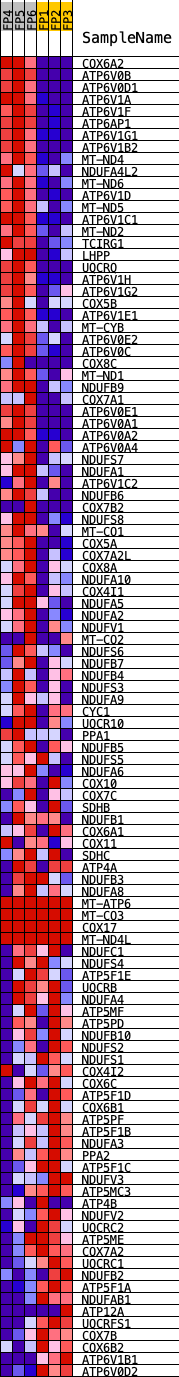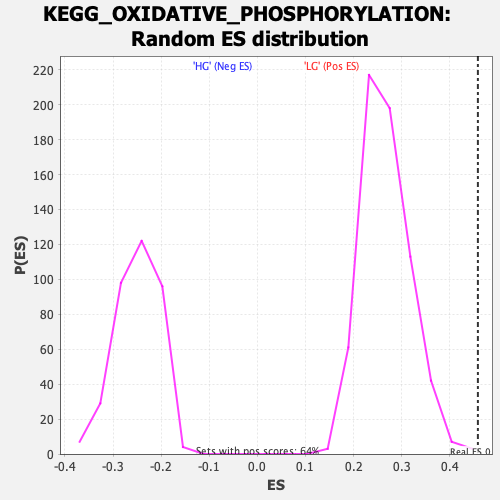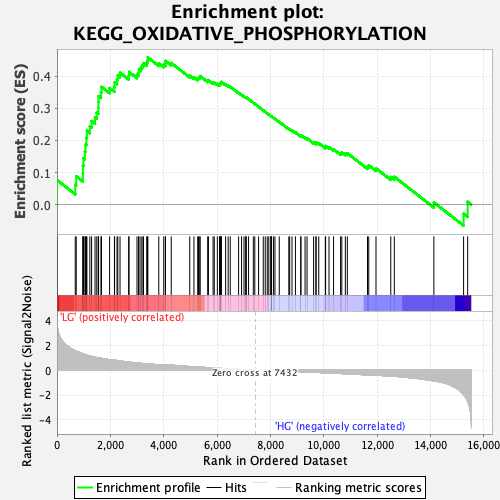
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | 0.4585911 |
| Normalized Enrichment Score (NES) | 1.7159932 |
| Nominal p-value | 0.0015527951 |
| FDR q-value | 0.064198524 |
| FWER p-Value | 0.236 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COX6A2 | Cox6a2 | 3 | 4.063 | 0.0763 | Yes | ||
| 2 | ATP6V0B | Atp6v0b | 684 | 1.564 | 0.0616 | Yes | ||
| 3 | ATP6V0D1 | Atp6v0d1 | 720 | 1.529 | 0.0881 | Yes | ||
| 4 | ATP6V1A | Atp6v1a | 968 | 1.319 | 0.0969 | Yes | ||
| 5 | ATP6V1F | Atp6v1f | 973 | 1.317 | 0.1214 | Yes | ||
| 6 | ATP6AP1 | Atp6ap1 | 994 | 1.304 | 0.1447 | Yes | ||
| 7 | ATP6V1G1 | Atp6v1g1 | 1046 | 1.261 | 0.1651 | Yes | ||
| 8 | ATP6V1B2 | Atp6v1b2 | 1067 | 1.247 | 0.1873 | Yes | ||
| 9 | MT-ND4 | mt-Nd4 | 1100 | 1.223 | 0.2082 | Yes | ||
| 10 | NDUFA4L2 | Ndufa4l2 | 1121 | 1.210 | 0.2297 | Yes | ||
| 11 | MT-ND6 | mt-Nd6 | 1232 | 1.133 | 0.2439 | Yes | ||
| 12 | ATP6V1D | Atp6v1d | 1297 | 1.100 | 0.2605 | Yes | ||
| 13 | MT-ND5 | mt-Nd5 | 1425 | 1.042 | 0.2718 | Yes | ||
| 14 | ATP6V1C1 | Atp6v1c1 | 1491 | 1.008 | 0.2866 | Yes | ||
| 15 | MT-ND2 | mt-Nd2 | 1548 | 0.986 | 0.3015 | Yes | ||
| 16 | TCIRG1 | Tcirg1 | 1552 | 0.985 | 0.3199 | Yes | ||
| 17 | LHPP | Lhpp | 1558 | 0.982 | 0.3380 | Yes | ||
| 18 | UQCRQ | Uqcrq | 1648 | 0.955 | 0.3502 | Yes | ||
| 19 | ATP6V1H | Atp6v1h | 1666 | 0.945 | 0.3669 | Yes | ||
| 20 | ATP6V1G2 | Atp6v1g2 | 1971 | 0.843 | 0.3630 | Yes | ||
| 21 | COX5B | Cox5b | 2152 | 0.799 | 0.3664 | Yes | ||
| 22 | ATP6V1E1 | Atp6v1e1 | 2158 | 0.797 | 0.3811 | Yes | ||
| 23 | MT-CYB | mt-Cytb | 2250 | 0.773 | 0.3897 | Yes | ||
| 24 | ATP6V0E2 | Atp6v0e2 | 2279 | 0.764 | 0.4023 | Yes | ||
| 25 | ATP6V0C | Atp6v0c | 2362 | 0.737 | 0.4108 | Yes | ||
| 26 | COX8C | Cox8b | 2685 | 0.646 | 0.4021 | Yes | ||
| 27 | MT-ND1 | mt-Nd1 | 2700 | 0.642 | 0.4133 | Yes | ||
| 28 | NDUFB9 | Ndufb9 | 2998 | 0.573 | 0.4048 | Yes | ||
| 29 | COX7A1 | Cox7a1 | 3057 | 0.561 | 0.4116 | Yes | ||
| 30 | ATP6V0E1 | Atp6v0e | 3073 | 0.557 | 0.4211 | Yes | ||
| 31 | ATP6V0A1 | Atp6v0a1 | 3146 | 0.543 | 0.4266 | Yes | ||
| 32 | ATP6V0A2 | Atp6v0a2 | 3190 | 0.536 | 0.4339 | Yes | ||
| 33 | ATP6V0A4 | Atp6v0a4 | 3241 | 0.528 | 0.4406 | Yes | ||
| 34 | NDUFS7 | Ndufs7 | 3362 | 0.504 | 0.4423 | Yes | ||
| 35 | NDUFA1 | Ndufa1 | 3395 | 0.498 | 0.4496 | Yes | ||
| 36 | ATP6V1C2 | Atp6v1c2 | 3402 | 0.497 | 0.4586 | Yes | ||
| 37 | NDUFB6 | Ndufb6 | 3814 | 0.432 | 0.4401 | No | ||
| 38 | COX7B2 | Cox7b2 | 4000 | 0.421 | 0.4360 | No | ||
| 39 | NDUFS8 | Ndufs8 | 4063 | 0.417 | 0.4398 | No | ||
| 40 | MT-CO1 | mt-Co1 | 4064 | 0.417 | 0.4476 | No | ||
| 41 | COX5A | Cox5a | 4281 | 0.397 | 0.4411 | No | ||
| 42 | COX7A2L | Cox7a2l | 4974 | 0.289 | 0.4017 | No | ||
| 43 | COX8A | Cox8a | 5132 | 0.271 | 0.3966 | No | ||
| 44 | NDUFA10 | Ndufa10 | 5273 | 0.251 | 0.3922 | No | ||
| 45 | COX4I1 | Cox4i1 | 5301 | 0.248 | 0.3951 | No | ||
| 46 | NDUFA5 | Ndufa5 | 5352 | 0.242 | 0.3964 | No | ||
| 47 | NDUFA2 | Ndufa2 | 5361 | 0.240 | 0.4004 | No | ||
| 48 | NDUFV1 | Ndufv1 | 5648 | 0.201 | 0.3856 | No | ||
| 49 | MT-CO2 | mt-Co2 | 5674 | 0.198 | 0.3877 | No | ||
| 50 | NDUFS6 | Ndufs6 | 5849 | 0.174 | 0.3797 | No | ||
| 51 | NDUFB7 | Ndufb7 | 5894 | 0.168 | 0.3800 | No | ||
| 52 | NDUFB4 | Ndufb4 | 6009 | 0.155 | 0.3756 | No | ||
| 53 | NDUFS3 | Ndufs3 | 6090 | 0.146 | 0.3731 | No | ||
| 54 | NDUFA9 | Ndufa9 | 6103 | 0.145 | 0.3751 | No | ||
| 55 | CYC1 | Cyc1 | 6122 | 0.143 | 0.3766 | No | ||
| 56 | UQCR10 | Uqcr10 | 6139 | 0.141 | 0.3782 | No | ||
| 57 | PPA1 | Ppa1 | 6144 | 0.140 | 0.3806 | No | ||
| 58 | NDUFB5 | Ndufb5 | 6160 | 0.139 | 0.3823 | No | ||
| 59 | NDUFS5 | Ndufs5 | 6319 | 0.122 | 0.3743 | No | ||
| 60 | NDUFA6 | Ndufa6 | 6414 | 0.110 | 0.3703 | No | ||
| 61 | COX10 | Cox10 | 6492 | 0.102 | 0.3672 | No | ||
| 62 | COX7C | Cox7c | 6805 | 0.065 | 0.3482 | No | ||
| 63 | SDHB | Sdhb | 6917 | 0.055 | 0.3420 | No | ||
| 64 | NDUFB1 | Ndufb1-ps | 7008 | 0.045 | 0.3370 | No | ||
| 65 | COX6A1 | Cox6a1 | 7077 | 0.036 | 0.3333 | No | ||
| 66 | COX11 | Cox11 | 7084 | 0.036 | 0.3336 | No | ||
| 67 | SDHC | Sdhc | 7086 | 0.036 | 0.3342 | No | ||
| 68 | ATP4A | Atp4a | 7179 | 0.026 | 0.3287 | No | ||
| 69 | NDUFB3 | Ndufb3 | 7356 | 0.007 | 0.3174 | No | ||
| 70 | NDUFA8 | Ndufa8 | 7412 | 0.002 | 0.3139 | No | ||
| 71 | MT-ATP6 | mt-Atp6 | 7555 | 0.000 | 0.3047 | No | ||
| 72 | MT-CO3 | mt-Co3 | 7729 | 0.000 | 0.2935 | No | ||
| 73 | COX17 | Cox17 | 7803 | 0.000 | 0.2888 | No | ||
| 74 | MT-ND4L | mt-Nd4l | 7879 | 0.000 | 0.2839 | No | ||
| 75 | NDUFC1 | Ndufc1 | 7937 | -0.003 | 0.2802 | No | ||
| 76 | NDUFS4 | Ndufs4 | 8020 | -0.010 | 0.2751 | No | ||
| 77 | ATP5F1E | Atp5e | 8023 | -0.010 | 0.2752 | No | ||
| 78 | UQCRB | Uqcrb | 8025 | -0.011 | 0.2753 | No | ||
| 79 | NDUFA4 | Ndufa4 | 8027 | -0.011 | 0.2755 | No | ||
| 80 | ATP5MF | Atp5j2 | 8110 | -0.019 | 0.2705 | No | ||
| 81 | ATP5PD | Atp5h | 8167 | -0.025 | 0.2673 | No | ||
| 82 | NDUFB10 | Ndufb10 | 8329 | -0.039 | 0.2576 | No | ||
| 83 | NDUFS2 | Ndufs2 | 8687 | -0.073 | 0.2358 | No | ||
| 84 | NDUFS1 | Ndufs1 | 8719 | -0.076 | 0.2353 | No | ||
| 85 | COX4I2 | Cox4i2 | 8807 | -0.084 | 0.2312 | No | ||
| 86 | COX6C | Cox6c | 8935 | -0.096 | 0.2248 | No | ||
| 87 | ATP5F1D | Atp5d | 9134 | -0.118 | 0.2141 | No | ||
| 88 | COX6B1 | Cox6b1 | 9141 | -0.119 | 0.2160 | No | ||
| 89 | ATP5PF | Atp5j | 9296 | -0.134 | 0.2085 | No | ||
| 90 | ATP5F1B | Atp5b | 9370 | -0.141 | 0.2064 | No | ||
| 91 | NDUFA3 | Ndufa3 | 9616 | -0.166 | 0.1937 | No | ||
| 92 | PPA2 | Ppa2 | 9704 | -0.174 | 0.1913 | No | ||
| 93 | ATP5F1C | Atp5c1 | 9706 | -0.174 | 0.1945 | No | ||
| 94 | NDUFV3 | Ndufv3 | 9809 | -0.184 | 0.1914 | No | ||
| 95 | ATP5MC3 | Atp5g3 | 10054 | -0.209 | 0.1795 | No | ||
| 96 | ATP4B | Atp4b | 10066 | -0.211 | 0.1827 | No | ||
| 97 | NDUFV2 | Ndufv2 | 10193 | -0.222 | 0.1787 | No | ||
| 98 | UQCRC2 | Uqcrc2 | 10364 | -0.240 | 0.1722 | No | ||
| 99 | ATP5ME | Atp5k | 10627 | -0.267 | 0.1603 | No | ||
| 100 | COX7A2 | Cox7a2 | 10673 | -0.273 | 0.1625 | No | ||
| 101 | UQCRC1 | Uqcrc1 | 10805 | -0.287 | 0.1594 | No | ||
| 102 | NDUFB2 | Ndufb2 | 10880 | -0.294 | 0.1601 | No | ||
| 103 | ATP5F1A | Atp5a1 | 11636 | -0.380 | 0.1183 | No | ||
| 104 | NDUFAB1 | Ndufab1 | 11683 | -0.388 | 0.1226 | No | ||
| 105 | ATP12A | Atp12a | 11954 | -0.418 | 0.1130 | No | ||
| 106 | UQCRFS1 | Uqcrfs1 | 12508 | -0.475 | 0.0860 | No | ||
| 107 | COX7B | Cox7b | 12640 | -0.494 | 0.0868 | No | ||
| 108 | COX6B2 | Cox6b2 | 14123 | -0.857 | 0.0068 | No | ||
| 109 | ATP6V1B1 | Atp6v1b1 | 15237 | -1.959 | -0.0285 | No | ||
| 110 | ATP6V0D2 | Atp6v0d2 | 15389 | -2.504 | 0.0088 | No |