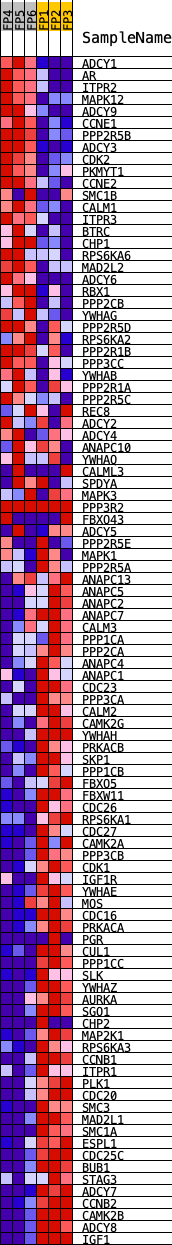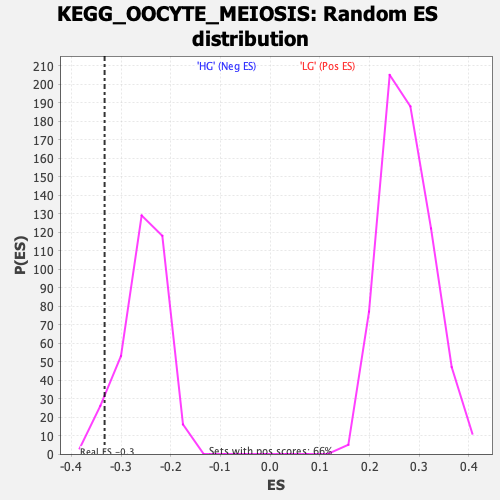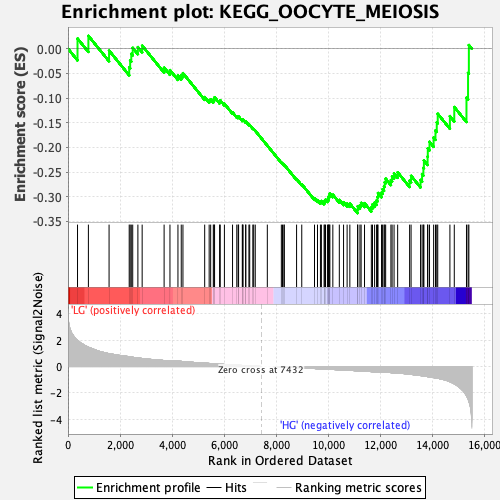
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_OOCYTE_MEIOSIS |
| Enrichment Score (ES) | -0.33316225 |
| Normalized Enrichment Score (NES) | -1.3119867 |
| Nominal p-value | 0.046376813 |
| FDR q-value | 0.3620415 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADCY1 | Adcy1 | 366 | 1.982 | 0.0204 | No | ||
| 2 | AR | Ar | 784 | 1.461 | 0.0259 | No | ||
| 3 | ITPR2 | Itpr2 | 1579 | 0.974 | -0.0039 | No | ||
| 4 | MAPK12 | Mapk12 | 2353 | 0.740 | -0.0375 | No | ||
| 5 | ADCY9 | Adcy9 | 2390 | 0.729 | -0.0236 | No | ||
| 6 | CCNE1 | Ccne1 | 2433 | 0.714 | -0.0104 | No | ||
| 7 | PPP2R5B | Ppp2r5b | 2488 | 0.698 | 0.0016 | No | ||
| 8 | ADCY3 | Adcy3 | 2687 | 0.646 | 0.0032 | No | ||
| 9 | CDK2 | Cdk2 | 2851 | 0.607 | 0.0061 | No | ||
| 10 | PKMYT1 | Pkmyt1 | 3695 | 0.453 | -0.0385 | No | ||
| 11 | CCNE2 | Ccne2 | 3920 | 0.425 | -0.0435 | No | ||
| 12 | SMC1B | Smc1b | 4227 | 0.406 | -0.0543 | No | ||
| 13 | CALM1 | Calm1 | 4354 | 0.386 | -0.0539 | No | ||
| 14 | ITPR3 | Itpr3 | 4419 | 0.376 | -0.0496 | No | ||
| 15 | BTRC | Btrc | 5255 | 0.254 | -0.0981 | No | ||
| 16 | CHP1 | Chp1 | 5427 | 0.231 | -0.1040 | No | ||
| 17 | RPS6KA6 | Rps6ka6 | 5485 | 0.222 | -0.1028 | No | ||
| 18 | MAD2L2 | Mad2l2 | 5586 | 0.211 | -0.1046 | No | ||
| 19 | ADCY6 | Adcy6 | 5613 | 0.207 | -0.1017 | No | ||
| 20 | RBX1 | Rbx1 | 5635 | 0.202 | -0.0985 | No | ||
| 21 | PPP2CB | Ppp2cb | 5828 | 0.177 | -0.1070 | No | ||
| 22 | YWHAG | Ywhag | 5853 | 0.174 | -0.1047 | No | ||
| 23 | PPP2R5D | Ppp2r5d | 6013 | 0.154 | -0.1116 | No | ||
| 24 | RPS6KA2 | Rps6ka2 | 6326 | 0.121 | -0.1291 | No | ||
| 25 | PPP2R1B | Ppp2r1b | 6489 | 0.102 | -0.1373 | No | ||
| 26 | PPP3CC | Ppp3cc | 6542 | 0.096 | -0.1386 | No | ||
| 27 | YWHAB | Ywhab | 6559 | 0.094 | -0.1375 | No | ||
| 28 | PPP2R1A | Ppp2r1a | 6701 | 0.078 | -0.1449 | No | ||
| 29 | PPP2R5C | Ppp2r5c | 6712 | 0.076 | -0.1439 | No | ||
| 30 | REC8 | Rec8 | 6730 | 0.074 | -0.1433 | No | ||
| 31 | ADCY2 | Adcy2 | 6832 | 0.063 | -0.1485 | No | ||
| 32 | ADCY4 | Adcy4 | 6834 | 0.063 | -0.1471 | No | ||
| 33 | ANAPC10 | Anapc10 | 6956 | 0.051 | -0.1538 | No | ||
| 34 | YWHAQ | Ywhaq | 6986 | 0.048 | -0.1547 | No | ||
| 35 | CALML3 | Calml3 | 7106 | 0.034 | -0.1616 | No | ||
| 36 | SPDYA | Spdya | 7140 | 0.031 | -0.1631 | No | ||
| 37 | MAPK3 | Mapk3 | 7199 | 0.024 | -0.1663 | No | ||
| 38 | PPP3R2 | Ppp3r2 | 7663 | 0.000 | -0.1963 | No | ||
| 39 | FBXO43 | Fbxo43 | 8202 | -0.028 | -0.2305 | No | ||
| 40 | ADCY5 | Adcy5 | 8247 | -0.030 | -0.2327 | No | ||
| 41 | PPP2R5E | Ppp2r5e | 8301 | -0.036 | -0.2354 | No | ||
| 42 | MAPK1 | Mapk1 | 8316 | -0.037 | -0.2354 | No | ||
| 43 | PPP2R5A | Ppp2r5a | 8788 | -0.083 | -0.2641 | No | ||
| 44 | ANAPC13 | Anapc13 | 8987 | -0.102 | -0.2747 | No | ||
| 45 | ANAPC5 | Anapc5 | 9477 | -0.151 | -0.3030 | No | ||
| 46 | ANAPC2 | Anapc2 | 9592 | -0.163 | -0.3068 | No | ||
| 47 | ANAPC7 | Anapc7 | 9705 | -0.174 | -0.3102 | No | ||
| 48 | CALM3 | Calm3 | 9742 | -0.178 | -0.3085 | No | ||
| 49 | PPP1CA | Ppp1ca | 9846 | -0.188 | -0.3110 | No | ||
| 50 | PPP2CA | Ppp2ca | 9869 | -0.190 | -0.3082 | No | ||
| 51 | ANAPC4 | Anapc4 | 9900 | -0.193 | -0.3059 | No | ||
| 52 | ANAPC1 | Anapc1 | 9975 | -0.201 | -0.3062 | No | ||
| 53 | CDC23 | Cdc23 | 10009 | -0.204 | -0.3038 | No | ||
| 54 | PPP3CA | Ppp3ca | 10018 | -0.206 | -0.2997 | No | ||
| 55 | CALM2 | Calm2 | 10048 | -0.209 | -0.2969 | No | ||
| 56 | CAMK2G | Camk2g | 10068 | -0.211 | -0.2935 | No | ||
| 57 | YWHAH | Ywhah | 10182 | -0.221 | -0.2959 | No | ||
| 58 | PRKACB | Prkacb | 10431 | -0.249 | -0.3064 | No | ||
| 59 | SKP1 | Skp1a | 10594 | -0.265 | -0.3110 | No | ||
| 60 | PPP1CB | Ppp1cb | 10731 | -0.279 | -0.3136 | No | ||
| 61 | FBXO5 | Fbxo5 | 10836 | -0.290 | -0.3139 | No | ||
| 62 | FBXW11 | Fbxw11 | 11134 | -0.323 | -0.3260 | Yes | ||
| 63 | CDC26 | Cdc26 | 11141 | -0.324 | -0.3191 | Yes | ||
| 64 | RPS6KA1 | Rps6ka1 | 11224 | -0.333 | -0.3170 | Yes | ||
| 65 | CDC27 | Cdc27 | 11269 | -0.337 | -0.3124 | Yes | ||
| 66 | CAMK2A | Camk2a | 11400 | -0.353 | -0.3130 | Yes | ||
| 67 | PPP3CB | Ppp3cb | 11662 | -0.384 | -0.3213 | Yes | ||
| 68 | CDK1 | Cdk1 | 11707 | -0.390 | -0.3155 | Yes | ||
| 69 | IGF1R | Igf1r | 11788 | -0.400 | -0.3117 | Yes | ||
| 70 | YWHAE | Ywhae | 11863 | -0.410 | -0.3074 | Yes | ||
| 71 | MOS | Mos | 11899 | -0.415 | -0.3004 | Yes | ||
| 72 | CDC16 | Cdc16 | 11919 | -0.418 | -0.2924 | Yes | ||
| 73 | PRKACA | Prkaca | 12054 | -0.424 | -0.2916 | Yes | ||
| 74 | PGR | Pgr | 12095 | -0.424 | -0.2848 | Yes | ||
| 75 | CUL1 | Cul1 | 12150 | -0.428 | -0.2787 | Yes | ||
| 76 | PPP1CC | Ppp1cc | 12176 | -0.431 | -0.2707 | Yes | ||
| 77 | SLK | Slk | 12213 | -0.436 | -0.2634 | Yes | ||
| 78 | YWHAZ | Ywhaz | 12415 | -0.460 | -0.2662 | Yes | ||
| 79 | AURKA | Aurka | 12460 | -0.468 | -0.2586 | Yes | ||
| 80 | SGO1 | Sgo1 | 12536 | -0.481 | -0.2527 | Yes | ||
| 81 | CHP2 | Chp2 | 12674 | -0.499 | -0.2505 | Yes | ||
| 82 | MAP2K1 | Map2k1 | 13134 | -0.581 | -0.2673 | Yes | ||
| 83 | RPS6KA3 | Rps6ka3 | 13190 | -0.597 | -0.2576 | Yes | ||
| 84 | CCNB1 | Ccnb1 | 13554 | -0.683 | -0.2659 | Yes | ||
| 85 | ITPR1 | Itpr1 | 13613 | -0.701 | -0.2541 | Yes | ||
| 86 | PLK1 | Plk1 | 13663 | -0.714 | -0.2413 | Yes | ||
| 87 | CDC20 | Cdc20 | 13683 | -0.722 | -0.2265 | Yes | ||
| 88 | SMC3 | Smc3 | 13825 | -0.764 | -0.2186 | Yes | ||
| 89 | MAD2L1 | Mad2l1 | 13833 | -0.766 | -0.2020 | Yes | ||
| 90 | SMC1A | Smc1a | 13899 | -0.788 | -0.1887 | Yes | ||
| 91 | ESPL1 | Espl1 | 14057 | -0.839 | -0.1802 | Yes | ||
| 92 | CDC25C | Cdc25c | 14128 | -0.858 | -0.1656 | Yes | ||
| 93 | BUB1 | Bub1 | 14178 | -0.872 | -0.1494 | Yes | ||
| 94 | STAG3 | Stag3 | 14215 | -0.888 | -0.1319 | Yes | ||
| 95 | ADCY7 | Adcy7 | 14683 | -1.147 | -0.1367 | Yes | ||
| 96 | CCNB2 | Ccnb2 | 14851 | -1.313 | -0.1182 | Yes | ||
| 97 | CAMK2B | Camk2b | 15320 | -2.213 | -0.0993 | Yes | ||
| 98 | ADCY8 | Adcy8 | 15381 | -2.433 | -0.0490 | Yes | ||
| 99 | IGF1 | Igf1 | 15412 | -2.616 | 0.0073 | Yes |