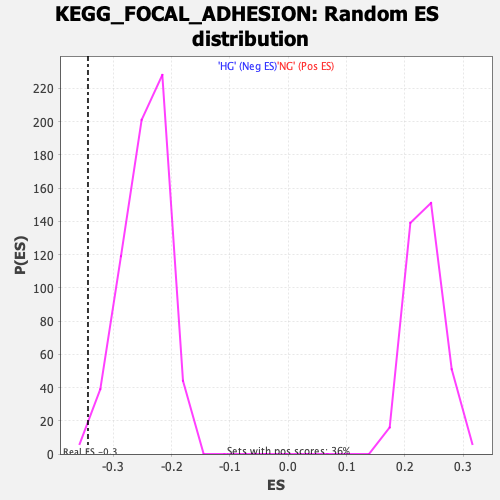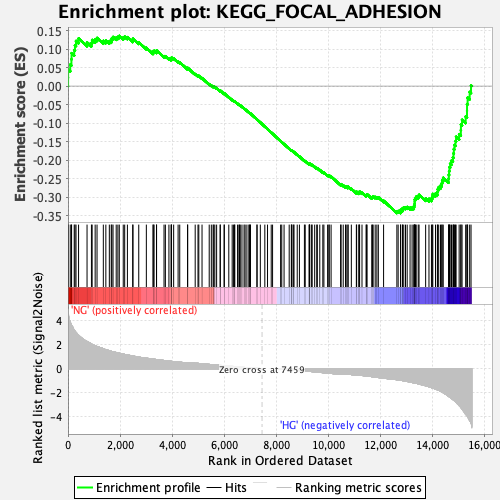
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_FOCAL_ADHESION |
| Enrichment Score (ES) | -0.34347287 |
| Normalized Enrichment Score (NES) | -1.3968558 |
| Nominal p-value | 0.0094191525 |
| FDR q-value | 0.21992494 |
| FWER p-Value | 0.991 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VWF | Vwf | 3 | 4.877 | 0.0227 | No | ||
| 2 | KDR | Kdr | 5 | 4.757 | 0.0451 | No | ||
| 3 | COL4A4 | Col4a4 | 86 | 3.885 | 0.0581 | No | ||
| 4 | FLT4 | Flt4 | 131 | 3.633 | 0.0723 | No | ||
| 5 | COL11A2 | Col11a2 | 138 | 3.605 | 0.0889 | No | ||
| 6 | LAMB3 | Lamb3 | 239 | 3.202 | 0.0974 | No | ||
| 7 | LAMA1 | Lama1 | 268 | 3.121 | 0.1103 | No | ||
| 8 | CHAD | Chad | 308 | 3.010 | 0.1219 | No | ||
| 9 | PARVG | Parvg | 406 | 2.785 | 0.1287 | No | ||
| 10 | THBS3 | Thbs3 | 732 | 2.258 | 0.1181 | No | ||
| 11 | ITGB5 | Itgb5 | 901 | 2.055 | 0.1168 | No | ||
| 12 | PDGFB | Pdgfb | 928 | 2.013 | 0.1246 | No | ||
| 13 | LAMA3 | Lama3 | 1044 | 1.897 | 0.1260 | No | ||
| 14 | ITGA9 | Itga9 | 1116 | 1.845 | 0.1301 | No | ||
| 15 | RELN | Reln | 1358 | 1.655 | 0.1222 | No | ||
| 16 | ITGB7 | Itgb7 | 1455 | 1.573 | 0.1233 | No | ||
| 17 | VEGFB | Vegfb | 1589 | 1.494 | 0.1217 | No | ||
| 18 | ITGB8 | Itgb8 | 1662 | 1.445 | 0.1238 | No | ||
| 19 | VEGFC | Vegfc | 1684 | 1.429 | 0.1291 | No | ||
| 20 | ELK1 | Elk1 | 1731 | 1.412 | 0.1328 | No | ||
| 21 | LAMB2 | Lamb2 | 1853 | 1.339 | 0.1312 | No | ||
| 22 | MYL12B | Myl12b | 1918 | 1.307 | 0.1331 | No | ||
| 23 | VEGFA | Vegfa | 1972 | 1.282 | 0.1357 | No | ||
| 24 | TNXB | Tnxb | 2124 | 1.208 | 0.1315 | No | ||
| 25 | SHC1 | Shc1 | 2179 | 1.183 | 0.1336 | No | ||
| 26 | CTNNB1 | Ctnnb1 | 2284 | 1.137 | 0.1322 | No | ||
| 27 | FN1 | Fn1 | 2488 | 1.047 | 0.1238 | No | ||
| 28 | GSK3B | Gsk3b | 2498 | 1.045 | 0.1282 | No | ||
| 29 | ITGB6 | Itgb6 | 2720 | 0.961 | 0.1183 | No | ||
| 30 | LAMA5 | Lama5 | 3012 | 0.866 | 0.1034 | No | ||
| 31 | COL5A2 | Col5a2 | 3272 | 0.793 | 0.0902 | No | ||
| 32 | ITGA10 | Gm42957 | 3275 | 0.792 | 0.0938 | No | ||
| 33 | PRKCG | Prkcg | 3309 | 0.781 | 0.0953 | No | ||
| 34 | BRAF | Braf | 3403 | 0.752 | 0.0928 | No | ||
| 35 | ERBB2 | Erbb2 | 3405 | 0.752 | 0.0963 | No | ||
| 36 | DIAPH1 | Diaph1 | 3692 | 0.678 | 0.0808 | No | ||
| 37 | SOS2 | Sos2 | 3744 | 0.664 | 0.0806 | No | ||
| 38 | MAPK10 | Mapk10 | 3880 | 0.629 | 0.0748 | No | ||
| 39 | FYN | Fyn | 3963 | 0.606 | 0.0723 | No | ||
| 40 | COL2A1 | Col2a1 | 3976 | 0.604 | 0.0743 | No | ||
| 41 | PARVB | Parvb | 3980 | 0.603 | 0.0770 | No | ||
| 42 | MAPK3 | Mapk3 | 4062 | 0.582 | 0.0744 | No | ||
| 43 | FLNA | Flna | 4233 | 0.542 | 0.0659 | No | ||
| 44 | ARHGAP35 | Arhgap35 | 4296 | 0.532 | 0.0644 | No | ||
| 45 | IGF1R | Igf1r | 4597 | 0.491 | 0.0471 | No | ||
| 46 | PTEN | Pten | 4601 | 0.490 | 0.0492 | No | ||
| 47 | MYLK3 | Mylk3 | 4890 | 0.441 | 0.0325 | No | ||
| 48 | PIK3CG | Pik3cg | 4998 | 0.436 | 0.0276 | No | ||
| 49 | ROCK1 | Rock1 | 5029 | 0.430 | 0.0276 | No | ||
| 50 | PIK3CD | Pik3cd | 5153 | 0.402 | 0.0215 | No | ||
| 51 | ITGA11 | Itga11 | 5429 | 0.351 | 0.0052 | No | ||
| 52 | PDPK1 | Pdpk1 | 5510 | 0.337 | 0.0016 | No | ||
| 53 | CRK | Crk | 5578 | 0.323 | -0.0013 | No | ||
| 54 | RAC3 | Rac3 | 5603 | 0.316 | -0.0013 | No | ||
| 55 | MAPK8 | Mapk8 | 5645 | 0.310 | -0.0026 | No | ||
| 56 | ACTN2 | Actn2 | 5714 | 0.296 | -0.0056 | No | ||
| 57 | ITGA7 | Itga7 | 5851 | 0.275 | -0.0132 | No | ||
| 58 | THBS4 | Thbs4 | 5854 | 0.275 | -0.0120 | No | ||
| 59 | CAPN2 | Capn2 | 6003 | 0.243 | -0.0205 | No | ||
| 60 | PIK3R5 | Pik3r5 | 6009 | 0.242 | -0.0197 | No | ||
| 61 | BAD | Bad | 6182 | 0.214 | -0.0299 | No | ||
| 62 | PAK3 | Pak3 | 6312 | 0.188 | -0.0374 | No | ||
| 63 | PDGFD | Pdgfd | 6366 | 0.178 | -0.0401 | No | ||
| 64 | PXN | Pxn | 6405 | 0.170 | -0.0417 | No | ||
| 65 | CCND3 | Ccnd3 | 6410 | 0.169 | -0.0412 | No | ||
| 66 | PIP5K1C | Pip5k1c | 6519 | 0.148 | -0.0476 | No | ||
| 67 | RAP1B | Rap1b | 6570 | 0.138 | -0.0502 | No | ||
| 68 | RHOA | Rhoa | 6593 | 0.134 | -0.0510 | No | ||
| 69 | HRAS | Hras | 6623 | 0.128 | -0.0523 | No | ||
| 70 | BCAR1 | Bcar1 | 6688 | 0.118 | -0.0559 | No | ||
| 71 | VAV2 | Vav2 | 6769 | 0.101 | -0.0606 | No | ||
| 72 | AKT2 | Akt2 | 6809 | 0.094 | -0.0627 | No | ||
| 73 | PDGFA | Pdgfa | 6878 | 0.084 | -0.0668 | No | ||
| 74 | ITGA3 | Itga3 | 6954 | 0.072 | -0.0713 | No | ||
| 75 | CAV3 | Cav3 | 6975 | 0.070 | -0.0723 | No | ||
| 76 | MYL10 | Myl10 | 7002 | 0.065 | -0.0737 | No | ||
| 77 | PIK3CA | Pik3ca | 7015 | 0.064 | -0.0742 | No | ||
| 78 | AKT1 | Akt1 | 7265 | 0.031 | -0.0903 | No | ||
| 79 | FLNB | Flnb | 7268 | 0.030 | -0.0902 | No | ||
| 80 | SOS1 | Sos1 | 7394 | 0.010 | -0.0983 | No | ||
| 81 | VAV1 | Vav1 | 7567 | 0.000 | -0.1096 | No | ||
| 82 | IBSP | Ibsp | 7671 | 0.000 | -0.1163 | No | ||
| 83 | VTN | Vtn | 7813 | 0.000 | -0.1255 | No | ||
| 84 | PAK5 | Pak7 | 7865 | 0.000 | -0.1288 | No | ||
| 85 | COMP | Comp | 8174 | -0.027 | -0.1488 | No | ||
| 86 | MAP2K1 | Map2k1 | 8218 | -0.037 | -0.1514 | No | ||
| 87 | LAMC3 | Lamc3 | 8307 | -0.054 | -0.1569 | No | ||
| 88 | FLT1 | Flt1 | 8504 | -0.090 | -0.1692 | No | ||
| 89 | XIAP | Xiap | 8586 | -0.105 | -0.1740 | No | ||
| 90 | RAPGEF1 | Rapgef1 | 8597 | -0.107 | -0.1742 | No | ||
| 91 | ITGAV | Itgav | 8611 | -0.109 | -0.1745 | No | ||
| 92 | ITGB1 | Itgb1 | 8679 | -0.122 | -0.1783 | No | ||
| 93 | LAMA2 | Lama2 | 8683 | -0.123 | -0.1779 | No | ||
| 94 | FLNC | Flnc | 8813 | -0.146 | -0.1856 | No | ||
| 95 | PAK6 | Pak6 | 8897 | -0.161 | -0.1903 | No | ||
| 96 | PIK3CB | Pik3cb | 9089 | -0.193 | -0.2019 | No | ||
| 97 | SRC | Src | 9120 | -0.199 | -0.2029 | No | ||
| 98 | PPP1R12A | Ppp1r12a | 9267 | -0.224 | -0.2113 | No | ||
| 99 | CDC42 | Cdc42 | 9285 | -0.228 | -0.2114 | No | ||
| 100 | CRKL | Crkl | 9288 | -0.229 | -0.2104 | No | ||
| 101 | ITGA5 | Itga5 | 9293 | -0.230 | -0.2096 | No | ||
| 102 | ROCK2 | Rock2 | 9367 | -0.243 | -0.2132 | No | ||
| 103 | PPP1CB | Ppp1cb | 9385 | -0.246 | -0.2132 | No | ||
| 104 | LAMB1 | Lamb1 | 9484 | -0.268 | -0.2183 | No | ||
| 105 | MAPK1 | Mapk1 | 9564 | -0.283 | -0.2221 | No | ||
| 106 | PAK2 | Pak2 | 9585 | -0.288 | -0.2221 | No | ||
| 107 | MYLK2 | Mylk2 | 9680 | -0.307 | -0.2268 | No | ||
| 108 | BCL2 | Bcl2 | 9797 | -0.330 | -0.2328 | No | ||
| 109 | COL5A3 | Col5a3 | 9838 | -0.340 | -0.2338 | No | ||
| 110 | MAPK9 | Mapk9 | 9981 | -0.369 | -0.2413 | No | ||
| 111 | VASP | Vasp | 10030 | -0.377 | -0.2427 | No | ||
| 112 | ITGA2B | Itga2b | 10043 | -0.379 | -0.2417 | No | ||
| 113 | TLN1 | Tln1 | 10115 | -0.393 | -0.2445 | No | ||
| 114 | ACTN1 | Actn1 | 10481 | -0.433 | -0.2662 | No | ||
| 115 | MYLPF | Mylpf | 10499 | -0.436 | -0.2653 | No | ||
| 116 | MYL2 | Myl2 | 10581 | -0.440 | -0.2685 | No | ||
| 117 | PAK4 | Pak4 | 10670 | -0.450 | -0.2721 | No | ||
| 118 | PDGFC | Pdgfc | 10694 | -0.456 | -0.2715 | No | ||
| 119 | BIRC2 | Birc2 | 10763 | -0.470 | -0.2737 | No | ||
| 120 | LAMC2 | Lamc2 | 10767 | -0.471 | -0.2717 | No | ||
| 121 | GRB2 | Grb2 | 10893 | -0.492 | -0.2775 | No | ||
| 122 | AKT3 | Akt3 | 11086 | -0.522 | -0.2876 | No | ||
| 123 | RAC2 | Rac2 | 11097 | -0.524 | -0.2858 | No | ||
| 124 | MYL7 | Myl7 | 11186 | -0.543 | -0.2889 | No | ||
| 125 | VAV3 | Vav3 | 11188 | -0.543 | -0.2864 | No | ||
| 126 | RAF1 | Raf1 | 11211 | -0.549 | -0.2853 | No | ||
| 127 | MYL12A | Myl12a | 11303 | -0.569 | -0.2886 | No | ||
| 128 | ARHGAP5 | Arhgap5 | 11461 | -0.606 | -0.2959 | No | ||
| 129 | RAC1 | Rac1 | 11474 | -0.610 | -0.2939 | No | ||
| 130 | PTK2 | Ptk2 | 11499 | -0.614 | -0.2925 | No | ||
| 131 | PPP1CA | Ppp1ca | 11678 | -0.663 | -0.3010 | No | ||
| 132 | RAP1A | Rap1a | 11705 | -0.672 | -0.2995 | No | ||
| 133 | TLN2 | Tln2 | 11731 | -0.681 | -0.2980 | No | ||
| 134 | LAMC1 | Lamc1 | 11810 | -0.703 | -0.2998 | No | ||
| 135 | DOCK1 | Dock1 | 11875 | -0.720 | -0.3005 | No | ||
| 136 | ITGA8 | Itga8 | 11931 | -0.738 | -0.3007 | No | ||
| 137 | COL6A6 | Col6a6 | 12133 | -0.793 | -0.3100 | No | ||
| 138 | ITGA4 | Itga4 | 12644 | -0.922 | -0.3390 | Yes | ||
| 139 | BIRC3 | Birc3 | 12693 | -0.938 | -0.3377 | Yes | ||
| 140 | VCL | Vcl | 12783 | -0.967 | -0.3389 | Yes | ||
| 141 | COL4A2 | Col4a2 | 12787 | -0.968 | -0.3346 | Yes | ||
| 142 | COL1A2 | Col1a2 | 12855 | -0.990 | -0.3343 | Yes | ||
| 143 | PPP1CC | Ppp1cc | 12863 | -0.993 | -0.3301 | Yes | ||
| 144 | THBS1 | Thbs1 | 12909 | -1.012 | -0.3282 | Yes | ||
| 145 | COL4A1 | Col4a1 | 12976 | -1.034 | -0.3277 | Yes | ||
| 146 | PIK3R1 | Pik3r1 | 13040 | -1.061 | -0.3268 | Yes | ||
| 147 | ACTB | Actb | 13151 | -1.112 | -0.3287 | Yes | ||
| 148 | CAV2 | Cav2 | 13229 | -1.147 | -0.3284 | Yes | ||
| 149 | EGF | Egf | 13295 | -1.179 | -0.3271 | Yes | ||
| 150 | MYL9 | Myl9 | 13308 | -1.184 | -0.3223 | Yes | ||
| 151 | COL4A6 | Col4a6 | 13326 | -1.191 | -0.3178 | Yes | ||
| 152 | ITGB4 | Itgb4 | 13329 | -1.194 | -0.3123 | Yes | ||
| 153 | MET | Met | 13331 | -1.194 | -0.3067 | Yes | ||
| 154 | ACTG1 | Actg1 | 13360 | -1.209 | -0.3029 | Yes | ||
| 155 | PAK1 | Pak1 | 13387 | -1.226 | -0.2988 | Yes | ||
| 156 | PRKCA | Prkca | 13464 | -1.260 | -0.2978 | Yes | ||
| 157 | ACTN4 | Actn4 | 13496 | -1.278 | -0.2938 | Yes | ||
| 158 | PGF | Pgf | 13750 | -1.425 | -0.3036 | Yes | ||
| 159 | ITGA6 | Itga6 | 13878 | -1.505 | -0.3048 | Yes | ||
| 160 | PARVA | Parva | 13973 | -1.578 | -0.3035 | Yes | ||
| 161 | COL11A1 | Col11a1 | 14005 | -1.603 | -0.2980 | Yes | ||
| 162 | ITGA2 | Itga2 | 14025 | -1.616 | -0.2917 | Yes | ||
| 163 | ITGA1 | Itga1 | 14129 | -1.695 | -0.2904 | Yes | ||
| 164 | PDGFRB | Pdgfrb | 14200 | -1.757 | -0.2867 | Yes | ||
| 165 | EGFR | Egfr | 14212 | -1.772 | -0.2791 | Yes | ||
| 166 | SHC4 | Shc4 | 14252 | -1.803 | -0.2731 | Yes | ||
| 167 | VEGFD | Vegfd | 14322 | -1.872 | -0.2688 | Yes | ||
| 168 | ZYX | Zyx | 14368 | -1.920 | -0.2627 | Yes | ||
| 169 | COL5A1 | Col5a1 | 14386 | -1.943 | -0.2547 | Yes | ||
| 170 | CCND1 | Ccnd1 | 14426 | -2.005 | -0.2478 | Yes | ||
| 171 | LAMA4 | Lama4 | 14633 | -2.302 | -0.2504 | Yes | ||
| 172 | MYLK | Mylk | 14640 | -2.317 | -0.2399 | Yes | ||
| 173 | JUN | Jun | 14653 | -2.336 | -0.2297 | Yes | ||
| 174 | COL6A3 | Col6a3 | 14666 | -2.360 | -0.2194 | Yes | ||
| 175 | ILK | Ilk | 14692 | -2.387 | -0.2098 | Yes | ||
| 176 | SHC3 | Shc3 | 14744 | -2.492 | -0.2014 | Yes | ||
| 177 | PDGFRA | Pdgfra | 14799 | -2.561 | -0.1929 | Yes | ||
| 178 | COL1A1 | Col1a1 | 14821 | -2.614 | -0.1819 | Yes | ||
| 179 | TNR | Tnr | 14834 | -2.636 | -0.1703 | Yes | ||
| 180 | CAV1 | Cav1 | 14857 | -2.675 | -0.1592 | Yes | ||
| 181 | ACTN3 | Actn3 | 14899 | -2.753 | -0.1489 | Yes | ||
| 182 | COL6A1 | Col6a1 | 14920 | -2.813 | -0.1370 | Yes | ||
| 183 | SPP1 | Spp1 | 15038 | -3.069 | -0.1302 | Yes | ||
| 184 | IGF1 | Igf1 | 15103 | -3.222 | -0.1192 | Yes | ||
| 185 | COL6A2 | Col6a2 | 15107 | -3.232 | -0.1042 | Yes | ||
| 186 | RASGRF1 | Rasgrf1 | 15150 | -3.350 | -0.0911 | Yes | ||
| 187 | HGF | Hgf | 15289 | -3.790 | -0.0823 | Yes | ||
| 188 | COL3A1 | Col3a1 | 15340 | -3.917 | -0.0671 | Yes | ||
| 189 | THBS2 | Thbs2 | 15342 | -3.926 | -0.0487 | Yes | ||
| 190 | TNN | Tnn | 15364 | -4.000 | -0.0313 | Yes | ||
| 191 | TNC | Tnc | 15443 | -4.338 | -0.0160 | Yes | ||
| 192 | PRKCB | Prkcb | 15495 | -4.518 | 0.0020 | Yes |