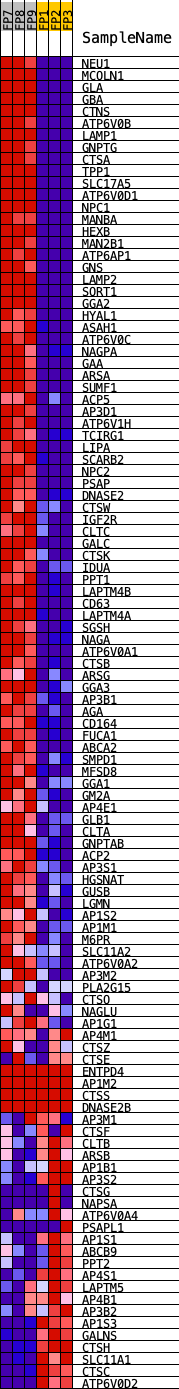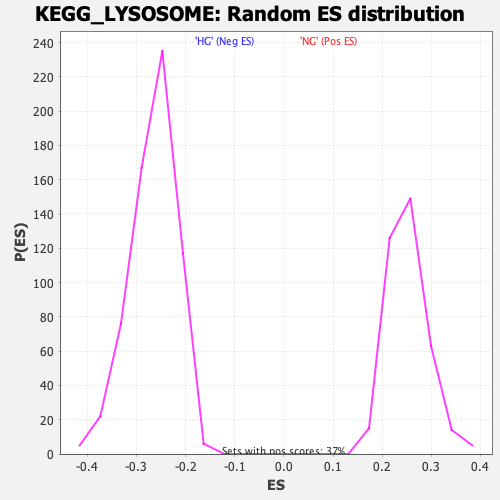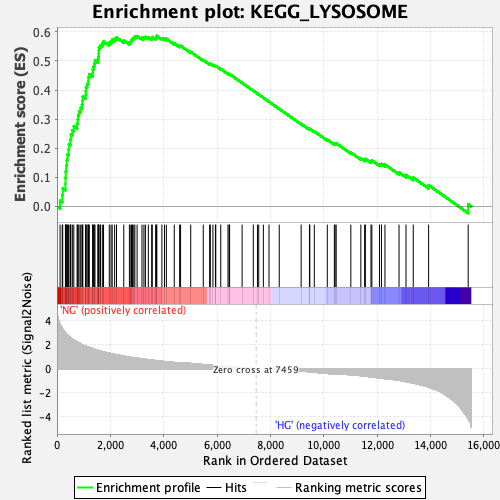
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_LYSOSOME |
| Enrichment Score (ES) | 0.5862246 |
| Normalized Enrichment Score (NES) | 2.3299654 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NEU1 | Neu1 | 115 | 3.729 | 0.0197 | Yes | ||
| 2 | MCOLN1 | Mcoln1 | 195 | 3.401 | 0.0393 | Yes | ||
| 3 | GLA | Gla | 214 | 3.309 | 0.0622 | Yes | ||
| 4 | GBA | Gba | 316 | 2.987 | 0.0774 | Yes | ||
| 5 | CTNS | Ctns | 318 | 2.982 | 0.0990 | Yes | ||
| 6 | ATP6V0B | Atp6v0b | 326 | 2.960 | 0.1201 | Yes | ||
| 7 | LAMP1 | Lamp1 | 346 | 2.915 | 0.1401 | Yes | ||
| 8 | GNPTG | Gnptg | 368 | 2.858 | 0.1595 | Yes | ||
| 9 | CTSA | Ctsa | 391 | 2.811 | 0.1786 | Yes | ||
| 10 | TPP1 | Tpp1 | 437 | 2.717 | 0.1954 | Yes | ||
| 11 | SLC17A5 | Slc17a5 | 446 | 2.699 | 0.2145 | Yes | ||
| 12 | ATP6V0D1 | Atp6v0d1 | 505 | 2.590 | 0.2296 | Yes | ||
| 13 | NPC1 | Npc1 | 518 | 2.575 | 0.2476 | Yes | ||
| 14 | MANBA | Manba | 575 | 2.473 | 0.2620 | Yes | ||
| 15 | HEXB | Hexb | 630 | 2.389 | 0.2758 | Yes | ||
| 16 | MAN2B1 | Man2b1 | 753 | 2.240 | 0.2842 | Yes | ||
| 17 | ATP6AP1 | Atp6ap1 | 777 | 2.216 | 0.2989 | Yes | ||
| 18 | GNS | Gns | 789 | 2.195 | 0.3141 | Yes | ||
| 19 | LAMP2 | Lamp2 | 827 | 2.152 | 0.3274 | Yes | ||
| 20 | SORT1 | Sort1 | 888 | 2.074 | 0.3386 | Yes | ||
| 21 | GGA2 | Gga2 | 935 | 2.007 | 0.3502 | Yes | ||
| 22 | HYAL1 | Hyal1 | 959 | 1.983 | 0.3631 | Yes | ||
| 23 | ASAH1 | Asah1 | 967 | 1.977 | 0.3771 | Yes | ||
| 24 | ATP6V0C | Atp6v0c | 1074 | 1.875 | 0.3838 | Yes | ||
| 25 | NAGPA | Nagpa | 1078 | 1.873 | 0.3973 | Yes | ||
| 26 | GAA | Gaa | 1086 | 1.865 | 0.4104 | Yes | ||
| 27 | ARSA | Arsa | 1129 | 1.833 | 0.4210 | Yes | ||
| 28 | SUMF1 | Sumf1 | 1175 | 1.794 | 0.4311 | Yes | ||
| 29 | ACP5 | Acp5 | 1176 | 1.792 | 0.4442 | Yes | ||
| 30 | AP3D1 | Ap3d1 | 1211 | 1.773 | 0.4549 | Yes | ||
| 31 | ATP6V1H | Atp6v1h | 1338 | 1.675 | 0.4589 | Yes | ||
| 32 | TCIRG1 | Tcirg1 | 1341 | 1.672 | 0.4709 | Yes | ||
| 33 | LIPA | Lipa | 1372 | 1.642 | 0.4809 | Yes | ||
| 34 | SCARB2 | Scarb2 | 1396 | 1.620 | 0.4912 | Yes | ||
| 35 | NPC2 | Npc2 | 1423 | 1.600 | 0.5012 | Yes | ||
| 36 | PSAP | Psap | 1529 | 1.533 | 0.5055 | Yes | ||
| 37 | DNASE2 | Dnase2a | 1553 | 1.517 | 0.5151 | Yes | ||
| 38 | CTSW | Ctsw | 1561 | 1.514 | 0.5257 | Yes | ||
| 39 | IGF2R | Igf2r | 1564 | 1.512 | 0.5365 | Yes | ||
| 40 | CLTC | Cltc | 1572 | 1.504 | 0.5470 | Yes | ||
| 41 | GALC | Galc | 1630 | 1.468 | 0.5540 | Yes | ||
| 42 | CTSK | Ctsk | 1699 | 1.423 | 0.5599 | Yes | ||
| 43 | IDUA | Idua | 1744 | 1.404 | 0.5673 | Yes | ||
| 44 | PPT1 | Ppt1 | 1959 | 1.286 | 0.5628 | Yes | ||
| 45 | LAPTM4B | Laptm4b | 2024 | 1.253 | 0.5677 | Yes | ||
| 46 | CD63 | Cd63 | 2068 | 1.235 | 0.5739 | Yes | ||
| 47 | LAPTM4A | Laptm4a | 2160 | 1.190 | 0.5767 | Yes | ||
| 48 | SGSH | Sgsh | 2229 | 1.158 | 0.5807 | Yes | ||
| 49 | NAGA | Naga | 2502 | 1.042 | 0.5707 | Yes | ||
| 50 | ATP6V0A1 | Atp6v0a1 | 2713 | 0.963 | 0.5640 | Yes | ||
| 51 | CTSB | Ctsb | 2767 | 0.943 | 0.5675 | Yes | ||
| 52 | ARSG | Arsg | 2792 | 0.934 | 0.5727 | Yes | ||
| 53 | GGA3 | Gga3 | 2831 | 0.922 | 0.5769 | Yes | ||
| 54 | AP3B1 | Ap3b1 | 2877 | 0.905 | 0.5806 | Yes | ||
| 55 | AGA | Aga | 2917 | 0.891 | 0.5846 | Yes | ||
| 56 | CD164 | Cd164 | 3002 | 0.869 | 0.5854 | Yes | ||
| 57 | FUCA1 | Fuca1 | 3185 | 0.819 | 0.5796 | Yes | ||
| 58 | ABCA2 | Abca2 | 3258 | 0.796 | 0.5807 | Yes | ||
| 59 | SMPD1 | Smpd1 | 3313 | 0.780 | 0.5829 | Yes | ||
| 60 | MFSD8 | Mfsd8 | 3422 | 0.746 | 0.5813 | Yes | ||
| 61 | GGA1 | Gga1 | 3549 | 0.713 | 0.5783 | Yes | ||
| 62 | GM2A | Gm2a | 3569 | 0.708 | 0.5823 | Yes | ||
| 63 | AP4E1 | Ap4e1 | 3696 | 0.677 | 0.5790 | Yes | ||
| 64 | GLB1 | Glb1 | 3735 | 0.665 | 0.5814 | Yes | ||
| 65 | CLTA | Clta | 3736 | 0.665 | 0.5862 | Yes | ||
| 66 | GNPTAB | Gnptab | 3930 | 0.614 | 0.5782 | No | ||
| 67 | ACP2 | Acp2 | 4027 | 0.590 | 0.5762 | No | ||
| 68 | AP3S1 | Ap3s1 | 4098 | 0.573 | 0.5759 | No | ||
| 69 | HGSNAT | Hgsnat | 4394 | 0.514 | 0.5605 | No | ||
| 70 | GUSB | Gusb | 4596 | 0.491 | 0.5510 | No | ||
| 71 | LGMN | Lgmn | 4631 | 0.483 | 0.5523 | No | ||
| 72 | AP1S2 | Ap1s2 | 5011 | 0.434 | 0.5309 | No | ||
| 73 | AP1M1 | Ap1m1 | 5481 | 0.341 | 0.5029 | No | ||
| 74 | M6PR | M6pr | 5721 | 0.296 | 0.4896 | No | ||
| 75 | SLC11A2 | Slc11a2 | 5745 | 0.291 | 0.4902 | No | ||
| 76 | ATP6V0A2 | Atp6v0a2 | 5843 | 0.277 | 0.4859 | No | ||
| 77 | AP3M2 | Ap3m2 | 5937 | 0.256 | 0.4818 | No | ||
| 78 | PLA2G15 | Pla2g15 | 5950 | 0.253 | 0.4828 | No | ||
| 79 | CTSO | Ctso | 6136 | 0.221 | 0.4724 | No | ||
| 80 | NAGLU | Naglu | 6408 | 0.169 | 0.4561 | No | ||
| 81 | AP1G1 | Ap1g1 | 6452 | 0.162 | 0.4545 | No | ||
| 82 | AP4M1 | Ap4m1 | 6468 | 0.157 | 0.4546 | No | ||
| 83 | CTSZ | Ctsz | 6937 | 0.074 | 0.4248 | No | ||
| 84 | CTSE | Ctse | 7356 | 0.015 | 0.3978 | No | ||
| 85 | ENTPD4 | Entpd4b | 7512 | 0.000 | 0.3878 | No | ||
| 86 | AP1M2 | Ap1m2 | 7560 | 0.000 | 0.3847 | No | ||
| 87 | CTSS | Ctss | 7733 | 0.000 | 0.3736 | No | ||
| 88 | DNASE2B | Dnase2b | 7945 | 0.000 | 0.3599 | No | ||
| 89 | AP3M1 | Ap3m1 | 8328 | -0.058 | 0.3355 | No | ||
| 90 | CTSF | Ctsf | 9149 | -0.203 | 0.2838 | No | ||
| 91 | CLTB | Cltb | 9460 | -0.261 | 0.2656 | No | ||
| 92 | ARSB | Arsb | 9474 | -0.265 | 0.2667 | No | ||
| 93 | AP1B1 | Ap1b1 | 9642 | -0.299 | 0.2580 | No | ||
| 94 | AP3S2 | Ap3s2 | 10129 | -0.396 | 0.2294 | No | ||
| 95 | CTSG | Ctsg | 10393 | -0.424 | 0.2154 | No | ||
| 96 | NAPSA | Napsa | 10421 | -0.424 | 0.2167 | No | ||
| 97 | ATP6V0A4 | Atp6v0a4 | 10462 | -0.428 | 0.2173 | No | ||
| 98 | PSAPL1 | Psapl1 | 11011 | -0.512 | 0.1854 | No | ||
| 99 | AP1S1 | Ap1s1 | 11386 | -0.585 | 0.1654 | No | ||
| 100 | ABCB9 | Abcb9 | 11529 | -0.624 | 0.1608 | No | ||
| 101 | PPT2 | Ppt2 | 11556 | -0.631 | 0.1637 | No | ||
| 102 | AP4S1 | Ap4s1 | 11766 | -0.692 | 0.1551 | No | ||
| 103 | LAPTM5 | Laptm5 | 11799 | -0.702 | 0.1582 | No | ||
| 104 | AP4B1 | Ap4b1 | 12084 | -0.782 | 0.1454 | No | ||
| 105 | AP3B2 | Ap3b2 | 12159 | -0.799 | 0.1464 | No | ||
| 106 | AP1S3 | Ap1s3 | 12290 | -0.836 | 0.1441 | No | ||
| 107 | GALNS | Galns | 12817 | -0.977 | 0.1171 | No | ||
| 108 | CTSH | Ctsh | 13079 | -1.083 | 0.1080 | No | ||
| 109 | SLC11A1 | Slc11a1 | 13350 | -1.202 | 0.0993 | No | ||
| 110 | CTSC | Ctsc | 13926 | -1.547 | 0.0732 | No | ||
| 111 | ATP6V0D2 | Atp6v0d2 | 15409 | -4.183 | 0.0075 | No |