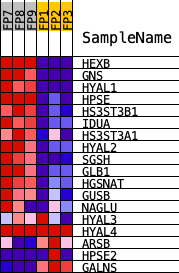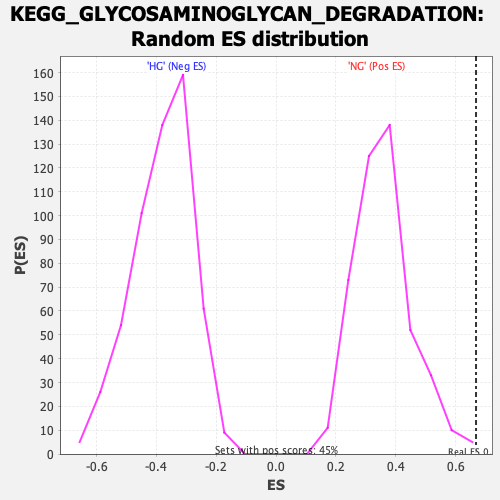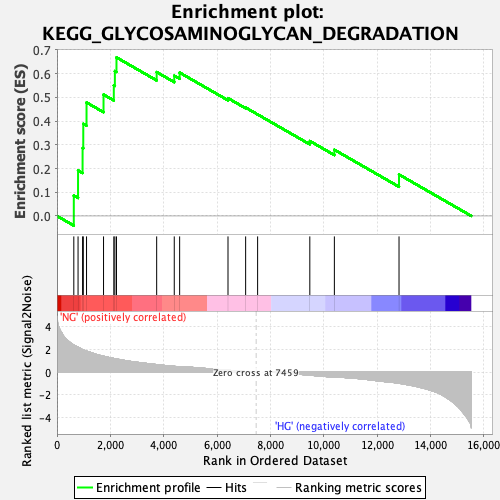
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_GLYCOSAMINOGLYCAN_DEGRADATION |
| Enrichment Score (ES) | 0.66819304 |
| Normalized Enrichment Score (NES) | 1.8551693 |
| Nominal p-value | 0.004474273 |
| FDR q-value | 0.013535163 |
| FWER p-Value | 0.035 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HEXB | Hexb | 630 | 2.389 | 0.0859 | Yes | ||
| 2 | GNS | Gns | 789 | 2.195 | 0.1919 | Yes | ||
| 3 | HYAL1 | Hyal1 | 959 | 1.983 | 0.2861 | Yes | ||
| 4 | HPSE | Hpse | 985 | 1.957 | 0.3881 | Yes | ||
| 5 | HS3ST3B1 | Hs3st3b1 | 1107 | 1.850 | 0.4782 | Yes | ||
| 6 | IDUA | Idua | 1744 | 1.404 | 0.5116 | Yes | ||
| 7 | HS3ST3A1 | Hs3st3a1 | 2129 | 1.202 | 0.5504 | Yes | ||
| 8 | HYAL2 | Hyal2 | 2169 | 1.186 | 0.6107 | Yes | ||
| 9 | SGSH | Sgsh | 2229 | 1.158 | 0.6682 | Yes | ||
| 10 | GLB1 | Glb1 | 3735 | 0.665 | 0.6064 | No | ||
| 11 | HGSNAT | Hgsnat | 4394 | 0.514 | 0.5911 | No | ||
| 12 | GUSB | Gusb | 4596 | 0.491 | 0.6042 | No | ||
| 13 | NAGLU | Naglu | 6408 | 0.169 | 0.4963 | No | ||
| 14 | HYAL3 | Hyal3 | 7069 | 0.056 | 0.4568 | No | ||
| 15 | HYAL4 | Hyal4 | 7518 | 0.000 | 0.4279 | No | ||
| 16 | ARSB | Arsb | 9474 | -0.265 | 0.3158 | No | ||
| 17 | HPSE2 | Hpse2 | 10394 | -0.424 | 0.2790 | No | ||
| 18 | GALNS | Galns | 12817 | -0.977 | 0.1746 | No |