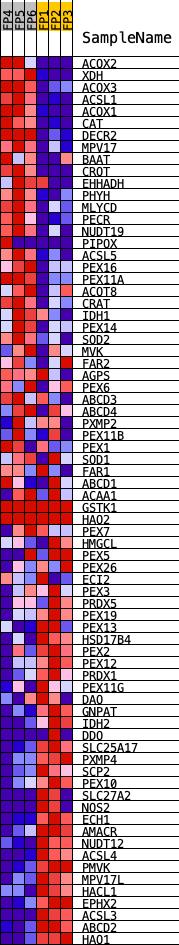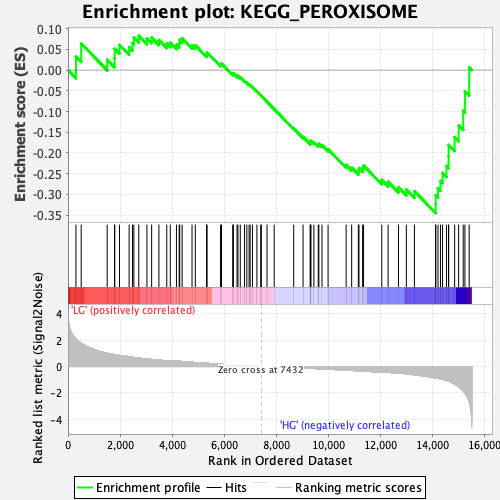
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
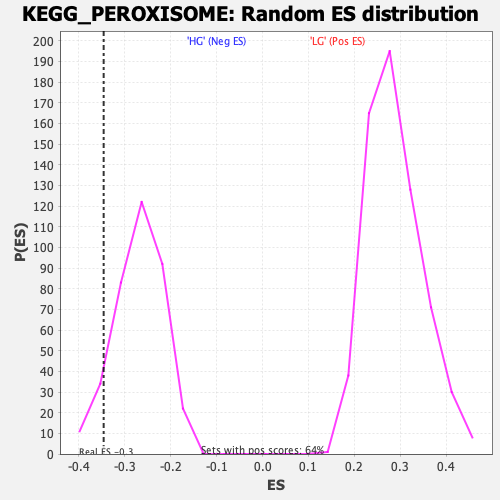
| GeneSet | KEGG_PEROXISOME |
| Enrichment Score (ES) | -0.34536755 |
| Normalized Enrichment Score (NES) | -1.2858629 |
| Nominal p-value | 0.07967033 |
| FDR q-value | 0.38332382 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ACOX2 | Acox2 | 306 | 2.076 | 0.0326 | No | ||
| 2 | XDH | Xdh | 507 | 1.760 | 0.0640 | No | ||
| 3 | ACOX3 | Acox3 | 1506 | 1.003 | 0.0247 | No | ||
| 4 | ACSL1 | Acsl1 | 1790 | 0.899 | 0.0290 | No | ||
| 5 | ACOX1 | Acox1 | 1797 | 0.895 | 0.0512 | No | ||
| 6 | CAT | Cat | 1979 | 0.839 | 0.0607 | No | ||
| 7 | DECR2 | Decr2 | 2350 | 0.740 | 0.0554 | No | ||
| 8 | MPV17 | Mpv17 | 2479 | 0.701 | 0.0648 | No | ||
| 9 | BAAT | Baat | 2532 | 0.683 | 0.0786 | No | ||
| 10 | CROT | Crot | 2721 | 0.637 | 0.0825 | No | ||
| 11 | EHHADH | Ehhadh | 3035 | 0.564 | 0.0765 | No | ||
| 12 | PHYH | Phyh | 3216 | 0.532 | 0.0783 | No | ||
| 13 | MLYCD | Mlycd | 3495 | 0.486 | 0.0725 | No | ||
| 14 | PECR | Pecr | 3797 | 0.434 | 0.0640 | No | ||
| 15 | NUDT19 | Nudt19 | 3932 | 0.424 | 0.0660 | No | ||
| 16 | PIPOX | Pipox | 4169 | 0.407 | 0.0610 | No | ||
| 17 | ACSL5 | Acsl5 | 4274 | 0.398 | 0.0643 | No | ||
| 18 | PEX16 | Pex16 | 4296 | 0.395 | 0.0729 | No | ||
| 19 | PEX11A | Pex11a | 4388 | 0.382 | 0.0767 | No | ||
| 20 | ACOT8 | Acot8 | 4768 | 0.317 | 0.0601 | No | ||
| 21 | CRAT | Crat | 4894 | 0.298 | 0.0596 | No | ||
| 22 | IDH1 | Idh1 | 5327 | 0.245 | 0.0378 | No | ||
| 23 | PEX14 | Pex14 | 5347 | 0.243 | 0.0427 | No | ||
| 24 | SOD2 | Sod2 | 5865 | 0.173 | 0.0136 | No | ||
| 25 | MVK | Mvk | 5901 | 0.168 | 0.0155 | No | ||
| 26 | FAR2 | Far2 | 6332 | 0.120 | -0.0093 | No | ||
| 27 | AGPS | Agps | 6360 | 0.117 | -0.0081 | No | ||
| 28 | PEX6 | Pex6 | 6499 | 0.101 | -0.0145 | No | ||
| 29 | ABCD3 | Abcd3 | 6534 | 0.097 | -0.0142 | No | ||
| 30 | ABCD4 | Abcd4 | 6623 | 0.087 | -0.0177 | No | ||
| 31 | PXMP2 | Pxmp2 | 6787 | 0.068 | -0.0266 | No | ||
| 32 | PEX11B | Pex11b | 6875 | 0.059 | -0.0307 | No | ||
| 33 | PEX1 | Pex1 | 6962 | 0.050 | -0.0350 | No | ||
| 34 | SOD1 | Sod1 | 7004 | 0.046 | -0.0365 | No | ||
| 35 | FAR1 | Far1 | 7092 | 0.035 | -0.0412 | No | ||
| 36 | ABCD1 | Abcd1 | 7263 | 0.018 | -0.0518 | No | ||
| 37 | ACAA1 | Acaa1a | 7409 | 0.003 | -0.0611 | No | ||
| 38 | GSTK1 | Gstk1 | 7432 | 0.000 | -0.0625 | No | ||
| 39 | HAO2 | Hao2 | 7652 | 0.000 | -0.0767 | No | ||
| 40 | PEX7 | Pex7 | 7927 | -0.002 | -0.0944 | No | ||
| 41 | HMGCL | Hmgcl | 8678 | -0.072 | -0.1411 | No | ||
| 42 | PEX5 | Pex5 | 9037 | -0.109 | -0.1615 | No | ||
| 43 | PEX26 | Pex26 | 9313 | -0.136 | -0.1759 | No | ||
| 44 | ECI2 | Eci2 | 9318 | -0.136 | -0.1727 | No | ||
| 45 | PEX3 | Pex3 | 9342 | -0.138 | -0.1707 | No | ||
| 46 | PRDX5 | Prdx5 | 9453 | -0.148 | -0.1741 | No | ||
| 47 | PEX19 | Pex19 | 9621 | -0.166 | -0.1807 | No | ||
| 48 | PEX13 | Pex13 | 9645 | -0.169 | -0.1779 | No | ||
| 49 | HSD17B4 | Hsd17b4 | 9765 | -0.180 | -0.1811 | No | ||
| 50 | PEX2 | Pex2 | 9999 | -0.203 | -0.1911 | No | ||
| 51 | PEX12 | Pex12 | 10693 | -0.275 | -0.2290 | No | ||
| 52 | PRDX1 | Prdx1 | 10906 | -0.296 | -0.2352 | No | ||
| 53 | PEX11G | Pex11g | 11162 | -0.326 | -0.2435 | No | ||
| 54 | DAO | Dao | 11187 | -0.329 | -0.2368 | No | ||
| 55 | GNPAT | Gnpat | 11327 | -0.344 | -0.2371 | No | ||
| 56 | IDH2 | Idh2 | 11364 | -0.349 | -0.2306 | No | ||
| 57 | DDO | Ddo | 12063 | -0.424 | -0.2651 | No | ||
| 58 | SLC25A17 | Slc25a17 | 12306 | -0.442 | -0.2696 | No | ||
| 59 | PXMP4 | Pxmp4 | 12707 | -0.502 | -0.2829 | No | ||
| 60 | SCP2 | Scp2 | 13008 | -0.554 | -0.2883 | No | ||
| 61 | PEX10 | Pex10 | 13319 | -0.624 | -0.2926 | No | ||
| 62 | SLC27A2 | Slc27a2 | 14135 | -0.860 | -0.3237 | Yes | ||
| 63 | NOS2 | Nos2 | 14138 | -0.860 | -0.3021 | Yes | ||
| 64 | ECH1 | Ech1 | 14225 | -0.892 | -0.2852 | Yes | ||
| 65 | AMACR | Amacr | 14320 | -0.934 | -0.2677 | Yes | ||
| 66 | NUDT12 | Nudt12 | 14400 | -0.971 | -0.2483 | Yes | ||
| 67 | ACSL4 | Acsl4 | 14550 | -1.055 | -0.2314 | Yes | ||
| 68 | PMVK | Pmvk | 14631 | -1.107 | -0.2087 | Yes | ||
| 69 | MPV17L | Mpv17l | 14635 | -1.111 | -0.1808 | Yes | ||
| 70 | HACL1 | Hacl1 | 14865 | -1.331 | -0.1621 | Yes | ||
| 71 | EPHX2 | Ephx2 | 15019 | -1.529 | -0.1334 | Yes | ||
| 72 | ACSL3 | Acsl3 | 15194 | -1.845 | -0.0981 | Yes | ||
| 73 | ABCD2 | Abcd2 | 15260 | -2.018 | -0.0515 | Yes | ||
| 74 | HAO1 | Hao1 | 15424 | -2.718 | 0.0065 | Yes |