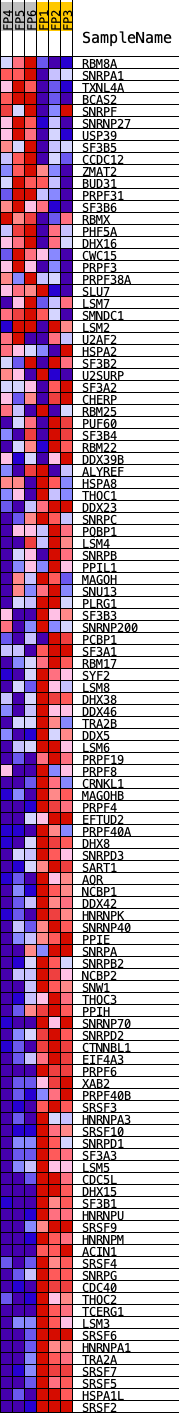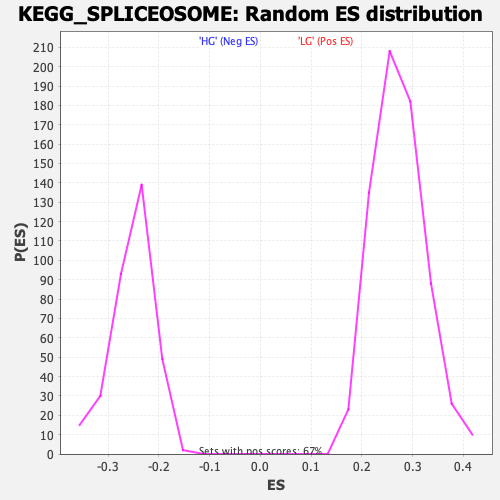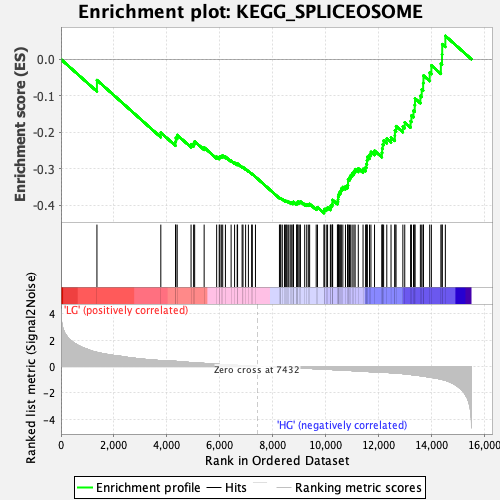
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_SPLICEOSOME |
| Enrichment Score (ES) | -0.42162797 |
| Normalized Enrichment Score (NES) | -1.6674242 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14665551 |
| FWER p-Value | 0.323 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RBM8A | Rbm8a | 1359 | 1.070 | -0.0571 | No | ||
| 2 | SNRPA1 | Snrpa1 | 3775 | 0.440 | -0.2010 | No | ||
| 3 | TXNL4A | Txnl4a | 4331 | 0.390 | -0.2257 | No | ||
| 4 | BCAS2 | Bcas2 | 4344 | 0.387 | -0.2152 | No | ||
| 5 | SNRPF | Snrpf | 4395 | 0.380 | -0.2074 | No | ||
| 6 | SNRNP27 | Snrnp27 | 4919 | 0.296 | -0.2328 | No | ||
| 7 | USP39 | Usp39 | 5016 | 0.285 | -0.2307 | No | ||
| 8 | SF3B5 | Sf3b5 | 5055 | 0.280 | -0.2251 | No | ||
| 9 | CCDC12 | Ccdc12 | 5414 | 0.232 | -0.2415 | No | ||
| 10 | ZMAT2 | Zmat2 | 5887 | 0.169 | -0.2673 | No | ||
| 11 | BUD31 | Bud31 | 5984 | 0.158 | -0.2689 | No | ||
| 12 | PRPF31 | Prpf31 | 6008 | 0.156 | -0.2659 | No | ||
| 13 | SF3B6 | Sf3b6 | 6063 | 0.149 | -0.2651 | No | ||
| 14 | RBMX | Rbmx | 6116 | 0.144 | -0.2643 | No | ||
| 15 | PHF5A | Phf5a | 6219 | 0.133 | -0.2670 | No | ||
| 16 | DHX16 | Dhx16 | 6434 | 0.108 | -0.2778 | No | ||
| 17 | CWC15 | Cwc15 | 6565 | 0.093 | -0.2835 | No | ||
| 18 | PRPF3 | Prpf3 | 6664 | 0.082 | -0.2875 | No | ||
| 19 | PRPF38A | Prpf38a | 6678 | 0.080 | -0.2860 | No | ||
| 20 | SLU7 | Slu7 | 6850 | 0.062 | -0.2953 | No | ||
| 21 | LSM7 | Lsm7 | 6883 | 0.058 | -0.2957 | No | ||
| 22 | SMNDC1 | Smndc1 | 6982 | 0.048 | -0.3006 | No | ||
| 23 | LSM2 | Lsm2 | 7088 | 0.036 | -0.3064 | No | ||
| 24 | U2AF2 | U2af2 | 7211 | 0.022 | -0.3137 | No | ||
| 25 | HSPA2 | Hspa2 | 7235 | 0.019 | -0.3146 | No | ||
| 26 | SF3B2 | Sf3b2 | 7355 | 0.007 | -0.3221 | No | ||
| 27 | U2SURP | U2surp | 8262 | -0.032 | -0.3800 | No | ||
| 28 | SF3A2 | Sf3a2 | 8292 | -0.035 | -0.3809 | No | ||
| 29 | CHERP | Cherp | 8314 | -0.037 | -0.3812 | No | ||
| 30 | RBM25 | Rbm25 | 8369 | -0.043 | -0.3834 | No | ||
| 31 | PUF60 | Puf60 | 8452 | -0.049 | -0.3873 | No | ||
| 32 | SF3B4 | Sf3b4 | 8485 | -0.052 | -0.3879 | No | ||
| 33 | RBM22 | Rbm22 | 8522 | -0.055 | -0.3886 | No | ||
| 34 | DDX39B | Ddx39b | 8554 | -0.059 | -0.3889 | No | ||
| 35 | ALYREF | 4931428L18Rik | 8612 | -0.066 | -0.3907 | No | ||
| 36 | HSPA8 | Hspa8 | 8667 | -0.071 | -0.3921 | No | ||
| 37 | THOC1 | Thoc1 | 8709 | -0.075 | -0.3926 | No | ||
| 38 | DDX23 | Ddx23 | 8774 | -0.081 | -0.3944 | No | ||
| 39 | SNRPC | Snrpc | 8784 | -0.082 | -0.3926 | No | ||
| 40 | PQBP1 | Pqbp1 | 8792 | -0.083 | -0.3906 | No | ||
| 41 | LSM4 | Lsm4 | 8913 | -0.093 | -0.3957 | No | ||
| 42 | SNRPB | Snrpb | 8924 | -0.094 | -0.3936 | No | ||
| 43 | PPIL1 | Ppil1 | 8961 | -0.099 | -0.3931 | No | ||
| 44 | MAGOH | Magoh | 8962 | -0.099 | -0.3902 | No | ||
| 45 | SNU13 | Snu13 | 9032 | -0.108 | -0.3915 | No | ||
| 46 | PLRG1 | Plrg1 | 9055 | -0.110 | -0.3898 | No | ||
| 47 | SF3B3 | Sf3b3 | 9217 | -0.126 | -0.3965 | No | ||
| 48 | SNRNP200 | Snrnp200 | 9294 | -0.134 | -0.3976 | No | ||
| 49 | PCBP1 | Pcbp1 | 9354 | -0.139 | -0.3974 | No | ||
| 50 | SF3A1 | Sf3a1 | 9404 | -0.144 | -0.3964 | No | ||
| 51 | RBM17 | Rbm17 | 9652 | -0.170 | -0.4075 | No | ||
| 52 | SYF2 | Syf2 | 9700 | -0.174 | -0.4055 | No | ||
| 53 | LSM8 | Lsm8 | 9950 | -0.198 | -0.4159 | Yes | ||
| 54 | DHX38 | Dhx38 | 9955 | -0.199 | -0.4104 | Yes | ||
| 55 | DDX46 | Ddx46 | 10031 | -0.207 | -0.4092 | Yes | ||
| 56 | TRA2B | Tra2b | 10082 | -0.212 | -0.4063 | Yes | ||
| 57 | DDX5 | Ddx5 | 10188 | -0.222 | -0.4067 | Yes | ||
| 58 | LSM6 | Lsm6 | 10204 | -0.223 | -0.4012 | Yes | ||
| 59 | PRPF19 | Prpf19 | 10252 | -0.229 | -0.3976 | Yes | ||
| 60 | PRPF8 | Prpf8 | 10271 | -0.230 | -0.3921 | Yes | ||
| 61 | CRNKL1 | Crnkl1 | 10272 | -0.231 | -0.3854 | Yes | ||
| 62 | MAGOHB | Magohb | 10454 | -0.250 | -0.3899 | Yes | ||
| 63 | PRPF4 | Prpf4 | 10477 | -0.252 | -0.3840 | Yes | ||
| 64 | EFTUD2 | Eftud2 | 10483 | -0.253 | -0.3770 | Yes | ||
| 65 | PRPF40A | Prpf40a | 10498 | -0.254 | -0.3705 | Yes | ||
| 66 | DHX8 | Dhx8 | 10528 | -0.257 | -0.3649 | Yes | ||
| 67 | SNRPD3 | Snrpd3 | 10578 | -0.263 | -0.3605 | Yes | ||
| 68 | SART1 | Sart1 | 10602 | -0.265 | -0.3543 | Yes | ||
| 69 | AQR | Aqr | 10660 | -0.272 | -0.3501 | Yes | ||
| 70 | NCBP1 | Ncbp1 | 10759 | -0.283 | -0.3482 | Yes | ||
| 71 | DDX42 | Ddx42 | 10838 | -0.290 | -0.3449 | Yes | ||
| 72 | HNRNPK | Hnrnpk | 10856 | -0.291 | -0.3375 | Yes | ||
| 73 | SNRNP40 | Snrnp40 | 10857 | -0.291 | -0.3291 | Yes | ||
| 74 | PPIE | Ppie | 10912 | -0.296 | -0.3240 | Yes | ||
| 75 | SNRPA | Snrpa | 10956 | -0.302 | -0.3180 | Yes | ||
| 76 | SNRPB2 | Snrpb2 | 11011 | -0.308 | -0.3125 | Yes | ||
| 77 | NCBP2 | Ncbp2 | 11078 | -0.317 | -0.3076 | Yes | ||
| 78 | SNW1 | Snw1 | 11124 | -0.322 | -0.3012 | Yes | ||
| 79 | THOC3 | Thoc3 | 11244 | -0.335 | -0.2992 | Yes | ||
| 80 | PPIH | Gm7879 | 11424 | -0.356 | -0.3004 | Yes | ||
| 81 | SNRNP70 | Snrnp70 | 11521 | -0.367 | -0.2960 | Yes | ||
| 82 | SNRPD2 | Snrpd2 | 11549 | -0.370 | -0.2870 | Yes | ||
| 83 | CTNNBL1 | Ctnnbl1 | 11572 | -0.373 | -0.2776 | Yes | ||
| 84 | EIF4A3 | Eif4a3 | 11584 | -0.374 | -0.2675 | Yes | ||
| 85 | PRPF6 | Prpf6 | 11669 | -0.385 | -0.2618 | Yes | ||
| 86 | XAB2 | Xab2 | 11722 | -0.392 | -0.2538 | Yes | ||
| 87 | PRPF40B | Prpf40b | 11858 | -0.410 | -0.2506 | Yes | ||
| 88 | SRSF3 | Srsf3 | 12136 | -0.425 | -0.2563 | Yes | ||
| 89 | HNRNPA3 | Gm17190 | 12147 | -0.428 | -0.2445 | Yes | ||
| 90 | SRSF10 | Srsf10 | 12168 | -0.430 | -0.2333 | Yes | ||
| 91 | SNRPD1 | Snrpd1 | 12202 | -0.435 | -0.2228 | Yes | ||
| 92 | SF3A3 | Sf3a3 | 12322 | -0.445 | -0.2176 | Yes | ||
| 93 | LSM5 | Lsm5 | 12482 | -0.471 | -0.2143 | Yes | ||
| 94 | CDC5L | Cdc5l | 12626 | -0.492 | -0.2093 | Yes | ||
| 95 | DHX15 | Dhx15 | 12630 | -0.492 | -0.1952 | Yes | ||
| 96 | SF3B1 | Sf3b1 | 12672 | -0.499 | -0.1833 | Yes | ||
| 97 | HNRNPU | Hnrnpu | 12930 | -0.537 | -0.1844 | Yes | ||
| 98 | SRSF9 | Srsf9 | 13001 | -0.553 | -0.1729 | Yes | ||
| 99 | HNRNPM | Hnrnpm | 13222 | -0.602 | -0.1697 | Yes | ||
| 100 | ACIN1 | Acin1 | 13254 | -0.611 | -0.1540 | Yes | ||
| 101 | SRSF4 | Srsf4 | 13332 | -0.627 | -0.1408 | Yes | ||
| 102 | SNRPG | Snrpg | 13379 | -0.638 | -0.1252 | Yes | ||
| 103 | CDC40 | Cdc40 | 13389 | -0.640 | -0.1072 | Yes | ||
| 104 | THOC2 | Thoc2 | 13592 | -0.695 | -0.1002 | Yes | ||
| 105 | TCERG1 | Tcerg1 | 13644 | -0.710 | -0.0829 | Yes | ||
| 106 | LSM3 | Lsm3 | 13697 | -0.727 | -0.0651 | Yes | ||
| 107 | SRSF6 | Srsf6 | 13709 | -0.731 | -0.0446 | Yes | ||
| 108 | HNRNPA1 | Hnrnpa1 | 13945 | -0.801 | -0.0366 | Yes | ||
| 109 | TRA2A | Tra2a | 14008 | -0.823 | -0.0167 | Yes | ||
| 110 | SRSF7 | Srsf7 | 14368 | -0.957 | -0.0122 | Yes | ||
| 111 | SRSF5 | Srsf5 | 14411 | -0.976 | 0.0134 | Yes | ||
| 112 | HSPA1L | Hspa1l | 14418 | -0.978 | 0.0414 | Yes | ||
| 113 | SRSF2 | Srsf2 | 14537 | -1.046 | 0.0641 | Yes |