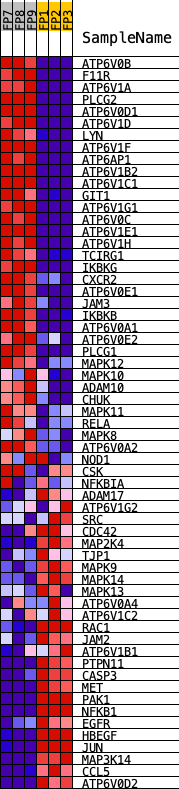
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_EPITHELIAL_CELL_SIGNALING_IN_HELICOBACTER_PYLORI_INFECTION |
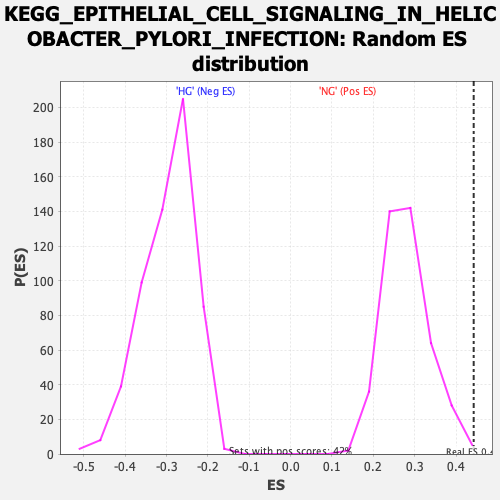
| Enrichment Score (ES) | 0.44259438 |
| Normalized Enrichment Score (NES) | 1.589257 |
| Nominal p-value | 0.0023980816 |
| FDR q-value | 0.13239923 |
| FWER p-Value | 0.56 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP6V0B | Atp6v0b | 326 | 2.960 | 0.0162 | Yes | ||
| 2 | F11R | F11r | 338 | 2.941 | 0.0525 | Yes | ||
| 3 | ATP6V1A | Atp6v1a | 398 | 2.800 | 0.0840 | Yes | ||
| 4 | PLCG2 | Plcg2 | 488 | 2.619 | 0.1112 | Yes | ||
| 5 | ATP6V0D1 | Atp6v0d1 | 505 | 2.590 | 0.1428 | Yes | ||
| 6 | ATP6V1D | Atp6v1d | 606 | 2.422 | 0.1668 | Yes | ||
| 7 | LYN | Lyn | 678 | 2.307 | 0.1912 | Yes | ||
| 8 | ATP6V1F | Atp6v1f | 685 | 2.295 | 0.2198 | Yes | ||
| 9 | ATP6AP1 | Atp6ap1 | 777 | 2.216 | 0.2418 | Yes | ||
| 10 | ATP6V1B2 | Atp6v1b2 | 810 | 2.172 | 0.2671 | Yes | ||
| 11 | ATP6V1C1 | Atp6v1c1 | 916 | 2.032 | 0.2859 | Yes | ||
| 12 | GIT1 | Git1 | 1003 | 1.942 | 0.3047 | Yes | ||
| 13 | ATP6V1G1 | Atp6v1g1 | 1072 | 1.876 | 0.3240 | Yes | ||
| 14 | ATP6V0C | Atp6v0c | 1074 | 1.875 | 0.3475 | Yes | ||
| 15 | ATP6V1E1 | Atp6v1e1 | 1257 | 1.740 | 0.3577 | Yes | ||
| 16 | ATP6V1H | Atp6v1h | 1338 | 1.675 | 0.3736 | Yes | ||
| 17 | TCIRG1 | Tcirg1 | 1341 | 1.672 | 0.3945 | Yes | ||
| 18 | IKBKG | Ikbkg | 1361 | 1.649 | 0.4140 | Yes | ||
| 19 | CXCR2 | Cxcr2 | 1416 | 1.605 | 0.4308 | Yes | ||
| 20 | ATP6V0E1 | Atp6v0e | 1575 | 1.503 | 0.4395 | Yes | ||
| 21 | JAM3 | Jam3 | 1795 | 1.373 | 0.4426 | Yes | ||
| 22 | IKBKB | Ikbkb | 2487 | 1.048 | 0.4111 | No | ||
| 23 | ATP6V0A1 | Atp6v0a1 | 2713 | 0.963 | 0.4087 | No | ||
| 24 | ATP6V0E2 | Atp6v0e2 | 2895 | 0.900 | 0.4083 | No | ||
| 25 | PLCG1 | Plcg1 | 3156 | 0.826 | 0.4019 | No | ||
| 26 | MAPK12 | Mapk12 | 3239 | 0.803 | 0.4067 | No | ||
| 27 | MAPK10 | Mapk10 | 3880 | 0.629 | 0.3733 | No | ||
| 28 | ADAM10 | Adam10 | 3918 | 0.620 | 0.3787 | No | ||
| 29 | CHUK | Chuk | 4201 | 0.548 | 0.3673 | No | ||
| 30 | MAPK11 | Mapk11 | 4459 | 0.501 | 0.3570 | No | ||
| 31 | RELA | Rela | 5054 | 0.426 | 0.3240 | No | ||
| 32 | MAPK8 | Mapk8 | 5645 | 0.310 | 0.2897 | No | ||
| 33 | ATP6V0A2 | Atp6v0a2 | 5843 | 0.277 | 0.2805 | No | ||
| 34 | NOD1 | Nod1 | 6097 | 0.227 | 0.2670 | No | ||
| 35 | CSK | Csk | 6745 | 0.106 | 0.2265 | No | ||
| 36 | NFKBIA | Nfkbia | 7373 | 0.013 | 0.1861 | No | ||
| 37 | ADAM17 | Adam17 | 8776 | -0.140 | 0.0972 | No | ||
| 38 | ATP6V1G2 | Atp6v1g2 | 9118 | -0.199 | 0.0777 | No | ||
| 39 | SRC | Src | 9120 | -0.199 | 0.0801 | No | ||
| 40 | CDC42 | Cdc42 | 9285 | -0.228 | 0.0724 | No | ||
| 41 | MAP2K4 | Map2k4 | 9633 | -0.297 | 0.0537 | No | ||
| 42 | TJP1 | Tjp1 | 9842 | -0.340 | 0.0445 | No | ||
| 43 | MAPK9 | Mapk9 | 9981 | -0.369 | 0.0402 | No | ||
| 44 | MAPK14 | Mapk14 | 10069 | -0.384 | 0.0394 | No | ||
| 45 | MAPK13 | Mapk13 | 10342 | -0.423 | 0.0272 | No | ||
| 46 | ATP6V0A4 | Atp6v0a4 | 10462 | -0.428 | 0.0249 | No | ||
| 47 | ATP6V1C2 | Atp6v1c2 | 11120 | -0.529 | -0.0109 | No | ||
| 48 | RAC1 | Rac1 | 11474 | -0.610 | -0.0261 | No | ||
| 49 | JAM2 | Jam2 | 12081 | -0.780 | -0.0554 | No | ||
| 50 | ATP6V1B1 | Atp6v1b1 | 12128 | -0.791 | -0.0485 | No | ||
| 51 | PTPN11 | Ptpn11 | 12241 | -0.819 | -0.0454 | No | ||
| 52 | CASP3 | Casp3 | 13319 | -1.187 | -0.1001 | No | ||
| 53 | MET | Met | 13331 | -1.194 | -0.0858 | No | ||
| 54 | PAK1 | Pak1 | 13387 | -1.226 | -0.0739 | No | ||
| 55 | NFKB1 | Nfkb1 | 13962 | -1.572 | -0.0912 | No | ||
| 56 | EGFR | Egfr | 14212 | -1.772 | -0.0850 | No | ||
| 57 | HBEGF | Hbegf | 14288 | -1.837 | -0.0667 | No | ||
| 58 | JUN | Jun | 14653 | -2.336 | -0.0608 | No | ||
| 59 | MAP3K14 | Map3k14 | 14707 | -2.419 | -0.0338 | No | ||
| 60 | CCL5 | Ccl5 | 14866 | -2.689 | -0.0101 | No | ||
| 61 | ATP6V0D2 | Atp6v0d2 | 15409 | -4.183 | 0.0075 | No |