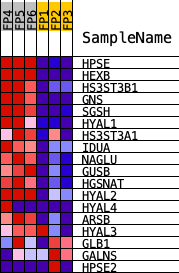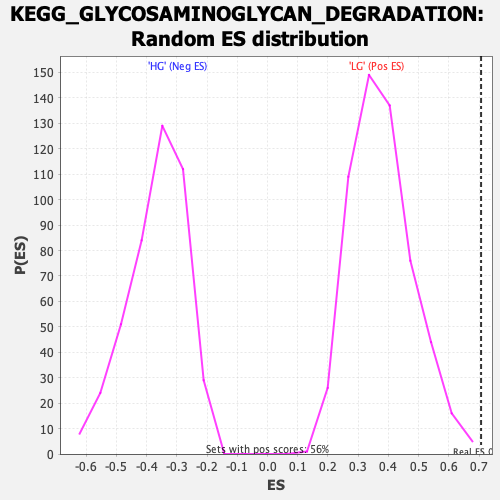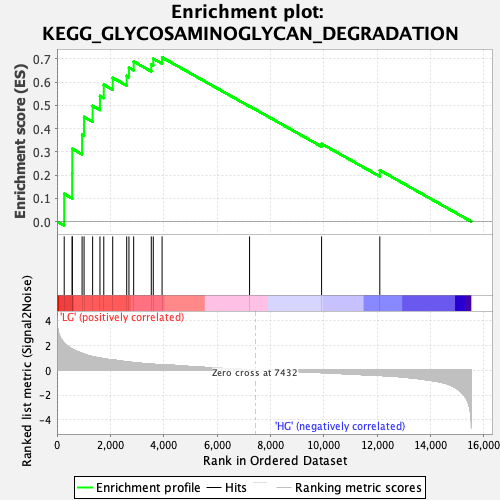
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_GLYCOSAMINOGLYCAN_DEGRADATION |
| Enrichment Score (ES) | 0.7066219 |
| Normalized Enrichment Score (NES) | 1.8751518 |
| Nominal p-value | 0.001776199 |
| FDR q-value | 0.0078560645 |
| FWER p-Value | 0.025 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HPSE | Hpse | 270 | 2.166 | 0.1191 | Yes | ||
| 2 | HEXB | Hexb | 568 | 1.696 | 0.2068 | Yes | ||
| 3 | HS3ST3B1 | Hs3st3b1 | 573 | 1.691 | 0.3131 | Yes | ||
| 4 | GNS | Gns | 940 | 1.341 | 0.3740 | Yes | ||
| 5 | SGSH | Sgsh | 1014 | 1.287 | 0.4504 | Yes | ||
| 6 | HYAL1 | Hyal1 | 1335 | 1.080 | 0.4978 | Yes | ||
| 7 | HS3ST3A1 | Hs3st3a1 | 1610 | 0.965 | 0.5409 | Yes | ||
| 8 | IDUA | Idua | 1753 | 0.911 | 0.5892 | Yes | ||
| 9 | NAGLU | Naglu | 2088 | 0.812 | 0.6188 | Yes | ||
| 10 | GUSB | Gusb | 2612 | 0.665 | 0.6270 | Yes | ||
| 11 | HGSNAT | Hgsnat | 2696 | 0.644 | 0.6622 | Yes | ||
| 12 | HYAL2 | Hyal2 | 2873 | 0.603 | 0.6889 | Yes | ||
| 13 | HYAL4 | Hyal4 | 3533 | 0.477 | 0.6765 | Yes | ||
| 14 | ARSB | Arsb | 3608 | 0.469 | 0.7012 | Yes | ||
| 15 | HYAL3 | Hyal3 | 3938 | 0.423 | 0.7066 | Yes | ||
| 16 | GLB1 | Glb1 | 7216 | 0.021 | 0.4967 | No | ||
| 17 | GALNS | Galns | 9915 | -0.195 | 0.3350 | No | ||
| 18 | HPSE2 | Hpse2 | 12097 | -0.424 | 0.2210 | No |