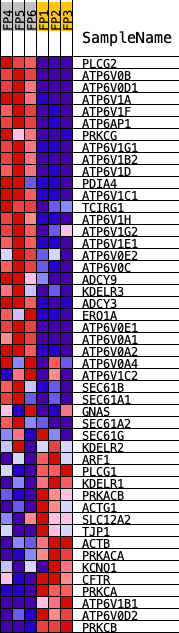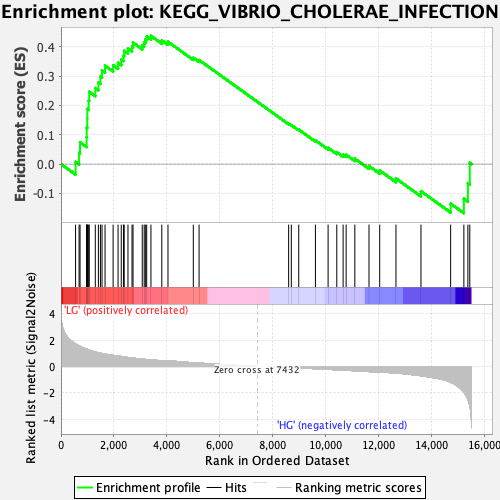
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#LG_versus_HG.phenotype.cls#LG_versus_HG_repos |
| Phenotype | phenotype.cls#LG_versus_HG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_VIBRIO_CHOLERAE_INFECTION |
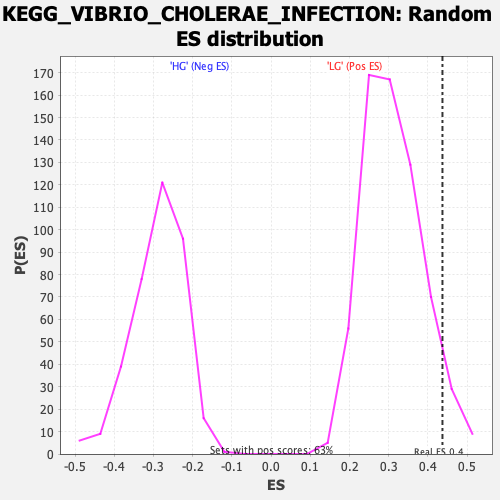
| Enrichment Score (ES) | 0.43690822 |
| Normalized Enrichment Score (NES) | 1.4064605 |
| Nominal p-value | 0.055205047 |
| FDR q-value | 0.26459995 |
| FWER p-Value | 0.996 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLCG2 | Plcg2 | 551 | 1.717 | 0.0074 | Yes | ||
| 2 | ATP6V0B | Atp6v0b | 684 | 1.564 | 0.0381 | Yes | ||
| 3 | ATP6V0D1 | Atp6v0d1 | 720 | 1.529 | 0.0741 | Yes | ||
| 4 | ATP6V1A | Atp6v1a | 968 | 1.319 | 0.0912 | Yes | ||
| 5 | ATP6V1F | Atp6v1f | 973 | 1.317 | 0.1240 | Yes | ||
| 6 | ATP6AP1 | Atp6ap1 | 994 | 1.304 | 0.1553 | Yes | ||
| 7 | PRKCG | Prkcg | 996 | 1.303 | 0.1879 | Yes | ||
| 8 | ATP6V1G1 | Atp6v1g1 | 1046 | 1.261 | 0.2164 | Yes | ||
| 9 | ATP6V1B2 | Atp6v1b2 | 1067 | 1.247 | 0.2463 | Yes | ||
| 10 | ATP6V1D | Atp6v1d | 1297 | 1.100 | 0.2591 | Yes | ||
| 11 | PDIA4 | Pdia4 | 1414 | 1.049 | 0.2779 | Yes | ||
| 12 | ATP6V1C1 | Atp6v1c1 | 1491 | 1.008 | 0.2982 | Yes | ||
| 13 | TCIRG1 | Tcirg1 | 1552 | 0.985 | 0.3190 | Yes | ||
| 14 | ATP6V1H | Atp6v1h | 1666 | 0.945 | 0.3354 | Yes | ||
| 15 | ATP6V1G2 | Atp6v1g2 | 1971 | 0.843 | 0.3369 | Yes | ||
| 16 | ATP6V1E1 | Atp6v1e1 | 2158 | 0.797 | 0.3448 | Yes | ||
| 17 | ATP6V0E2 | Atp6v0e2 | 2279 | 0.764 | 0.3562 | Yes | ||
| 18 | ATP6V0C | Atp6v0c | 2362 | 0.737 | 0.3694 | Yes | ||
| 19 | ADCY9 | Adcy9 | 2390 | 0.729 | 0.3859 | Yes | ||
| 20 | KDELR3 | Kdelr3 | 2533 | 0.683 | 0.3938 | Yes | ||
| 21 | ADCY3 | Adcy3 | 2687 | 0.646 | 0.4001 | Yes | ||
| 22 | ERO1A | Ero1l | 2724 | 0.637 | 0.4137 | Yes | ||
| 23 | ATP6V0E1 | Atp6v0e | 3073 | 0.557 | 0.4052 | Yes | ||
| 24 | ATP6V0A1 | Atp6v0a1 | 3146 | 0.543 | 0.4142 | Yes | ||
| 25 | ATP6V0A2 | Atp6v0a2 | 3190 | 0.536 | 0.4248 | Yes | ||
| 26 | ATP6V0A4 | Atp6v0a4 | 3241 | 0.528 | 0.4348 | Yes | ||
| 27 | ATP6V1C2 | Atp6v1c2 | 3402 | 0.497 | 0.4369 | Yes | ||
| 28 | SEC61B | Gm10320 | 3812 | 0.432 | 0.4213 | No | ||
| 29 | SEC61A1 | Sec61a1 | 4046 | 0.419 | 0.4167 | No | ||
| 30 | GNAS | Gnas | 5004 | 0.286 | 0.3621 | No | ||
| 31 | SEC61A2 | Sec61a2 | 5222 | 0.259 | 0.3545 | No | ||
| 32 | SEC61G | Sec61g | 8610 | -0.066 | 0.1374 | No | ||
| 33 | KDELR2 | Kdelr2 | 8711 | -0.075 | 0.1328 | No | ||
| 34 | ARF1 | Arf1 | 8992 | -0.103 | 0.1173 | No | ||
| 35 | PLCG1 | Plcg1 | 9622 | -0.167 | 0.0808 | No | ||
| 36 | KDELR1 | Kdelr1 | 10102 | -0.214 | 0.0552 | No | ||
| 37 | PRKACB | Prkacb | 10431 | -0.249 | 0.0403 | No | ||
| 38 | ACTG1 | Actg1 | 10670 | -0.272 | 0.0317 | No | ||
| 39 | SLC12A2 | Slc12a2 | 10784 | -0.285 | 0.0316 | No | ||
| 40 | TJP1 | Tjp1 | 11113 | -0.321 | 0.0184 | No | ||
| 41 | ACTB | Actb | 11649 | -0.382 | -0.0066 | No | ||
| 42 | PRKACA | Prkaca | 12054 | -0.424 | -0.0220 | No | ||
| 43 | KCNQ1 | Kcnq1 | 12669 | -0.499 | -0.0492 | No | ||
| 44 | CFTR | Cftr | 13614 | -0.701 | -0.0926 | No | ||
| 45 | PRKCA | Prkca | 14737 | -1.199 | -0.1351 | No | ||
| 46 | ATP6V1B1 | Atp6v1b1 | 15237 | -1.959 | -0.1182 | No | ||
| 47 | ATP6V0D2 | Atp6v0d2 | 15389 | -2.504 | -0.0653 | No | ||
| 48 | PRKCB | Prkcb | 15461 | -2.954 | 0.0041 | No |