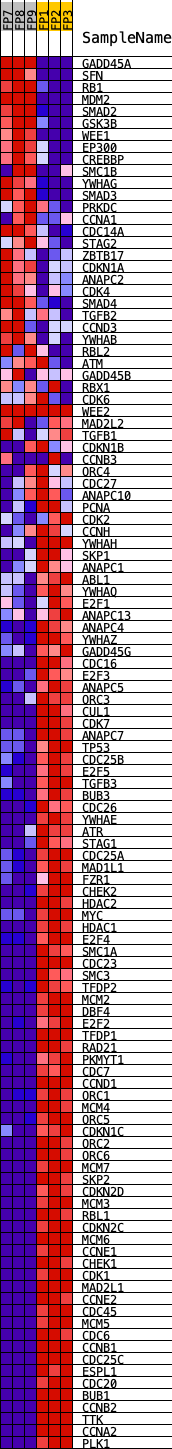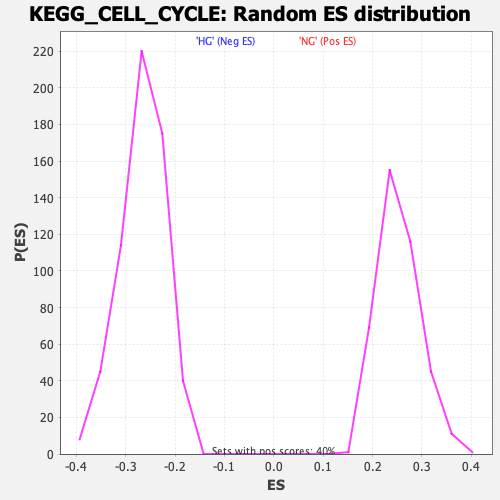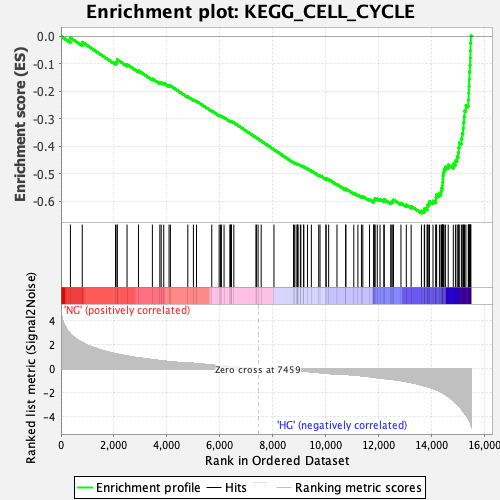
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.6438858 |
| Normalized Enrichment Score (NES) | -2.4225614 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GADD45A | Gadd45a | 356 | 2.893 | -0.0056 | No | ||
| 2 | SFN | Sfn | 804 | 2.179 | -0.0214 | No | ||
| 3 | RB1 | Rb1 | 2061 | 1.239 | -0.0953 | No | ||
| 4 | MDM2 | Mdm2 | 2120 | 1.210 | -0.0918 | No | ||
| 5 | SMAD2 | Smad2 | 2126 | 1.206 | -0.0848 | No | ||
| 6 | GSK3B | Gsk3b | 2498 | 1.045 | -0.1025 | No | ||
| 7 | WEE1 | Wee1 | 2932 | 0.886 | -0.1252 | No | ||
| 8 | EP300 | Ep300 | 3456 | 0.737 | -0.1547 | No | ||
| 9 | CREBBP | Crebbp | 3739 | 0.665 | -0.1690 | No | ||
| 10 | SMC1B | Smc1b | 3789 | 0.654 | -0.1682 | No | ||
| 11 | YWHAG | Ywhag | 3886 | 0.628 | -0.1706 | No | ||
| 12 | SMAD3 | Smad3 | 4087 | 0.576 | -0.1801 | No | ||
| 13 | PRKDC | Prkdc | 4138 | 0.562 | -0.1799 | No | ||
| 14 | CCNA1 | Ccna1 | 4796 | 0.445 | -0.2199 | No | ||
| 15 | CDC14A | Cdc14a | 5008 | 0.434 | -0.2309 | No | ||
| 16 | STAG2 | Stag2 | 5124 | 0.409 | -0.2359 | No | ||
| 17 | ZBTB17 | Zbtb17 | 5702 | 0.299 | -0.2716 | No | ||
| 18 | CDKN1A | Cdkn1a | 5982 | 0.248 | -0.2882 | No | ||
| 19 | ANAPC2 | Anapc2 | 6036 | 0.237 | -0.2902 | No | ||
| 20 | CDK4 | Cdk4 | 6067 | 0.233 | -0.2907 | No | ||
| 21 | SMAD4 | Smad4 | 6167 | 0.216 | -0.2958 | No | ||
| 22 | TGFB2 | Tgfb2 | 6387 | 0.174 | -0.3090 | No | ||
| 23 | CCND3 | Ccnd3 | 6410 | 0.169 | -0.3094 | No | ||
| 24 | YWHAB | Ywhab | 6436 | 0.165 | -0.3100 | No | ||
| 25 | RBL2 | Rbl2 | 6445 | 0.163 | -0.3095 | No | ||
| 26 | ATM | Atm | 6537 | 0.144 | -0.3146 | No | ||
| 27 | GADD45B | Gadd45b | 7375 | 0.013 | -0.3688 | No | ||
| 28 | RBX1 | Rbx1 | 7389 | 0.011 | -0.3696 | No | ||
| 29 | CDK6 | Cdk6 | 7454 | 0.001 | -0.3737 | No | ||
| 30 | WEE2 | Wee2 | 7571 | 0.000 | -0.3812 | No | ||
| 31 | MAD2L2 | Mad2l2 | 8054 | -0.005 | -0.4125 | No | ||
| 32 | TGFB1 | Tgfb1 | 8795 | -0.144 | -0.4596 | No | ||
| 33 | CDKN1B | Cdkn1b | 8825 | -0.147 | -0.4606 | No | ||
| 34 | CCNB3 | Ccnb3 | 8893 | -0.160 | -0.4640 | No | ||
| 35 | ORC4 | Orc4 | 8947 | -0.169 | -0.4664 | No | ||
| 36 | CDC27 | Cdc27 | 8952 | -0.170 | -0.4657 | No | ||
| 37 | ANAPC10 | Anapc10 | 8964 | -0.171 | -0.4653 | No | ||
| 38 | PCNA | Pcna | 9052 | -0.186 | -0.4699 | No | ||
| 39 | CDK2 | Cdk2 | 9072 | -0.189 | -0.4699 | No | ||
| 40 | CCNH | Ccnh | 9178 | -0.208 | -0.4755 | No | ||
| 41 | YWHAH | Ywhah | 9181 | -0.208 | -0.4744 | No | ||
| 42 | SKP1 | Skp1a | 9322 | -0.234 | -0.4820 | No | ||
| 43 | ANAPC1 | Anapc1 | 9469 | -0.264 | -0.4899 | No | ||
| 44 | ABL1 | Abl1 | 9747 | -0.322 | -0.5059 | No | ||
| 45 | YWHAQ | Ywhaq | 9796 | -0.330 | -0.5070 | No | ||
| 46 | E2F1 | E2f1 | 10020 | -0.376 | -0.5192 | No | ||
| 47 | ANAPC13 | Anapc13 | 10027 | -0.377 | -0.5173 | No | ||
| 48 | ANAPC4 | Anapc4 | 10123 | -0.395 | -0.5211 | No | ||
| 49 | YWHAZ | Ywhaz | 10438 | -0.425 | -0.5389 | No | ||
| 50 | GADD45G | Gadd45g | 10766 | -0.471 | -0.5573 | No | ||
| 51 | CDC16 | Cdc16 | 10775 | -0.472 | -0.5549 | No | ||
| 52 | E2F3 | E2f3 | 11074 | -0.521 | -0.5711 | No | ||
| 53 | ANAPC5 | Anapc5 | 11229 | -0.553 | -0.5778 | No | ||
| 54 | ORC3 | Orc3 | 11363 | -0.581 | -0.5829 | No | ||
| 55 | CUL1 | Cul1 | 11415 | -0.595 | -0.5826 | No | ||
| 56 | CDK7 | Cdk7 | 11668 | -0.661 | -0.5949 | No | ||
| 57 | ANAPC7 | Anapc7 | 11828 | -0.707 | -0.6009 | No | ||
| 58 | TP53 | Trp53 | 11831 | -0.708 | -0.5968 | No | ||
| 59 | CDC25B | Cdc25b | 11869 | -0.718 | -0.5948 | No | ||
| 60 | E2F5 | E2f5 | 11877 | -0.721 | -0.5909 | No | ||
| 61 | TGFB3 | Tgfb3 | 11962 | -0.747 | -0.5918 | No | ||
| 62 | BUB3 | Bub3 | 12066 | -0.775 | -0.5938 | No | ||
| 63 | CDC26 | Cdc26 | 12210 | -0.811 | -0.5982 | No | ||
| 64 | YWHAE | Ywhae | 12226 | -0.815 | -0.5942 | No | ||
| 65 | ATR | Atr | 12470 | -0.873 | -0.6047 | No | ||
| 66 | STAG1 | Stag1 | 12512 | -0.883 | -0.6020 | No | ||
| 67 | CDC25A | Cdc25a | 12552 | -0.893 | -0.5991 | No | ||
| 68 | MAD1L1 | Mad1l1 | 12575 | -0.901 | -0.5951 | No | ||
| 69 | FZR1 | Fzr1 | 12858 | -0.991 | -0.6074 | No | ||
| 70 | CHEK2 | Chek2 | 13060 | -1.073 | -0.6139 | No | ||
| 71 | HDAC2 | Hdac2 | 13243 | -1.156 | -0.6187 | No | ||
| 72 | MYC | Myc | 13632 | -1.357 | -0.6357 | Yes | ||
| 73 | HDAC1 | Hdac1 | 13740 | -1.420 | -0.6340 | Yes | ||
| 74 | E2F4 | E2f4 | 13753 | -1.426 | -0.6261 | Yes | ||
| 75 | SMC1A | Smc1a | 13851 | -1.489 | -0.6234 | Yes | ||
| 76 | CDC23 | Cdc23 | 13853 | -1.489 | -0.6144 | Yes | ||
| 77 | SMC3 | Smc3 | 13909 | -1.529 | -0.6087 | Yes | ||
| 78 | TFDP2 | Tfdp2 | 13937 | -1.555 | -0.6010 | Yes | ||
| 79 | MCM2 | Mcm2 | 14070 | -1.650 | -0.5996 | Yes | ||
| 80 | DBF4 | Dbf4 | 14177 | -1.738 | -0.5960 | Yes | ||
| 81 | E2F2 | E2f2 | 14180 | -1.744 | -0.5855 | Yes | ||
| 82 | TFDP1 | Tfdp1 | 14207 | -1.769 | -0.5765 | Yes | ||
| 83 | RAD21 | Rad21 | 14312 | -1.865 | -0.5719 | Yes | ||
| 84 | PKMYT1 | Pkmyt1 | 14372 | -1.926 | -0.5641 | Yes | ||
| 85 | CDC7 | Cdc7 | 14391 | -1.951 | -0.5534 | Yes | ||
| 86 | CCND1 | Ccnd1 | 14426 | -2.005 | -0.5435 | Yes | ||
| 87 | ORC1 | Orc1 | 14431 | -2.009 | -0.5315 | Yes | ||
| 88 | MCM4 | Mcm4 | 14442 | -2.032 | -0.5199 | Yes | ||
| 89 | ORC5 | Orc5 | 14446 | -2.039 | -0.5077 | Yes | ||
| 90 | CDKN1C | Cdkn1c | 14460 | -2.055 | -0.4961 | Yes | ||
| 91 | ORC2 | Orc2 | 14487 | -2.085 | -0.4851 | Yes | ||
| 92 | ORC6 | Orc6 | 14539 | -2.157 | -0.4754 | Yes | ||
| 93 | MCM7 | Mcm7 | 14645 | -2.323 | -0.4681 | Yes | ||
| 94 | SKP2 | Skp2 | 14837 | -2.642 | -0.4645 | Yes | ||
| 95 | CDKN2D | Cdkn2d | 14918 | -2.807 | -0.4526 | Yes | ||
| 96 | MCM3 | Mcm3 | 14982 | -2.950 | -0.4388 | Yes | ||
| 97 | RBL1 | Rbl1 | 15021 | -3.024 | -0.4230 | Yes | ||
| 98 | CDKN2C | Cdkn2c | 15043 | -3.083 | -0.4056 | Yes | ||
| 99 | MCM6 | Mcm6 | 15061 | -3.125 | -0.3878 | Yes | ||
| 100 | CCNE1 | Ccne1 | 15142 | -3.344 | -0.3727 | Yes | ||
| 101 | CHEK1 | Chek1 | 15169 | -3.426 | -0.3536 | Yes | ||
| 102 | CDK1 | Cdk1 | 15207 | -3.532 | -0.3346 | Yes | ||
| 103 | MAD2L1 | Mad2l1 | 15224 | -3.587 | -0.3139 | Yes | ||
| 104 | CCNE2 | Ccne2 | 15245 | -3.650 | -0.2931 | Yes | ||
| 105 | CDC45 | Cdc45 | 15263 | -3.711 | -0.2717 | Yes | ||
| 106 | MCM5 | Mcm5 | 15314 | -3.849 | -0.2516 | Yes | ||
| 107 | CDC6 | Cdc6 | 15404 | -4.168 | -0.2321 | Yes | ||
| 108 | CCNB1 | Ccnb1 | 15417 | -4.218 | -0.2073 | Yes | ||
| 109 | CDC25C | Cdc25c | 15428 | -4.264 | -0.1821 | Yes | ||
| 110 | ESPL1 | Espl1 | 15438 | -4.313 | -0.1565 | Yes | ||
| 111 | CDC20 | Cdc20 | 15447 | -4.345 | -0.1307 | Yes | ||
| 112 | BUB1 | Bub1 | 15466 | -4.393 | -0.1052 | Yes | ||
| 113 | CCNB2 | Ccnb2 | 15478 | -4.435 | -0.0790 | Yes | ||
| 114 | TTK | Ttk | 15484 | -4.462 | -0.0523 | Yes | ||
| 115 | CCNA2 | Ccna2 | 15489 | -4.484 | -0.0254 | Yes | ||
| 116 | PLK1 | Plk1 | 15502 | -4.560 | 0.0015 | Yes |