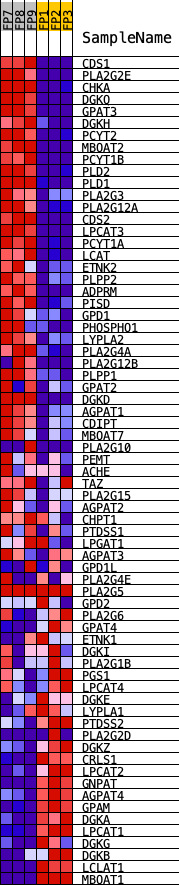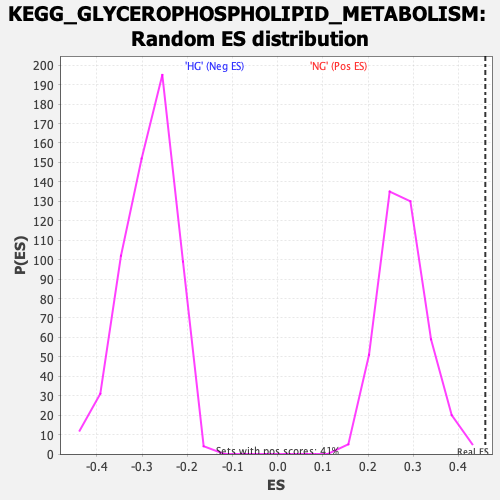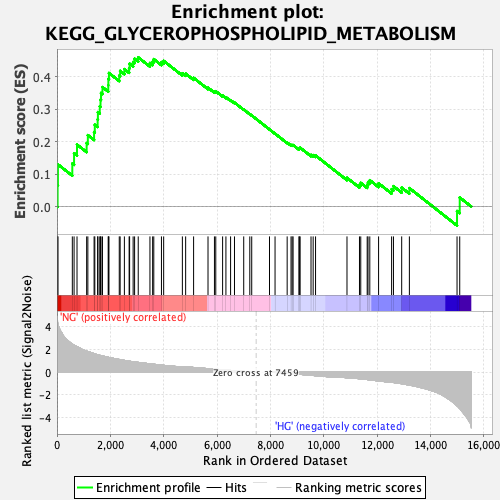
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_GLYCEROPHOSPHOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.45967278 |
| Normalized Enrichment Score (NES) | 1.6561913 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08117229 |
| FWER p-Value | 0.344 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDS1 | Cds1 | 23 | 4.472 | 0.0661 | Yes | ||
| 2 | PLA2G2E | Pla2g2e | 35 | 4.276 | 0.1300 | Yes | ||
| 3 | CHKA | Chka | 574 | 2.477 | 0.1326 | Yes | ||
| 4 | DGKQ | Dgkq | 640 | 2.370 | 0.1642 | Yes | ||
| 5 | GPAT3 | Gpat3 | 750 | 2.242 | 0.1910 | Yes | ||
| 6 | DGKH | Dgkh | 1111 | 1.847 | 0.1956 | Yes | ||
| 7 | PCYT2 | Pcyt2 | 1156 | 1.806 | 0.2201 | Yes | ||
| 8 | MBOAT2 | Mboat2 | 1390 | 1.624 | 0.2296 | Yes | ||
| 9 | PCYT1B | Pcyt1b | 1417 | 1.603 | 0.2521 | Yes | ||
| 10 | PLD2 | Pld2 | 1523 | 1.535 | 0.2685 | Yes | ||
| 11 | PLD1 | Pld1 | 1533 | 1.529 | 0.2910 | Yes | ||
| 12 | PLA2G3 | Pla2g3 | 1603 | 1.482 | 0.3089 | Yes | ||
| 13 | PLA2G12A | Pla2g12a | 1631 | 1.467 | 0.3294 | Yes | ||
| 14 | CDS2 | Cds2 | 1650 | 1.454 | 0.3502 | Yes | ||
| 15 | LPCAT3 | Lpcat3 | 1704 | 1.419 | 0.3682 | Yes | ||
| 16 | PCYT1A | Pcyt1a | 1922 | 1.306 | 0.3739 | Yes | ||
| 17 | LCAT | Lcat | 1925 | 1.304 | 0.3934 | Yes | ||
| 18 | ETNK2 | Etnk2 | 1947 | 1.292 | 0.4116 | Yes | ||
| 19 | PLPP2 | Plpp2 | 2333 | 1.116 | 0.4036 | Yes | ||
| 20 | ADPRM | Adprm | 2366 | 1.104 | 0.4182 | Yes | ||
| 21 | PISD | Pisd | 2525 | 1.033 | 0.4236 | Yes | ||
| 22 | GPD1 | Gpd1 | 2703 | 0.965 | 0.4267 | Yes | ||
| 23 | PHOSPHO1 | Phospho1 | 2721 | 0.961 | 0.4401 | Yes | ||
| 24 | LYPLA2 | Lypla2 | 2853 | 0.914 | 0.4454 | Yes | ||
| 25 | PLA2G4A | Pla2g4a | 2908 | 0.894 | 0.4555 | Yes | ||
| 26 | PLA2G12B | Pla2g12b | 3044 | 0.857 | 0.4597 | Yes | ||
| 27 | PLPP1 | Plpp1 | 3483 | 0.733 | 0.4424 | No | ||
| 28 | GPAT2 | Gpat2 | 3584 | 0.705 | 0.4466 | No | ||
| 29 | DGKD | Dgkd | 3627 | 0.692 | 0.4543 | No | ||
| 30 | AGPAT1 | Agpat1 | 3914 | 0.620 | 0.4452 | No | ||
| 31 | CDIPT | Cdipt | 3997 | 0.600 | 0.4490 | No | ||
| 32 | MBOAT7 | Mboat7 | 4698 | 0.469 | 0.4108 | No | ||
| 33 | PLA2G10 | Pla2g10 | 4821 | 0.443 | 0.4096 | No | ||
| 34 | PEMT | Pemt | 5119 | 0.410 | 0.3965 | No | ||
| 35 | ACHE | Ache | 5659 | 0.308 | 0.3663 | No | ||
| 36 | TAZ | Taz | 5900 | 0.263 | 0.3548 | No | ||
| 37 | PLA2G15 | Pla2g15 | 5950 | 0.253 | 0.3554 | No | ||
| 38 | AGPAT2 | Agpat2 | 6206 | 0.208 | 0.3421 | No | ||
| 39 | CHPT1 | Chpt1 | 6330 | 0.185 | 0.3369 | No | ||
| 40 | PTDSS1 | Ptdss1 | 6505 | 0.150 | 0.3279 | No | ||
| 41 | LPGAT1 | Lpgat1 | 6651 | 0.124 | 0.3204 | No | ||
| 42 | AGPAT3 | Agpat3 | 6997 | 0.066 | 0.2991 | No | ||
| 43 | GPD1L | Gpd1l | 7225 | 0.036 | 0.2850 | No | ||
| 44 | PLA2G4E | Pla2g4e | 7297 | 0.027 | 0.2808 | No | ||
| 45 | PLA2G5 | Pla2g5 | 7963 | 0.000 | 0.2377 | No | ||
| 46 | GPD2 | Gpd2 | 8171 | -0.026 | 0.2248 | No | ||
| 47 | PLA2G6 | Pla2g6 | 8625 | -0.112 | 0.1971 | No | ||
| 48 | GPAT4 | Gpat4 | 8770 | -0.139 | 0.1899 | No | ||
| 49 | ETNK1 | Etnk1 | 8811 | -0.146 | 0.1895 | No | ||
| 50 | DGKI | Dgki | 8847 | -0.152 | 0.1896 | No | ||
| 51 | PLA2G1B | Pla2g1b | 9067 | -0.189 | 0.1783 | No | ||
| 52 | PGS1 | Pgs1 | 9082 | -0.191 | 0.1802 | No | ||
| 53 | LPCAT4 | Lpcat4 | 9111 | -0.199 | 0.1814 | No | ||
| 54 | DGKE | Dgke | 9519 | -0.275 | 0.1592 | No | ||
| 55 | LYPLA1 | Lypla1 | 9599 | -0.291 | 0.1585 | No | ||
| 56 | PTDSS2 | Ptdss2 | 9689 | -0.309 | 0.1575 | No | ||
| 57 | PLA2G2D | Pla2g2d | 10867 | -0.489 | 0.0887 | No | ||
| 58 | DGKZ | Dgkz | 11335 | -0.575 | 0.0672 | No | ||
| 59 | CRLS1 | Crls1 | 11383 | -0.585 | 0.0730 | No | ||
| 60 | LPCAT2 | Lpcat2 | 11624 | -0.648 | 0.0672 | No | ||
| 61 | GNPAT | Gnpat | 11655 | -0.656 | 0.0752 | No | ||
| 62 | AGPAT4 | Agpat4 | 11722 | -0.678 | 0.0812 | No | ||
| 63 | GPAM | Gpam | 12051 | -0.770 | 0.0716 | No | ||
| 64 | DGKA | Dgka | 12539 | -0.890 | 0.0535 | No | ||
| 65 | LPCAT1 | Lpcat1 | 12608 | -0.910 | 0.0629 | No | ||
| 66 | DGKG | Dgkg | 12917 | -1.016 | 0.0583 | No | ||
| 67 | DGKB | Dgkb | 13205 | -1.134 | 0.0569 | No | ||
| 68 | LCLAT1 | Lclat1 | 14991 | -2.968 | -0.0138 | No | ||
| 69 | MBOAT1 | Mboat1 | 15092 | -3.193 | 0.0280 | No |