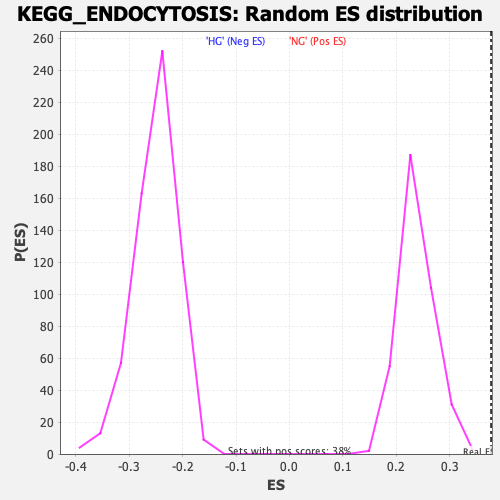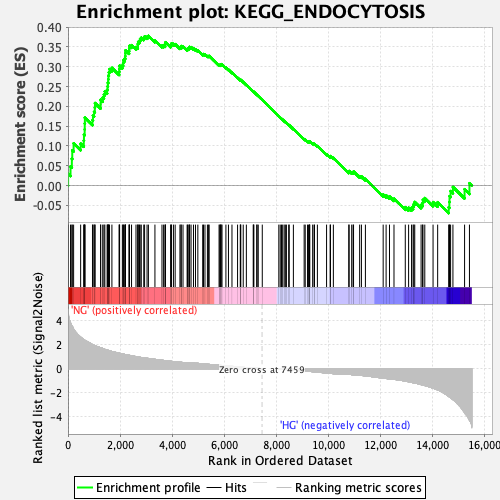
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_ENDOCYTOSIS |
| Enrichment Score (ES) | 0.37783647 |
| Normalized Enrichment Score (NES) | 1.5878602 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.11756118 |
| FWER p-Value | 0.564 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDR | Kdr | 5 | 4.757 | 0.0299 | Yes | ||
| 2 | ARAP2 | Arap2 | 97 | 3.810 | 0.0483 | Yes | ||
| 3 | CHMP4C | Chmp4c | 148 | 3.568 | 0.0677 | Yes | ||
| 4 | CXCR4 | Cxcr4 | 164 | 3.494 | 0.0890 | Yes | ||
| 5 | RET | Ret | 216 | 3.285 | 0.1066 | Yes | ||
| 6 | RAB11FIP4 | Rab11fip4 | 486 | 2.620 | 0.1058 | Yes | ||
| 7 | FGFR3 | Fgfr3 | 607 | 2.421 | 0.1134 | Yes | ||
| 8 | ASAP3 | Asap3 | 614 | 2.412 | 0.1283 | Yes | ||
| 9 | EHD3 | Ehd3 | 641 | 2.369 | 0.1417 | Yes | ||
| 10 | ARAP1 | Arap1 | 644 | 2.361 | 0.1566 | Yes | ||
| 11 | ADRB2 | Adrb2 | 649 | 2.357 | 0.1713 | Yes | ||
| 12 | GRK4 | Grk4 | 941 | 2.003 | 0.1652 | Yes | ||
| 13 | SH3GL3 | Sh3gl3 | 960 | 1.981 | 0.1766 | Yes | ||
| 14 | GIT1 | Git1 | 1003 | 1.942 | 0.1862 | Yes | ||
| 15 | PIKFYVE | Pikfyve | 1032 | 1.910 | 0.1966 | Yes | ||
| 16 | ADRB1 | Adrb1 | 1038 | 1.903 | 0.2083 | Yes | ||
| 17 | PARD6G | Pard6g | 1254 | 1.743 | 0.2054 | Yes | ||
| 18 | VPS25 | Vps25 | 1255 | 1.741 | 0.2165 | Yes | ||
| 19 | RNF41 | Rnf41 | 1335 | 1.675 | 0.2220 | Yes | ||
| 20 | SH3GL2 | Sh3gl2 | 1382 | 1.631 | 0.2294 | Yes | ||
| 21 | CXCR2 | Cxcr2 | 1416 | 1.605 | 0.2375 | Yes | ||
| 22 | LDLR | Ldlr | 1504 | 1.549 | 0.2417 | Yes | ||
| 23 | PLD2 | Pld2 | 1523 | 1.535 | 0.2503 | Yes | ||
| 24 | PLD1 | Pld1 | 1533 | 1.529 | 0.2594 | Yes | ||
| 25 | TRAF6 | Traf6 | 1548 | 1.520 | 0.2682 | Yes | ||
| 26 | HSPA8 | Hspa8 | 1554 | 1.517 | 0.2775 | Yes | ||
| 27 | CLTC | Cltc | 1572 | 1.504 | 0.2860 | Yes | ||
| 28 | PRKCZ | Prkcz | 1596 | 1.487 | 0.2940 | Yes | ||
| 29 | VPS37D | Vps37d | 1688 | 1.427 | 0.2971 | Yes | ||
| 30 | CHMP2B | Chmp2b | 1967 | 1.283 | 0.2872 | Yes | ||
| 31 | HGS | Hgs | 1974 | 1.281 | 0.2950 | Yes | ||
| 32 | SMURF1 | Smurf1 | 1984 | 1.276 | 0.3025 | Yes | ||
| 33 | IQSEC1 | Iqsec1 | 2090 | 1.225 | 0.3035 | Yes | ||
| 34 | MDM2 | Mdm2 | 2120 | 1.210 | 0.3093 | Yes | ||
| 35 | GIT2 | Git2 | 2130 | 1.202 | 0.3163 | Yes | ||
| 36 | STAM | Stam | 2181 | 1.182 | 0.3206 | Yes | ||
| 37 | CHMP1B | Chmp1b | 2201 | 1.173 | 0.3268 | Yes | ||
| 38 | VPS37B | Vps37b | 2205 | 1.172 | 0.3341 | Yes | ||
| 39 | FGFR2 | Fgfr2 | 2210 | 1.170 | 0.3413 | Yes | ||
| 40 | RAB31 | Rab31 | 2348 | 1.111 | 0.3394 | Yes | ||
| 41 | RAB11FIP5 | Rab11fip5 | 2354 | 1.109 | 0.3462 | Yes | ||
| 42 | STAM2 | Stam2 | 2363 | 1.105 | 0.3527 | Yes | ||
| 43 | PIP5K1B | Pip5k1b | 2443 | 1.065 | 0.3543 | Yes | ||
| 44 | PSD4 | Psd4 | 2614 | 1.003 | 0.3496 | Yes | ||
| 45 | CBLB | Cblb | 2684 | 0.974 | 0.3513 | Yes | ||
| 46 | PSD | Psd | 2686 | 0.973 | 0.3575 | Yes | ||
| 47 | ARFGAP3 | Arfgap3 | 2699 | 0.968 | 0.3629 | Yes | ||
| 48 | PARD6B | Pard6b | 2750 | 0.948 | 0.3656 | Yes | ||
| 49 | RAB11FIP1 | Rab11fip1 | 2773 | 0.940 | 0.3702 | Yes | ||
| 50 | ACAP3 | Acap3 | 2817 | 0.927 | 0.3733 | Yes | ||
| 51 | RAB22A | Rab22a | 2913 | 0.892 | 0.3728 | Yes | ||
| 52 | RABEP1 | Rabep1 | 2945 | 0.880 | 0.3764 | Yes | ||
| 53 | VPS4B | Vps4b | 3039 | 0.857 | 0.3758 | Yes | ||
| 54 | PIP5K1A | Pip5k1a | 3091 | 0.847 | 0.3778 | Yes | ||
| 55 | STAMBP | Stambp | 3342 | 0.773 | 0.3665 | No | ||
| 56 | MVB12A | Mvb12a | 3613 | 0.696 | 0.3533 | No | ||
| 57 | RAB11FIP3 | Rab11fip3 | 3676 | 0.681 | 0.3536 | No | ||
| 58 | SMURF2 | Smurf2 | 3733 | 0.666 | 0.3542 | No | ||
| 59 | CLTA | Clta | 3736 | 0.665 | 0.3583 | No | ||
| 60 | SMAP1 | Smap1 | 3749 | 0.663 | 0.3618 | No | ||
| 61 | IQSEC2 | Iqsec2 | 3947 | 0.610 | 0.3528 | No | ||
| 62 | FGFR4 | Fgfr4 | 3962 | 0.606 | 0.3558 | No | ||
| 63 | PSD2 | Psd2 | 3977 | 0.604 | 0.3587 | No | ||
| 64 | ASAP1 | Asap1 | 4053 | 0.583 | 0.3575 | No | ||
| 65 | SH3GLB1 | Sh3glb1 | 4120 | 0.567 | 0.3569 | No | ||
| 66 | CBL | Cbl | 4304 | 0.530 | 0.3483 | No | ||
| 67 | VPS37A | Vps37a | 4347 | 0.524 | 0.3489 | No | ||
| 68 | RAB11B | Rab11b | 4350 | 0.523 | 0.3521 | No | ||
| 69 | TSG101 | Tsg101 | 4422 | 0.503 | 0.3507 | No | ||
| 70 | KIT | Kit | 4580 | 0.493 | 0.3436 | No | ||
| 71 | IGF1R | Igf1r | 4597 | 0.491 | 0.3457 | No | ||
| 72 | PARD3 | Pard3 | 4647 | 0.479 | 0.3456 | No | ||
| 73 | RAB11A | Rab11a | 4662 | 0.477 | 0.3477 | No | ||
| 74 | DNAJC6 | Dnajc6 | 4675 | 0.474 | 0.3499 | No | ||
| 75 | EEA1 | Eea1 | 4731 | 0.461 | 0.3493 | No | ||
| 76 | ERBB4 | Erbb4 | 4824 | 0.443 | 0.3461 | No | ||
| 77 | CHMP4B | Chmp4b | 4901 | 0.439 | 0.3440 | No | ||
| 78 | AP2A1 | Ap2a1 | 4992 | 0.437 | 0.3409 | No | ||
| 79 | HSPA2 | Hspa2 | 5179 | 0.397 | 0.3313 | No | ||
| 80 | RAB5B | Rab5b | 5209 | 0.390 | 0.3319 | No | ||
| 81 | AGAP2 | Agap2 | 5266 | 0.379 | 0.3307 | No | ||
| 82 | SNF8 | Snf8 | 5360 | 0.362 | 0.3269 | No | ||
| 83 | ERBB3 | Erbb3 | 5398 | 0.355 | 0.3268 | No | ||
| 84 | SMAP2 | Smap2 | 5427 | 0.351 | 0.3272 | No | ||
| 85 | GRK5 | Grk5 | 5808 | 0.282 | 0.3042 | No | ||
| 86 | AGAP1 | Agap1 | 5834 | 0.278 | 0.3044 | No | ||
| 87 | MVB12B | Mvb12b | 5867 | 0.272 | 0.3040 | No | ||
| 88 | ARF6 | Arf6 | 5875 | 0.270 | 0.3053 | No | ||
| 89 | F2R | F2r | 5886 | 0.266 | 0.3063 | No | ||
| 90 | PIP4K2B | Pip4k2b | 5921 | 0.259 | 0.3058 | No | ||
| 91 | RAB5A | Rab5a | 6071 | 0.231 | 0.2975 | No | ||
| 92 | VPS37C | Vps37c | 6169 | 0.216 | 0.2926 | No | ||
| 93 | CHMP5 | Chmp5 | 6304 | 0.189 | 0.2851 | No | ||
| 94 | PIP5K1C | Pip5k1c | 6519 | 0.148 | 0.2721 | No | ||
| 95 | HRAS | Hras | 6623 | 0.128 | 0.2662 | No | ||
| 96 | DAB2 | Dab2 | 6636 | 0.126 | 0.2662 | No | ||
| 97 | RAB11FIP2 | Rab11fip2 | 6640 | 0.126 | 0.2668 | No | ||
| 98 | USP8 | Usp8 | 6737 | 0.107 | 0.2613 | No | ||
| 99 | LDLRAP1 | Ldlrap1 | 6857 | 0.086 | 0.2541 | No | ||
| 100 | IQSEC3 | Iqsec3 | 7125 | 0.050 | 0.2370 | No | ||
| 101 | GRK3 | Grk3 | 7127 | 0.050 | 0.2373 | No | ||
| 102 | ACAP2 | Acap2 | 7146 | 0.050 | 0.2364 | No | ||
| 103 | TFRC | Tfrc | 7255 | 0.033 | 0.2296 | No | ||
| 104 | SH3GLB2 | Sh3glb2 | 7271 | 0.030 | 0.2288 | No | ||
| 105 | DNM2 | Dnm2 | 7314 | 0.024 | 0.2262 | No | ||
| 106 | IL2RA | Il2ra | 7469 | 0.000 | 0.2162 | No | ||
| 107 | VTA1 | Vta1 | 8105 | -0.013 | 0.1750 | No | ||
| 108 | ARFGAP1 | Arfgap1 | 8159 | -0.024 | 0.1717 | No | ||
| 109 | AP2S1 | Ap2s1 | 8209 | -0.035 | 0.1687 | No | ||
| 110 | PARD6A | Pard6a | 8254 | -0.042 | 0.1661 | No | ||
| 111 | ASAP2 | Asap2 | 8311 | -0.055 | 0.1628 | No | ||
| 112 | CBLC | Cblc | 8356 | -0.063 | 0.1603 | No | ||
| 113 | EPN1 | Epn1 | 8397 | -0.070 | 0.1582 | No | ||
| 114 | AP2B1 | Ap2b1 | 8486 | -0.086 | 0.1530 | No | ||
| 115 | FLT1 | Flt1 | 8504 | -0.090 | 0.1525 | No | ||
| 116 | NEDD4L | Nedd4l | 8665 | -0.119 | 0.1428 | No | ||
| 117 | AP2M1 | Ap2m1 | 9079 | -0.191 | 0.1172 | No | ||
| 118 | SRC | Src | 9120 | -0.199 | 0.1158 | No | ||
| 119 | ARRB2 | Arrb2 | 9209 | -0.212 | 0.1114 | No | ||
| 120 | WWP1 | Wwp1 | 9244 | -0.220 | 0.1106 | No | ||
| 121 | VPS28 | Vps28 | 9282 | -0.227 | 0.1097 | No | ||
| 122 | CDC42 | Cdc42 | 9285 | -0.228 | 0.1110 | No | ||
| 123 | RAB5C | Rab5c | 9296 | -0.230 | 0.1118 | No | ||
| 124 | VPS36 | Vps36 | 9406 | -0.251 | 0.1063 | No | ||
| 125 | CLTB | Cltb | 9460 | -0.261 | 0.1045 | No | ||
| 126 | GRK2 | Grk2 | 9477 | -0.266 | 0.1052 | No | ||
| 127 | DNM1L | Dnm1l | 9588 | -0.288 | 0.0998 | No | ||
| 128 | EPN2 | Epn2 | 9941 | -0.363 | 0.0792 | No | ||
| 129 | RBSN | Rbsn | 10078 | -0.385 | 0.0728 | No | ||
| 130 | CHMP2A | Chmp2a | 10092 | -0.388 | 0.0745 | No | ||
| 131 | EHD1 | Ehd1 | 10202 | -0.416 | 0.0700 | No | ||
| 132 | EPS15 | Eps15 | 10795 | -0.477 | 0.0345 | No | ||
| 133 | ARFGAP2 | Arfgap2 | 10808 | -0.481 | 0.0368 | No | ||
| 134 | VPS45 | Vps45 | 10907 | -0.496 | 0.0336 | No | ||
| 135 | ITCH | Itch | 10961 | -0.501 | 0.0333 | No | ||
| 136 | EHD2 | Ehd2 | 10978 | -0.505 | 0.0355 | No | ||
| 137 | AP2A2 | Ap2a2 | 11213 | -0.550 | 0.0238 | No | ||
| 138 | SH3GL1 | Sh3gl1 | 11281 | -0.565 | 0.0230 | No | ||
| 139 | CSF1R | Csf1r | 11434 | -0.598 | 0.0169 | No | ||
| 140 | RAB4A | Rab4a | 12117 | -0.789 | -0.0225 | No | ||
| 141 | NEDD4 | Nedd4 | 12230 | -0.815 | -0.0246 | No | ||
| 142 | NTRK1 | Ntrk1 | 12360 | -0.851 | -0.0275 | No | ||
| 143 | PRKCI | Prkci | 12533 | -0.887 | -0.0331 | No | ||
| 144 | GRK1 | Grk1 | 12967 | -1.031 | -0.0547 | No | ||
| 145 | RUFY1 | Rufy1 | 13095 | -1.092 | -0.0560 | No | ||
| 146 | CHMP6 | Chmp6 | 13208 | -1.135 | -0.0561 | No | ||
| 147 | GRK6 | Grk6 | 13267 | -1.170 | -0.0524 | No | ||
| 148 | EGF | Egf | 13295 | -1.179 | -0.0467 | No | ||
| 149 | MET | Met | 13331 | -1.194 | -0.0414 | No | ||
| 150 | PSD3 | Psd3 | 13575 | -1.327 | -0.0487 | No | ||
| 151 | ARRB1 | Arrb1 | 13633 | -1.357 | -0.0438 | No | ||
| 152 | DNM1 | Dnm1 | 13645 | -1.366 | -0.0358 | No | ||
| 153 | IL2RB | Il2rb | 13717 | -1.405 | -0.0315 | No | ||
| 154 | EHD4 | Ehd4 | 14035 | -1.623 | -0.0418 | No | ||
| 155 | EGFR | Egfr | 14212 | -1.772 | -0.0420 | No | ||
| 156 | SH3KBP1 | Sh3kbp1 | 14636 | -2.304 | -0.0548 | No | ||
| 157 | DNM3 | Dnm3 | 14662 | -2.359 | -0.0415 | No | ||
| 158 | HSPA1L | Hspa1l | 14668 | -2.365 | -0.0267 | No | ||
| 159 | EPN3 | Epn3 | 14709 | -2.426 | -0.0139 | No | ||
| 160 | PDGFRA | Pdgfra | 14799 | -2.561 | -0.0034 | No | ||
| 161 | ARAP3 | Arap3 | 15247 | -3.652 | -0.0092 | No | ||
| 162 | ACAP1 | Acap1 | 15434 | -4.284 | 0.0059 | No |