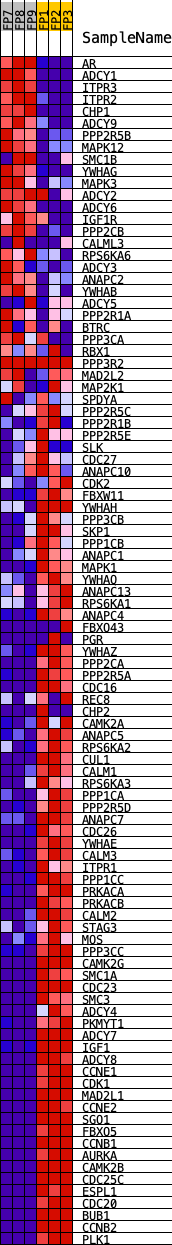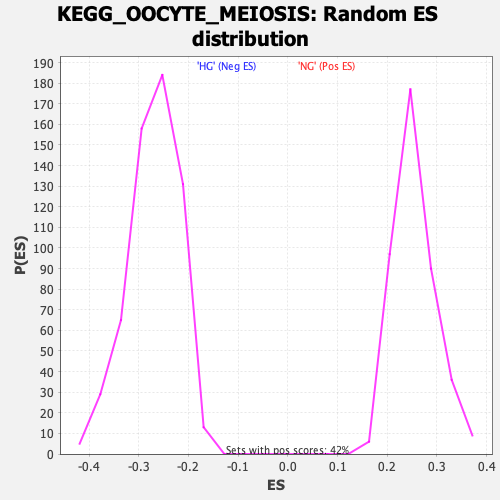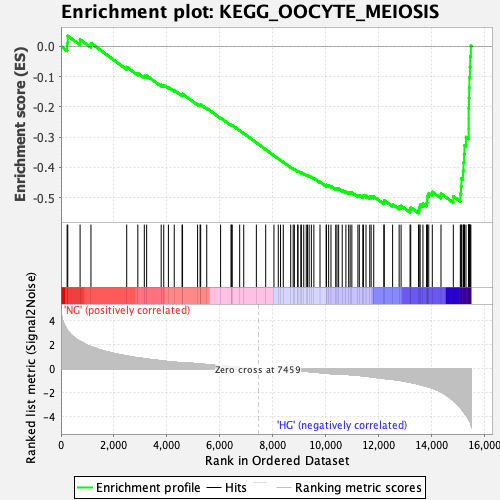
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_OOCYTE_MEIOSIS |
| Enrichment Score (ES) | -0.5513599 |
| Normalized Enrichment Score (NES) | -2.0498176 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AR | Ar | 232 | 3.222 | 0.0108 | No | ||
| 2 | ADCY1 | Adcy1 | 256 | 3.148 | 0.0346 | No | ||
| 3 | ITPR3 | Itpr3 | 722 | 2.272 | 0.0227 | No | ||
| 4 | ITPR2 | Itpr2 | 1133 | 1.827 | 0.0108 | No | ||
| 5 | CHP1 | Chp1 | 2485 | 1.048 | -0.0684 | No | ||
| 6 | ADCY9 | Adcy9 | 2906 | 0.895 | -0.0884 | No | ||
| 7 | PPP2R5B | Ppp2r5b | 3151 | 0.828 | -0.0976 | No | ||
| 8 | MAPK12 | Mapk12 | 3239 | 0.803 | -0.0968 | No | ||
| 9 | SMC1B | Smc1b | 3789 | 0.654 | -0.1272 | No | ||
| 10 | YWHAG | Ywhag | 3886 | 0.628 | -0.1283 | No | ||
| 11 | MAPK3 | Mapk3 | 4062 | 0.582 | -0.1350 | No | ||
| 12 | ADCY2 | Adcy2 | 4283 | 0.534 | -0.1450 | No | ||
| 13 | ADCY6 | Adcy6 | 4579 | 0.494 | -0.1601 | No | ||
| 14 | IGF1R | Igf1r | 4597 | 0.491 | -0.1573 | No | ||
| 15 | PPP2CB | Ppp2cb | 5166 | 0.399 | -0.1909 | No | ||
| 16 | CALML3 | Calml3 | 5261 | 0.379 | -0.1940 | No | ||
| 17 | RPS6KA6 | Rps6ka6 | 5283 | 0.376 | -0.1923 | No | ||
| 18 | ADCY3 | Adcy3 | 5512 | 0.336 | -0.2044 | No | ||
| 19 | ANAPC2 | Anapc2 | 6036 | 0.237 | -0.2364 | No | ||
| 20 | YWHAB | Ywhab | 6436 | 0.165 | -0.2609 | No | ||
| 21 | ADCY5 | Adcy5 | 6439 | 0.164 | -0.2598 | No | ||
| 22 | PPP2R1A | Ppp2r1a | 6477 | 0.156 | -0.2609 | No | ||
| 23 | BTRC | Btrc | 6758 | 0.104 | -0.2782 | No | ||
| 24 | PPP3CA | Ppp3ca | 6911 | 0.077 | -0.2875 | No | ||
| 25 | RBX1 | Rbx1 | 7389 | 0.011 | -0.3183 | No | ||
| 26 | PPP3R2 | Ppp3r2 | 7742 | 0.000 | -0.3411 | No | ||
| 27 | MAD2L2 | Mad2l2 | 8054 | -0.005 | -0.3612 | No | ||
| 28 | MAP2K1 | Map2k1 | 8218 | -0.037 | -0.3715 | No | ||
| 29 | SPDYA | Spdya | 8303 | -0.054 | -0.3765 | No | ||
| 30 | PPP2R5C | Ppp2r5c | 8407 | -0.072 | -0.3826 | No | ||
| 31 | PPP2R1B | Ppp2r1b | 8688 | -0.123 | -0.3998 | No | ||
| 32 | PPP2R5E | Ppp2r5e | 8782 | -0.141 | -0.4047 | No | ||
| 33 | SLK | Slk | 8826 | -0.147 | -0.4063 | No | ||
| 34 | CDC27 | Cdc27 | 8952 | -0.170 | -0.4130 | No | ||
| 35 | ANAPC10 | Anapc10 | 8964 | -0.171 | -0.4124 | No | ||
| 36 | CDK2 | Cdk2 | 9072 | -0.189 | -0.4178 | No | ||
| 37 | FBXW11 | Fbxw11 | 9091 | -0.194 | -0.4174 | No | ||
| 38 | YWHAH | Ywhah | 9181 | -0.208 | -0.4215 | No | ||
| 39 | PPP3CB | Ppp3cb | 9276 | -0.225 | -0.4258 | No | ||
| 40 | SKP1 | Skp1a | 9322 | -0.234 | -0.4268 | No | ||
| 41 | PPP1CB | Ppp1cb | 9385 | -0.246 | -0.4288 | No | ||
| 42 | ANAPC1 | Anapc1 | 9469 | -0.264 | -0.4321 | No | ||
| 43 | MAPK1 | Mapk1 | 9564 | -0.283 | -0.4359 | No | ||
| 44 | YWHAQ | Ywhaq | 9796 | -0.330 | -0.4483 | No | ||
| 45 | ANAPC13 | Anapc13 | 10027 | -0.377 | -0.4601 | No | ||
| 46 | RPS6KA1 | Rps6ka1 | 10042 | -0.379 | -0.4580 | No | ||
| 47 | ANAPC4 | Anapc4 | 10123 | -0.395 | -0.4600 | No | ||
| 48 | FBXO43 | Fbxo43 | 10211 | -0.418 | -0.4623 | No | ||
| 49 | PGR | Pgr | 10390 | -0.424 | -0.4704 | No | ||
| 50 | YWHAZ | Ywhaz | 10438 | -0.425 | -0.4701 | No | ||
| 51 | PPP2CA | Ppp2ca | 10501 | -0.437 | -0.4706 | No | ||
| 52 | PPP2R5A | Ppp2r5a | 10636 | -0.445 | -0.4757 | No | ||
| 53 | CDC16 | Cdc16 | 10775 | -0.472 | -0.4809 | No | ||
| 54 | REC8 | Rec8 | 10881 | -0.489 | -0.4838 | No | ||
| 55 | CHP2 | Chp2 | 10936 | -0.499 | -0.4832 | No | ||
| 56 | CAMK2A | Camk2a | 11001 | -0.510 | -0.4833 | No | ||
| 57 | ANAPC5 | Anapc5 | 11229 | -0.553 | -0.4936 | No | ||
| 58 | RPS6KA2 | Rps6ka2 | 11282 | -0.565 | -0.4924 | No | ||
| 59 | CUL1 | Cul1 | 11415 | -0.595 | -0.4962 | No | ||
| 60 | CALM1 | Calm1 | 11439 | -0.599 | -0.4929 | No | ||
| 61 | RPS6KA3 | Rps6ka3 | 11536 | -0.626 | -0.4941 | No | ||
| 62 | PPP1CA | Ppp1ca | 11678 | -0.663 | -0.4979 | No | ||
| 63 | PPP2R5D | Ppp2r5d | 11730 | -0.681 | -0.4957 | No | ||
| 64 | ANAPC7 | Anapc7 | 11828 | -0.707 | -0.4964 | No | ||
| 65 | CDC26 | Cdc26 | 12210 | -0.811 | -0.5145 | No | ||
| 66 | YWHAE | Ywhae | 12226 | -0.815 | -0.5090 | No | ||
| 67 | CALM3 | Calm3 | 12542 | -0.891 | -0.5223 | No | ||
| 68 | ITPR1 | Itpr1 | 12789 | -0.969 | -0.5304 | No | ||
| 69 | PPP1CC | Ppp1cc | 12863 | -0.993 | -0.5272 | No | ||
| 70 | PRKACA | Prkaca | 13202 | -1.133 | -0.5400 | No | ||
| 71 | PRKACB | Prkacb | 13228 | -1.145 | -0.5324 | No | ||
| 72 | CALM2 | Calm2 | 13521 | -1.294 | -0.5410 | Yes | ||
| 73 | STAG3 | Stag3 | 13545 | -1.312 | -0.5319 | Yes | ||
| 74 | MOS | Mos | 13584 | -1.334 | -0.5237 | Yes | ||
| 75 | PPP3CC | Ppp3cc | 13689 | -1.388 | -0.5193 | Yes | ||
| 76 | CAMK2G | Camk2g | 13828 | -1.470 | -0.5164 | Yes | ||
| 77 | SMC1A | Smc1a | 13851 | -1.489 | -0.5059 | Yes | ||
| 78 | CDC23 | Cdc23 | 13853 | -1.489 | -0.4940 | Yes | ||
| 79 | SMC3 | Smc3 | 13909 | -1.529 | -0.4853 | Yes | ||
| 80 | ADCY4 | Adcy4 | 14043 | -1.629 | -0.4809 | Yes | ||
| 81 | PKMYT1 | Pkmyt1 | 14372 | -1.926 | -0.4867 | Yes | ||
| 82 | ADCY7 | Adcy7 | 14838 | -2.643 | -0.4956 | Yes | ||
| 83 | IGF1 | Igf1 | 15103 | -3.222 | -0.4869 | Yes | ||
| 84 | ADCY8 | Adcy8 | 15131 | -3.320 | -0.4620 | Yes | ||
| 85 | CCNE1 | Ccne1 | 15142 | -3.344 | -0.4358 | Yes | ||
| 86 | CDK1 | Cdk1 | 15207 | -3.532 | -0.4116 | Yes | ||
| 87 | MAD2L1 | Mad2l1 | 15224 | -3.587 | -0.3839 | Yes | ||
| 88 | CCNE2 | Ccne2 | 15245 | -3.650 | -0.3559 | Yes | ||
| 89 | SGO1 | Sgo1 | 15261 | -3.699 | -0.3272 | Yes | ||
| 90 | FBXO5 | Fbxo5 | 15316 | -3.853 | -0.2998 | Yes | ||
| 91 | CCNB1 | Ccnb1 | 15417 | -4.218 | -0.2724 | Yes | ||
| 92 | AURKA | Aurka | 15418 | -4.222 | -0.2385 | Yes | ||
| 93 | CAMK2B | Camk2b | 15419 | -4.225 | -0.2046 | Yes | ||
| 94 | CDC25C | Cdc25c | 15428 | -4.264 | -0.1709 | Yes | ||
| 95 | ESPL1 | Espl1 | 15438 | -4.313 | -0.1369 | Yes | ||
| 96 | CDC20 | Cdc20 | 15447 | -4.345 | -0.1026 | Yes | ||
| 97 | BUB1 | Bub1 | 15466 | -4.393 | -0.0685 | Yes | ||
| 98 | CCNB2 | Ccnb2 | 15478 | -4.435 | -0.0336 | Yes | ||
| 99 | PLK1 | Plk1 | 15502 | -4.560 | 0.0015 | Yes |