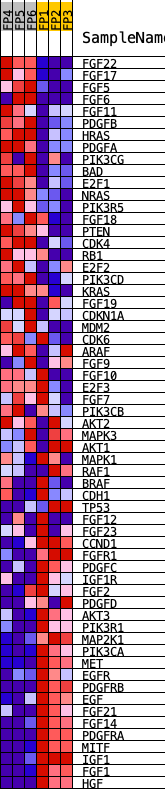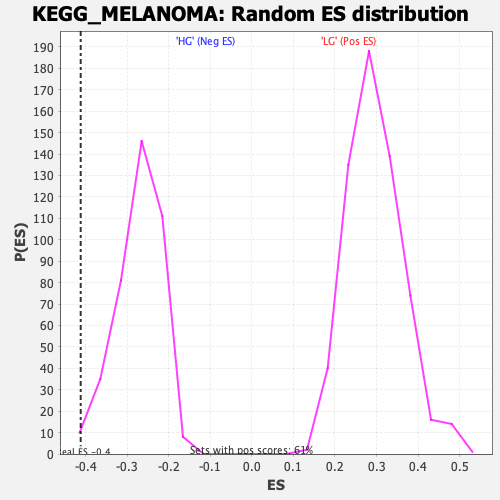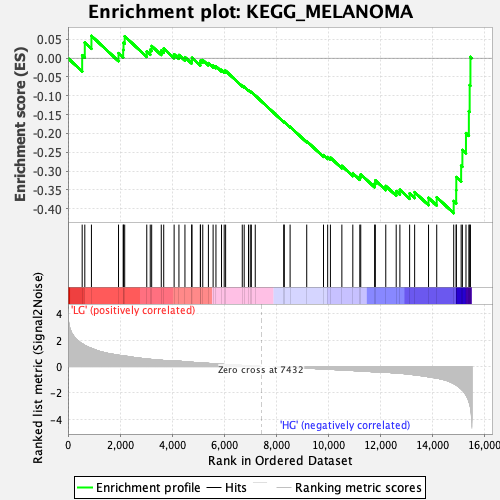
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_MELANOMA |
| Enrichment Score (ES) | -0.41178805 |
| Normalized Enrichment Score (NES) | -1.5119929 |
| Nominal p-value | 0.012787724 |
| FDR q-value | 0.21055491 |
| FWER p-Value | 0.775 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FGF22 | Fgf22 | 545 | 1.720 | 0.0078 | No | ||
| 2 | FGF17 | Fgf17 | 646 | 1.600 | 0.0414 | No | ||
| 3 | FGF5 | Fgf5 | 901 | 1.373 | 0.0593 | No | ||
| 4 | FGF6 | Fgf6 | 1942 | 0.850 | 0.0133 | No | ||
| 5 | FGF11 | Fgf11 | 2121 | 0.805 | 0.0220 | No | ||
| 6 | PDGFB | Pdgfb | 2135 | 0.802 | 0.0412 | No | ||
| 7 | HRAS | Hras | 2180 | 0.791 | 0.0582 | No | ||
| 8 | PDGFA | Pdgfa | 3028 | 0.566 | 0.0176 | No | ||
| 9 | PIK3CG | Pik3cg | 3162 | 0.541 | 0.0225 | No | ||
| 10 | BAD | Bad | 3217 | 0.532 | 0.0323 | No | ||
| 11 | E2F1 | E2f1 | 3586 | 0.473 | 0.0204 | No | ||
| 12 | NRAS | Nras | 3680 | 0.455 | 0.0257 | No | ||
| 13 | PIK3R5 | Pik3r5 | 4080 | 0.414 | 0.0103 | No | ||
| 14 | FGF18 | Fgf18 | 4265 | 0.401 | 0.0084 | No | ||
| 15 | PTEN | Pten | 4496 | 0.362 | 0.0026 | No | ||
| 16 | CDK4 | Cdk4 | 4753 | 0.318 | -0.0060 | No | ||
| 17 | RB1 | Rb1 | 4767 | 0.317 | 0.0011 | No | ||
| 18 | E2F2 | E2f2 | 5087 | 0.275 | -0.0126 | No | ||
| 19 | PIK3CD | Pik3cd | 5095 | 0.274 | -0.0062 | No | ||
| 20 | KRAS | Kras | 5181 | 0.266 | -0.0051 | No | ||
| 21 | FGF19 | Fgf15 | 5398 | 0.234 | -0.0132 | No | ||
| 22 | CDKN1A | Cdkn1a | 5579 | 0.211 | -0.0195 | No | ||
| 23 | MDM2 | Mdm2 | 5688 | 0.197 | -0.0216 | No | ||
| 24 | CDK6 | Cdk6 | 5902 | 0.167 | -0.0312 | No | ||
| 25 | ARAF | Araf | 6005 | 0.156 | -0.0339 | No | ||
| 26 | FGF9 | Fgf9 | 6060 | 0.149 | -0.0336 | No | ||
| 27 | FGF10 | Fgf10 | 6700 | 0.078 | -0.0730 | No | ||
| 28 | E2F3 | E2f3 | 6774 | 0.069 | -0.0760 | No | ||
| 29 | FGF7 | Fgf7 | 6940 | 0.053 | -0.0853 | No | ||
| 30 | PIK3CB | Pik3cb | 7007 | 0.045 | -0.0885 | No | ||
| 31 | AKT2 | Akt2 | 7049 | 0.039 | -0.0901 | No | ||
| 32 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.0992 | No | ||
| 33 | AKT1 | Akt1 | 8293 | -0.035 | -0.1690 | No | ||
| 34 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.1695 | No | ||
| 35 | RAF1 | Raf1 | 8539 | -0.058 | -0.1824 | No | ||
| 36 | BRAF | Braf | 9178 | -0.123 | -0.2206 | No | ||
| 37 | CDH1 | Cdh1 | 9823 | -0.185 | -0.2576 | No | ||
| 38 | TP53 | Trp53 | 9983 | -0.202 | -0.2628 | No | ||
| 39 | FGF12 | Fgf12 | 10089 | -0.213 | -0.2643 | No | ||
| 40 | FGF23 | Fgf23 | 10529 | -0.257 | -0.2862 | No | ||
| 41 | CCND1 | Ccnd1 | 10950 | -0.301 | -0.3058 | No | ||
| 42 | FGFR1 | Fgfr1 | 11219 | -0.332 | -0.3149 | No | ||
| 43 | PDGFC | Pdgfc | 11255 | -0.336 | -0.3087 | No | ||
| 44 | IGF1R | Igf1r | 11788 | -0.400 | -0.3331 | No | ||
| 45 | FGF2 | Fgf2 | 11816 | -0.404 | -0.3247 | No | ||
| 46 | PDGFD | Pdgfd | 12217 | -0.437 | -0.3397 | No | ||
| 47 | AKT3 | Akt3 | 12618 | -0.490 | -0.3533 | No | ||
| 48 | PIK3R1 | Pik3r1 | 12760 | -0.509 | -0.3497 | No | ||
| 49 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.3592 | No | ||
| 50 | PIK3CA | Pik3ca | 13327 | -0.625 | -0.3560 | No | ||
| 51 | MET | Met | 13861 | -0.774 | -0.3711 | No | ||
| 52 | EGFR | Egfr | 14176 | -0.870 | -0.3696 | No | ||
| 53 | PDGFRB | Pdgfrb | 14829 | -1.288 | -0.3795 | Yes | ||
| 54 | EGF | Egf | 14924 | -1.396 | -0.3507 | Yes | ||
| 55 | FGF21 | Fgf21 | 14929 | -1.400 | -0.3159 | Yes | ||
| 56 | FGF14 | Fgf14 | 15115 | -1.689 | -0.2856 | Yes | ||
| 57 | PDGFRA | Pdgfra | 15163 | -1.787 | -0.2439 | Yes | ||
| 58 | MITF | Mitf | 15301 | -2.140 | -0.1992 | Yes | ||
| 59 | IGF1 | Igf1 | 15412 | -2.616 | -0.1409 | Yes | ||
| 60 | FGF1 | Fgf1 | 15443 | -2.849 | -0.0715 | Yes | ||
| 61 | HGF | Hgf | 15472 | -3.068 | 0.0034 | Yes |