2633 | Yes | ||
| 8 | PISD | Pisd | 1216 | 1.142 | 0.2861 | Yes | ||
| 9 | PCYT1A | Pcyt1a | 1353 | 1.073 | 0.3035 | Yes | ||
| 10 | CDIPT | Cdipt | 1640 | 0.958 | 0.3084 | Yes | ||
| 11 | CHKA | Chka | 1644 | 0.956 | 0.3315 | Yes | ||
| 12 | DGKQ | Dgkq | 1814 | 0.891 | 0.3423 | Yes | ||
| 13 | PLD2 | Pld2 | 1861 | 0.875 | 0.3606 | Yes | ||
| 14 | PHOSPHO1 | Phospho1 | 1866 | 0.873 | 0.3816 | Yes | ||
| 15 | PLD1 | Pld1 | 1892 | 0.866 | 0.4012 | Yes | ||
| 16 | PLPP2 | Plpp2 | 1977 | 0.839 | 0.4162 | Yes | ||
| 17 | PLA2G15 | Pla2g15 | 2333 | 0.745 | 0.4114 | Yes | ||
| 18 | PLA2G4E | Pla2g4e | 2865 | 0.604 | 0.3918 | Yes | ||
| 19 | GPAT4 | Gpat4 | 2926 | 0.591 | 0.4023 | Yes | ||
| 20 | CDS2 | Cds2 | 2954 | 0.585 | 0.4148 | Yes | ||
| 21 | DGKH | Dgkh | 2967 | 0.581 | 0.4282 | Yes | ||
| 22 | PLA2G4A | Pla2g4a | 3193 | 0.536 | 0.4267 | Yes | ||
| 23 | MBOAT2 | Mboat2 | 3309 | 0.512 | 0.4318 | Yes | ||
| 24 | AGPAT1 | Agpat1 | 3359 | 0.505 | 0.4409 | Yes | ||
| 25 | PEMT | Pemt | 3413 | 0.494 | 0.4495 | Yes | ||
| 26 | PLA2G12B | Pla2g12b | 3419 | 0.494 | 0.4613 | Yes | ||
| 27 | PLA2G10 | Pla2g10 | 4141 | 0.407 | 0.4245 | No | ||
| 28 | LYPLA2 | Lypla2 | 4243 | 0.404 | 0.4278 | No | ||
| 29 | LPCAT4 | Lpcat4 | 4267 | 0.401 | 0.4361 | No | ||
| 30 | LPCAT3 | Lpcat3 | 4547 | 0.353 | 0.4267 | No | ||
| 31 | ADPRM | Adprm | 4715 | 0.324 | 0.4238 | No | ||
| 32 | ACHE | Ache | 4969 | 0.289 | 0.4145 | No | ||
| 33 | LPGAT1 | Lpgat1 | 5242 | 0.255 | 0.4031 | No | ||
| 34 | AGPAT2 | Agpat2 | 5445 | 0.228 | 0.3956 | No | ||
| 35 | MBOAT7 | Mboat7 | 5517 | 0.218 | 0.3963 | No | ||
| 36 | PCYT1B | Pcyt1b | 5664 | 0.199 | 0.3917 | No | ||
| 37 | TAZ | Taz | 5767 | 0.185 | 0.3896 | No | ||
| 38 | AGPAT3 | Agpat3 | 5889 | 0.169 | 0.3859 | No | ||
| 39 | GPAT2 | Gpat2 | 5978 | 0.159 | 0.3841 | No | ||
| 40 | PTDSS1 | Ptdss1 | 6036 | 0.152 | 0.3841 | No | ||
| 41 | DGKI | Dgki | 6290 | 0.125 | 0.3708 | No | ||
| 42 | CRLS1 | Crls1 | 6329 | 0.120 | 0.3713 | No | ||
| 43 | GPD2 | Gpd2 | 6858 | 0.061 | 0.3386 | No | ||
| 44 | LPCAT1 | Lpcat1 | 7083 | 0.036 | 0.3250 | No | ||
| 45 | GPD1L | Gpd1l | 7095 | 0.035 | 0.3252 | No | ||
| 46 | PLA2G5 | Pla2g5 | 7857 | 0.000 | 0.2759 | No | ||
| 47 | PTDSS2 | Ptdss2 | 8252 | -0.031 | 0.2512 | No | ||
| 48 | CHPT1 | Chpt1 | 8733 | -0.078 | 0.2220 | No | ||
| 49 | DGKB | Dgkb | 8801 | -0.083 | 0.2197 | No | ||
| 50 | PLA2G1B | Pla2g1b | 8815 | -0.085 | 0.2210 | No | ||
| 51 | GPAM | Gpam | 8818 | -0.085 | 0.2229 | No | ||
| 52 | LYPLA1 | Lypla1 | 9149 | -0.120 | 0.2045 | No | ||
| 53 | GPD1 | Gpd1 | 9441 | -0.147 | 0.1892 | No | ||
| 54 | PLA2G2D | Pla2g2d | 9718 | -0.175 | 0.1757 | No | ||
| 55 | PLA2G6 | Pla2g6 | 9756 | -0.179 | 0.1776 | No | ||
| 56 | DGKE | Dgke | 9801 | -0.184 | 0.1792 | No | ||
| 57 | DGKZ | Dgkz | 9981 | -0.202 | 0.1726 | No | ||
| 58 | PGS1 | Pgs1 | 10426 | -0.248 | 0.1499 | No | ||
| 59 | ETNK1 | Etnk1 | 10785 | -0.285 | 0.1337 | No | ||
| 60 | GNPAT | Gnpat | 11327 | -0.344 | 0.1071 | No | ||
| 61 | PCYT2 | Pcyt2 | 11608 | -0.377 | 0.0982 | No | ||
| 62 | DGKD | Dgkd | 11823 | -0.406 | 0.0942 | No | ||
| 63 | PLA2G3 | Pla2g3 | 11824 | -0.406 | 0.1041 | No | ||
| 64 | LPCAT2 | Lpcat2 | 12195 | -0.434 | 0.0907 | No | ||
| 65 | DGKG | Dgkg | 12526 | -0.479 | 0.0811 | No | ||
| 66 | AGPAT4 | Agpat4 | 13317 | -0.624 | 0.0452 | No | ||
| 67 | MBOAT1 | Mboat1 | 14127 | -0.857 | 0.0137 | No | ||
| 68 | DGKA | Dgka | 14909 | -1.378 | -0.0032 | No | ||
| 69 | LCLAT1 | Lclat1 | 15150 | -1.763 | 0.0243 | No |