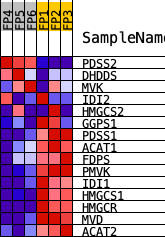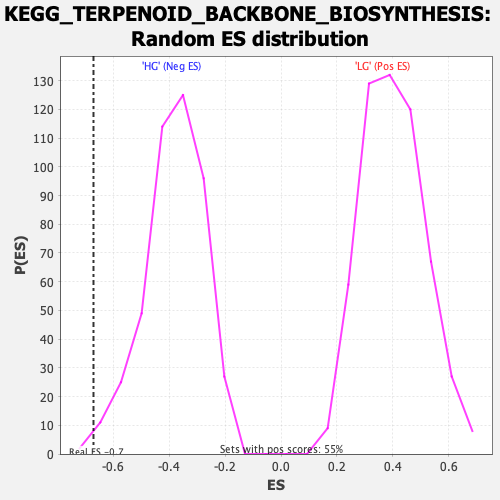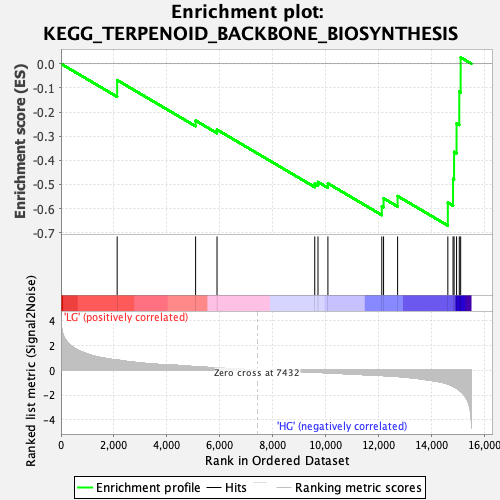
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_TERPENOID_BACKBONE_BIOSYNTHESIS |
| Enrichment Score (ES) | -0.6708528 |
| Normalized Enrichment Score (NES) | -1.7529922 |
| Nominal p-value | 0.004454343 |
| FDR q-value | 0.20118196 |
| FWER p-Value | 0.164 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDSS2 | Pdss2 | 2124 | 0.805 | -0.0678 | No | ||
| 2 | DHDDS | Dhdds | 5091 | 0.274 | -0.2354 | No | ||
| 3 | MVK | Mvk | 5901 | 0.168 | -0.2732 | No | ||
| 4 | IDI2 | Gm9745 | 9597 | -0.163 | -0.4973 | No | ||
| 5 | HMGCS2 | Hmgcs2 | 9720 | -0.175 | -0.4901 | No | ||
| 6 | GGPS1 | Ggps1 | 10095 | -0.213 | -0.4959 | No | ||
| 7 | PDSS1 | Pdss1 | 12133 | -0.425 | -0.5908 | Yes | ||
| 8 | ACAT1 | Acat1 | 12200 | -0.435 | -0.5577 | Yes | ||
| 9 | FDPS | Fdps | 12731 | -0.505 | -0.5484 | Yes | ||
| 10 | PMVK | Pmvk | 14631 | -1.107 | -0.5757 | Yes | ||
| 11 | IDI1 | Idi1 | 14828 | -1.287 | -0.4777 | Yes | ||
| 12 | HMGCS1 | Hmgcs1 | 14867 | -1.334 | -0.3656 | Yes | ||
| 13 | HMGCR | Hmgcr | 14961 | -1.447 | -0.2472 | Yes | ||
| 14 | MVD | Mvd | 15064 | -1.608 | -0.1156 | Yes | ||
| 15 | ACAT2 | Acat2 | 15116 | -1.691 | 0.0264 | Yes |