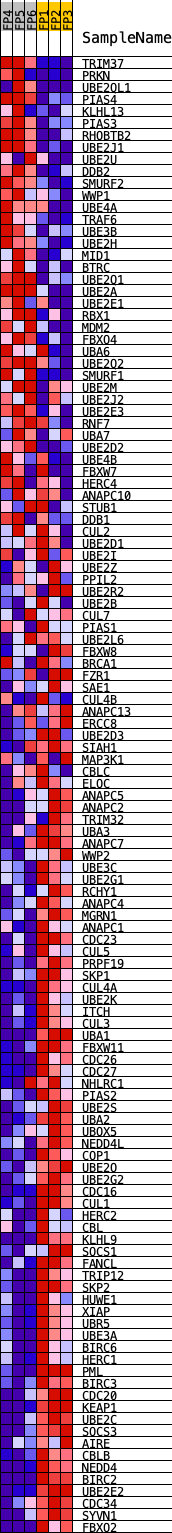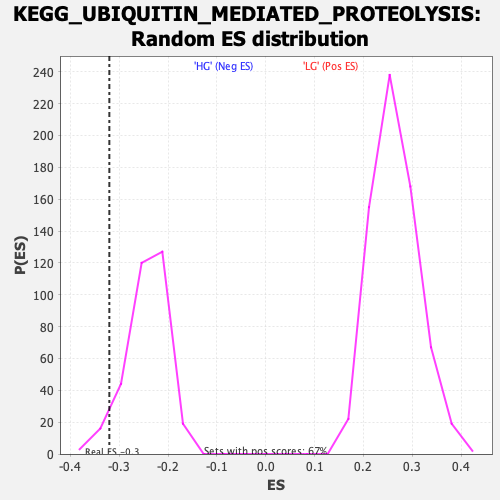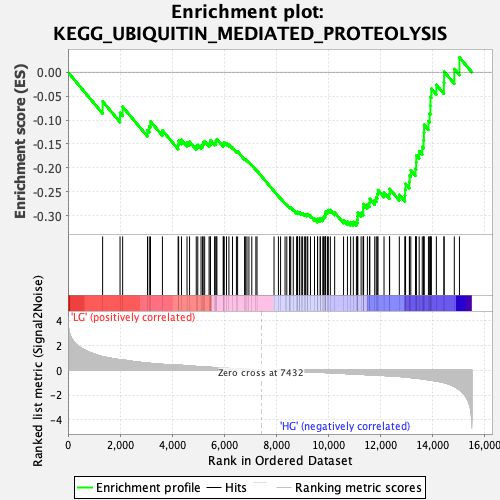
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_UBIQUITIN_MEDIATED_PROTEOLYSIS |
| Enrichment Score (ES) | -0.3204281 |
| Normalized Enrichment Score (NES) | -1.3184258 |
| Nominal p-value | 0.051671732 |
| FDR q-value | 0.36910596 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TRIM37 | Trim37 | 1331 | 1.081 | -0.0610 | No | ||
| 2 | PRKN | Prkn | 2001 | 0.835 | -0.0849 | No | ||
| 3 | UBE2QL1 | Ube2ql1 | 2101 | 0.810 | -0.0723 | No | ||
| 4 | PIAS4 | Pias4 | 3056 | 0.561 | -0.1211 | No | ||
| 5 | KLHL13 | Klhl13 | 3124 | 0.548 | -0.1126 | No | ||
| 6 | PIAS3 | Pias3 | 3172 | 0.540 | -0.1029 | No | ||
| 7 | RHOBTB2 | Rhobtb2 | 3630 | 0.465 | -0.1217 | No | ||
| 8 | UBE2J1 | Ube2j1 | 4234 | 0.404 | -0.1514 | No | ||
| 9 | UBE2U | Ube2u | 4254 | 0.402 | -0.1431 | No | ||
| 10 | DDB2 | Ddb2 | 4363 | 0.385 | -0.1411 | No | ||
| 11 | SMURF2 | Smurf2 | 4576 | 0.347 | -0.1467 | No | ||
| 12 | WWP1 | Wwp1 | 4673 | 0.331 | -0.1452 | No | ||
| 13 | UBE4A | Ube4a | 4931 | 0.294 | -0.1550 | No | ||
| 14 | TRAF6 | Traf6 | 4989 | 0.288 | -0.1519 | No | ||
| 15 | UBE3B | Ube3b | 5115 | 0.272 | -0.1537 | No | ||
| 16 | UBE2H | Ube2h | 5175 | 0.267 | -0.1512 | No | ||
| 17 | MID1 | Mid1 | 5197 | 0.263 | -0.1464 | No | ||
| 18 | BTRC | Btrc | 5255 | 0.254 | -0.1442 | No | ||
| 19 | UBE2Q1 | Ube2q1 | 5424 | 0.231 | -0.1496 | No | ||
| 20 | UBE2A | Ube2a | 5470 | 0.224 | -0.1473 | No | ||
| 21 | UBE2E1 | Ube2e1 | 5476 | 0.223 | -0.1424 | No | ||
| 22 | RBX1 | Rbx1 | 5635 | 0.202 | -0.1479 | No | ||
| 23 | MDM2 | Mdm2 | 5688 | 0.197 | -0.1466 | No | ||
| 24 | FBXO4 | Fbxo4 | 5690 | 0.197 | -0.1421 | No | ||
| 25 | UBA6 | Uba6 | 5725 | 0.193 | -0.1398 | No | ||
| 26 | UBE2Q2 | Ube2q2 | 5967 | 0.161 | -0.1516 | No | ||
| 27 | SMURF1 | Smurf1 | 5977 | 0.159 | -0.1485 | No | ||
| 28 | UBE2M | Ube2m | 6007 | 0.156 | -0.1467 | No | ||
| 29 | UBE2J2 | Ube2j2 | 6093 | 0.146 | -0.1488 | No | ||
| 30 | UBE2E3 | Ube2e3 | 6187 | 0.136 | -0.1516 | No | ||
| 31 | RNF7 | Gm7075 | 6324 | 0.121 | -0.1576 | No | ||
| 32 | UBA7 | Uba7 | 6475 | 0.103 | -0.1649 | No | ||
| 33 | UBE2D2 | Ube2d2a | 6521 | 0.098 | -0.1656 | No | ||
| 34 | UBE4B | Ube4b | 6788 | 0.068 | -0.1812 | No | ||
| 35 | FBXW7 | Fbxw7 | 6831 | 0.063 | -0.1825 | No | ||
| 36 | HERC4 | Herc4 | 6881 | 0.058 | -0.1843 | No | ||
| 37 | ANAPC10 | Anapc10 | 6956 | 0.051 | -0.1879 | No | ||
| 38 | STUB1 | Stub1 | 7066 | 0.038 | -0.1941 | No | ||
| 39 | DDB1 | Ddb1 | 7223 | 0.020 | -0.2038 | No | ||
| 40 | CUL2 | Cul2 | 7267 | 0.018 | -0.2061 | No | ||
| 41 | UBE2D1 | Ube2d1 | 7921 | -0.001 | -0.2485 | No | ||
| 42 | UBE2I | Ube2i | 8090 | -0.017 | -0.2590 | No | ||
| 43 | UBE2Z | Ube2z | 8173 | -0.026 | -0.2637 | No | ||
| 44 | PPIL2 | Ppil2 | 8345 | -0.040 | -0.2739 | No | ||
| 45 | UBE2R2 | Ube2r2 | 8412 | -0.045 | -0.2771 | No | ||
| 46 | UBE2B | Ube2b | 8521 | -0.055 | -0.2828 | No | ||
| 47 | CUL7 | Cul7 | 8529 | -0.057 | -0.2819 | No | ||
| 48 | PIAS1 | Pias1 | 8564 | -0.061 | -0.2827 | No | ||
| 49 | UBE2L6 | Ube2l6 | 8661 | -0.070 | -0.2873 | No | ||
| 50 | FBXW8 | Fbxw8 | 8789 | -0.083 | -0.2936 | No | ||
| 51 | BRCA1 | Brca1 | 8820 | -0.085 | -0.2936 | No | ||
| 52 | FZR1 | Fzr1 | 8822 | -0.085 | -0.2916 | No | ||
| 53 | SAE1 | Sae1 | 8894 | -0.091 | -0.2941 | No | ||
| 54 | CUL4B | Cul4b | 8910 | -0.093 | -0.2929 | No | ||
| 55 | ANAPC13 | Anapc13 | 8987 | -0.102 | -0.2954 | No | ||
| 56 | ERCC8 | Ercc8 | 9021 | -0.107 | -0.2951 | No | ||
| 57 | UBE2D3 | Ube2d3 | 9098 | -0.115 | -0.2973 | No | ||
| 58 | SIAH1 | Siah1a | 9135 | -0.118 | -0.2969 | No | ||
| 59 | MAP3K1 | Map3k1 | 9194 | -0.124 | -0.2977 | No | ||
| 60 | CBLC | Cblc | 9219 | -0.127 | -0.2963 | No | ||
| 61 | ELOC | Eloc | 9322 | -0.136 | -0.2997 | No | ||
| 62 | ANAPC5 | Anapc5 | 9477 | -0.151 | -0.3062 | No | ||
| 63 | ANAPC2 | Anapc2 | 9592 | -0.163 | -0.3098 | No | ||
| 64 | TRIM32 | Trim32 | 9598 | -0.164 | -0.3062 | No | ||
| 65 | UBA3 | Uba3 | 9692 | -0.174 | -0.3082 | No | ||
| 66 | ANAPC7 | Anapc7 | 9705 | -0.174 | -0.3049 | No | ||
| 67 | WWP2 | Wwp2 | 9797 | -0.183 | -0.3065 | No | ||
| 68 | UBE3C | Ube3c | 9819 | -0.185 | -0.3035 | No | ||
| 69 | UBE2G1 | Ube2g1 | 9844 | -0.188 | -0.3007 | No | ||
| 70 | RCHY1 | Rchy1 | 9884 | -0.191 | -0.2987 | No | ||
| 71 | ANAPC4 | Anapc4 | 9900 | -0.193 | -0.2952 | No | ||
| 72 | MGRN1 | Mgrn1 | 9906 | -0.194 | -0.2909 | No | ||
| 73 | ANAPC1 | Anapc1 | 9975 | -0.201 | -0.2906 | No | ||
| 74 | CDC23 | Cdc23 | 10009 | -0.204 | -0.2880 | No | ||
| 75 | CUL5 | Cul5 | 10086 | -0.213 | -0.2879 | No | ||
| 76 | PRPF19 | Prpf19 | 10252 | -0.229 | -0.2933 | No | ||
| 77 | SKP1 | Skp1a | 10594 | -0.265 | -0.3092 | No | ||
| 78 | CUL4A | Cul4a | 10743 | -0.281 | -0.3122 | No | ||
| 79 | UBE2K | Ube2k | 10867 | -0.292 | -0.3133 | Yes | ||
| 80 | ITCH | Itch | 10970 | -0.304 | -0.3128 | Yes | ||
| 81 | CUL3 | Cul3 | 11088 | -0.318 | -0.3130 | Yes | ||
| 82 | UBA1 | Uba1 | 11127 | -0.322 | -0.3079 | Yes | ||
| 83 | FBXW11 | Fbxw11 | 11134 | -0.323 | -0.3007 | Yes | ||
| 84 | CDC26 | Cdc26 | 11141 | -0.324 | -0.2935 | Yes | ||
| 85 | CDC27 | Cdc27 | 11269 | -0.337 | -0.2938 | Yes | ||
| 86 | NHLRC1 | Nhlrc1 | 11350 | -0.347 | -0.2908 | Yes | ||
| 87 | PIAS2 | Pias2 | 11351 | -0.348 | -0.2827 | Yes | ||
| 88 | UBE2S | Ube2s | 11365 | -0.349 | -0.2753 | Yes | ||
| 89 | UBA2 | Uba2 | 11510 | -0.366 | -0.2761 | Yes | ||
| 90 | UBOX5 | Ubox5 | 11590 | -0.374 | -0.2724 | Yes | ||
| 91 | NEDD4L | Nedd4l | 11605 | -0.376 | -0.2645 | Yes | ||
| 92 | COP1 | Cop1 | 11793 | -0.401 | -0.2673 | Yes | ||
| 93 | UBE2O | Ube2o | 11851 | -0.409 | -0.2614 | Yes | ||
| 94 | UBE2G2 | Ube2g2 | 11891 | -0.414 | -0.2542 | Yes | ||
| 95 | CDC16 | Cdc16 | 11919 | -0.418 | -0.2461 | Yes | ||
| 96 | CUL1 | Cul1 | 12150 | -0.428 | -0.2510 | Yes | ||
| 97 | HERC2 | Herc2 | 12359 | -0.450 | -0.2539 | Yes | ||
| 98 | CBL | Cbl | 12366 | -0.451 | -0.2437 | Yes | ||
| 99 | KLHL9 | Klhl9 | 12738 | -0.506 | -0.2559 | Yes | ||
| 100 | SOCS1 | Socs1 | 12954 | -0.541 | -0.2572 | Yes | ||
| 101 | FANCL | Fancl | 12963 | -0.542 | -0.2450 | Yes | ||
| 102 | TRIP12 | Trip12 | 12978 | -0.547 | -0.2331 | Yes | ||
| 103 | SKP2 | Skp2 | 13124 | -0.579 | -0.2289 | Yes | ||
| 104 | HUWE1 | Huwe1 | 13133 | -0.581 | -0.2158 | Yes | ||
| 105 | XIAP | Xiap | 13180 | -0.594 | -0.2048 | Yes | ||
| 106 | UBR5 | Ubr5 | 13360 | -0.633 | -0.2016 | Yes | ||
| 107 | UBE3A | Ube3a | 13385 | -0.639 | -0.1881 | Yes | ||
| 108 | BIRC6 | Birc6 | 13392 | -0.641 | -0.1735 | Yes | ||
| 109 | HERC1 | Herc1 | 13501 | -0.668 | -0.1648 | Yes | ||
| 110 | PML | Pml | 13619 | -0.702 | -0.1559 | Yes | ||
| 111 | BIRC3 | Birc3 | 13670 | -0.718 | -0.1423 | Yes | ||
| 112 | CDC20 | Cdc20 | 13683 | -0.722 | -0.1262 | Yes | ||
| 113 | KEAP1 | Keap1 | 13691 | -0.724 | -0.1096 | Yes | ||
| 114 | UBE2C | Ube2c | 13858 | -0.773 | -0.1022 | Yes | ||
| 115 | SOCS3 | Socs3 | 13904 | -0.790 | -0.0866 | Yes | ||
| 116 | AIRE | Aire | 13933 | -0.796 | -0.0697 | Yes | ||
| 117 | CBLB | Cblb | 13936 | -0.797 | -0.0511 | Yes | ||
| 118 | NEDD4 | Nedd4 | 13971 | -0.808 | -0.0344 | Yes | ||
| 119 | BIRC2 | Birc2 | 14162 | -0.865 | -0.0264 | Yes | ||
| 120 | UBE2E2 | Ube2e2 | 14451 | -0.995 | -0.0218 | Yes | ||
| 121 | CDC34 | Cdc34 | 14462 | -1.002 | 0.0011 | Yes | ||
| 122 | SYVN1 | Syvn1 | 14852 | -1.314 | 0.0067 | Yes | ||
| 123 | FBXO2 | Fbxo2 | 15050 | -1.574 | 0.0308 | Yes |