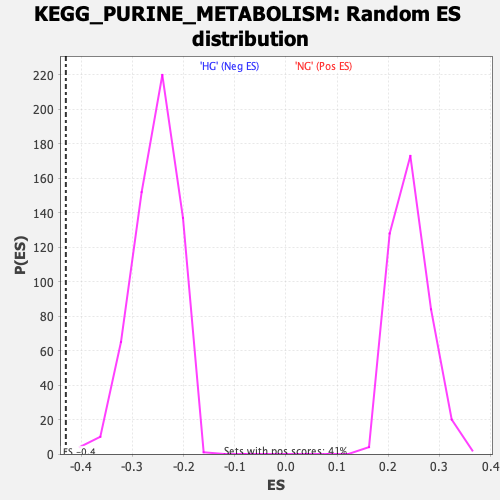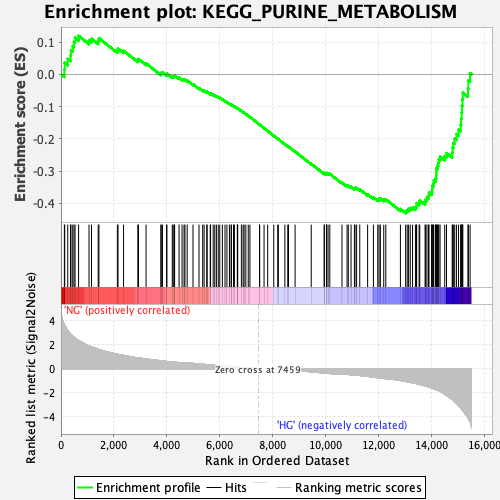
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_PURINE_METABOLISM |
| Enrichment Score (ES) | -0.43025044 |
| Normalized Enrichment Score (NES) | -1.6934788 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.027025063 |
| FWER p-Value | 0.278 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDE5A | Pde5a | 127 | 3.643 | 0.0142 | No | ||
| 2 | PDE6B | Pde6b | 141 | 3.596 | 0.0354 | No | ||
| 3 | ADCY1 | Adcy1 | 256 | 3.148 | 0.0474 | No | ||
| 4 | AMPD3 | Ampd3 | 360 | 2.875 | 0.0584 | No | ||
| 5 | GUCY1A2 | Gucy1a2 | 377 | 2.838 | 0.0748 | No | ||
| 6 | NPR1 | Npr1 | 444 | 2.703 | 0.0871 | No | ||
| 7 | PAPSS2 | Papss2 | 492 | 2.616 | 0.1002 | No | ||
| 8 | PDE3A | Pde3a | 531 | 2.539 | 0.1133 | No | ||
| 9 | PDE8A | Pde8a | 665 | 2.329 | 0.1190 | No | ||
| 10 | NPR2 | Npr2 | 1060 | 1.887 | 0.1050 | No | ||
| 11 | NME2 | Nme2 | 1154 | 1.809 | 0.1101 | No | ||
| 12 | ENTPD2 | Entpd2 | 1406 | 1.612 | 0.1037 | No | ||
| 13 | PNP | Pnp | 1435 | 1.591 | 0.1117 | No | ||
| 14 | ENTPD5 | Entpd5 | 2132 | 1.201 | 0.0738 | No | ||
| 15 | PDE10A | Pde10a | 2156 | 1.190 | 0.0796 | No | ||
| 16 | ADPRM | Adprm | 2366 | 1.104 | 0.0728 | No | ||
| 17 | ADCY9 | Adcy9 | 2906 | 0.895 | 0.0433 | No | ||
| 18 | POLR3A | Polr3a | 2931 | 0.886 | 0.0472 | No | ||
| 19 | ADA | Ada | 3217 | 0.809 | 0.0337 | No | ||
| 20 | NT5C | Nt5c | 3776 | 0.657 | 0.0014 | No | ||
| 21 | PRPS1L1 | Prps1l1 | 3792 | 0.654 | 0.0045 | No | ||
| 22 | POLR3F | Polr3f | 3837 | 0.640 | 0.0056 | No | ||
| 23 | PRUNE1 | Prune1 | 3998 | 0.600 | -0.0012 | No | ||
| 24 | NT5E | Nt5e | 4002 | 0.598 | 0.0023 | No | ||
| 25 | XDH | Xdh | 4208 | 0.547 | -0.0076 | No | ||
| 26 | RRM2B | Rrm2b | 4242 | 0.540 | -0.0065 | No | ||
| 27 | ADCY2 | Adcy2 | 4283 | 0.534 | -0.0058 | No | ||
| 28 | POLR3D | Polr3d | 4297 | 0.532 | -0.0033 | No | ||
| 29 | PKLR | Pklr | 4465 | 0.499 | -0.0111 | No | ||
| 30 | ADCY6 | Adcy6 | 4579 | 0.494 | -0.0154 | No | ||
| 31 | GUCY2C | Gucy2c | 4654 | 0.478 | -0.0173 | No | ||
| 32 | NME6 | Nme6 | 4688 | 0.472 | -0.0165 | No | ||
| 33 | NME5 | Nme5 | 4772 | 0.451 | -0.0192 | No | ||
| 34 | CANT1 | Cant1 | 4993 | 0.436 | -0.0308 | No | ||
| 35 | PRPS2 | Prps2 | 5219 | 0.388 | -0.0430 | No | ||
| 36 | POLR2L | Polr2l | 5359 | 0.362 | -0.0498 | No | ||
| 37 | GUCY2F | Gucy2f | 5413 | 0.353 | -0.0511 | No | ||
| 38 | ADCY3 | Adcy3 | 5512 | 0.336 | -0.0554 | No | ||
| 39 | AK4 | Ak4 | 5527 | 0.334 | -0.0543 | No | ||
| 40 | POLR1C | Polr1c | 5644 | 0.311 | -0.0599 | No | ||
| 41 | NUDT2 | Nudt2 | 5656 | 0.308 | -0.0587 | No | ||
| 42 | ADCY10 | Adcy10 | 5755 | 0.290 | -0.0633 | No | ||
| 43 | POLR2A | Polr2a | 5816 | 0.281 | -0.0655 | No | ||
| 44 | ATIC | Atic | 5877 | 0.269 | -0.0677 | No | ||
| 45 | PDE1C | Pde1c | 5948 | 0.253 | -0.0707 | No | ||
| 46 | NT5C2 | Nt5c2 | 6005 | 0.243 | -0.0729 | No | ||
| 47 | ENTPD8 | Entpd8 | 6103 | 0.227 | -0.0778 | No | ||
| 48 | GUK1 | Guk1 | 6203 | 0.210 | -0.0829 | No | ||
| 49 | PDE7A | Pde7a | 6276 | 0.193 | -0.0864 | No | ||
| 50 | POLR3B | Polr3b | 6371 | 0.177 | -0.0914 | No | ||
| 51 | ADCY5 | Adcy5 | 6439 | 0.164 | -0.0948 | No | ||
| 52 | POLR2F | Polr2f | 6440 | 0.164 | -0.0938 | No | ||
| 53 | AK5 | Ak5 | 6524 | 0.148 | -0.0982 | No | ||
| 54 | PDE6G | Pde6g | 6558 | 0.141 | -0.0995 | No | ||
| 55 | POLR2K | Polr2k | 6668 | 0.122 | -0.1059 | No | ||
| 56 | NME1 | Nme1 | 6678 | 0.121 | -0.1057 | No | ||
| 57 | POLE4 | Pole4 | 6682 | 0.120 | -0.1052 | No | ||
| 58 | POLR2E | Polr2e | 6822 | 0.093 | -0.1136 | No | ||
| 59 | NUDT9 | Nudt9 | 6887 | 0.081 | -0.1173 | No | ||
| 60 | AK7 | Ak7 | 6924 | 0.075 | -0.1192 | No | ||
| 61 | NME4 | Nme4 | 6990 | 0.068 | -0.1230 | No | ||
| 62 | GUCY2D | Gucy2e | 7082 | 0.054 | -0.1285 | No | ||
| 63 | POLR3C | Polr3c | 7145 | 0.050 | -0.1323 | No | ||
| 64 | ENTPD3 | Entpd3 | 7511 | 0.000 | -0.1560 | No | ||
| 65 | ENTPD4 | Entpd4b | 7512 | 0.000 | -0.1560 | No | ||
| 66 | ALLC | Allc | 7680 | 0.000 | -0.1668 | No | ||
| 67 | PDE2A | Gm45837 | 7817 | 0.000 | -0.1757 | No | ||
| 68 | POLR1D | Polr1d | 8047 | -0.003 | -0.1906 | No | ||
| 69 | AK1 | Ak1 | 8198 | -0.033 | -0.2001 | No | ||
| 70 | POLR2G | Polr2g | 8221 | -0.037 | -0.2013 | No | ||
| 71 | GUCY1B1 | Gucy1b1 | 8467 | -0.083 | -0.2167 | No | ||
| 72 | AMPD2 | Ampd2 | 8577 | -0.103 | -0.2232 | No | ||
| 73 | POLR2H | Polr2h | 8604 | -0.108 | -0.2242 | No | ||
| 74 | HPRT1 | Hprt | 8854 | -0.153 | -0.2394 | No | ||
| 75 | PDE4B | Pde4b | 9464 | -0.262 | -0.2774 | No | ||
| 76 | FHIT | Fhit | 9955 | -0.365 | -0.3070 | No | ||
| 77 | APRT | Aprt | 9959 | -0.366 | -0.3050 | No | ||
| 78 | PKM | Pkm | 10041 | -0.378 | -0.3079 | No | ||
| 79 | POLR3K | Polr3k | 10048 | -0.380 | -0.3060 | No | ||
| 80 | PDE6A | Pde6a | 10101 | -0.390 | -0.3069 | No | ||
| 81 | IMPDH1 | Impdh1 | 10167 | -0.407 | -0.3087 | No | ||
| 82 | PRPS1 | Prps1 | 10627 | -0.444 | -0.3358 | No | ||
| 83 | PDE9A | Pde9a | 10820 | -0.484 | -0.3453 | No | ||
| 84 | PDE11A | Pde11a | 10866 | -0.489 | -0.3452 | No | ||
| 85 | NT5C3A | Nt5c3 | 10970 | -0.503 | -0.3488 | No | ||
| 86 | POLR2I | Polr2i | 11098 | -0.525 | -0.3538 | No | ||
| 87 | PDE6D | Pde6d | 11119 | -0.529 | -0.3519 | No | ||
| 88 | PDE4A | Pde4a | 11180 | -0.541 | -0.3524 | No | ||
| 89 | ENTPD1 | Entpd1 | 11300 | -0.569 | -0.3567 | No | ||
| 90 | PDE6H | Pde6h | 11597 | -0.642 | -0.3720 | No | ||
| 91 | NME7 | Nme7 | 11816 | -0.705 | -0.3818 | No | ||
| 92 | DGUOK | Dguok | 11981 | -0.753 | -0.3878 | No | ||
| 93 | POLR2B | Polr2b | 12029 | -0.766 | -0.3862 | No | ||
| 94 | ZNRD1 | Znrd1 | 12079 | -0.779 | -0.3846 | No | ||
| 95 | NUDT5 | Nudt5 | 12207 | -0.810 | -0.3878 | No | ||
| 96 | PDE1B | Pde1b | 12285 | -0.834 | -0.3877 | No | ||
| 97 | POLR1E | Polr1e | 12837 | -0.986 | -0.4174 | No | ||
| 98 | AMPD1 | Ampd1 | 13035 | -1.060 | -0.4237 | Yes | ||
| 99 | POLR2C | Polr2c | 13094 | -1.090 | -0.4208 | Yes | ||
| 100 | POLR1A | Polr1a | 13147 | -1.110 | -0.4173 | Yes | ||
| 101 | PNPT1 | Pnpt1 | 13209 | -1.135 | -0.4143 | Yes | ||
| 102 | ENTPD6 | Entpd6 | 13296 | -1.179 | -0.4127 | Yes | ||
| 103 | PFAS | Pfas | 13405 | -1.234 | -0.4121 | Yes | ||
| 104 | ADK | Adk | 13447 | -1.251 | -0.4071 | Yes | ||
| 105 | NT5M | Nt5m | 13450 | -1.252 | -0.3995 | Yes | ||
| 106 | POLD2 | Pold2 | 13544 | -1.311 | -0.3975 | Yes | ||
| 107 | PAICS | Paics | 13569 | -1.325 | -0.3909 | Yes | ||
| 108 | GMPR | Gmpr | 13770 | -1.436 | -0.3950 | Yes | ||
| 109 | ADSL | Adsl | 13792 | -1.447 | -0.3875 | Yes | ||
| 110 | ADSS2 | Adss | 13842 | -1.482 | -0.3816 | Yes | ||
| 111 | POLR2D | Polr2d | 13903 | -1.524 | -0.3761 | Yes | ||
| 112 | GART | Gart | 13917 | -1.540 | -0.3675 | Yes | ||
| 113 | GMPR2 | Gmpr2 | 14015 | -1.608 | -0.3639 | Yes | ||
| 114 | GMPS | Gmps | 14039 | -1.627 | -0.3554 | Yes | ||
| 115 | ADCY4 | Adcy4 | 14043 | -1.629 | -0.3455 | Yes | ||
| 116 | GUCY1A1 | Gucy1a1 | 14085 | -1.664 | -0.3379 | Yes | ||
| 117 | PDE4D | Pde4d | 14100 | -1.677 | -0.3285 | Yes | ||
| 118 | PAPSS1 | Papss1 | 14168 | -1.733 | -0.3222 | Yes | ||
| 119 | POLE3 | Pole3 | 14191 | -1.751 | -0.3129 | Yes | ||
| 120 | POLR1B | Polr1b | 14194 | -1.751 | -0.3023 | Yes | ||
| 121 | POLD1 | Pold1 | 14198 | -1.754 | -0.2917 | Yes | ||
| 122 | POLR3H | Polr3h | 14239 | -1.790 | -0.2832 | Yes | ||
| 123 | ITPA | Itpa | 14268 | -1.822 | -0.2738 | Yes | ||
| 124 | POLE2 | Pole2 | 14290 | -1.839 | -0.2639 | Yes | ||
| 125 | PDE3B | Pde3b | 14337 | -1.891 | -0.2553 | Yes | ||
| 126 | AK2 | Ak2 | 14513 | -2.122 | -0.2536 | Yes | ||
| 127 | IMPDH2 | Impdh2 | 14580 | -2.224 | -0.2442 | Yes | ||
| 128 | ADSS1 | Adssl1 | 14796 | -2.558 | -0.2424 | Yes | ||
| 129 | PPAT | Ppat | 14814 | -2.607 | -0.2275 | Yes | ||
| 130 | ADCY7 | Adcy7 | 14838 | -2.643 | -0.2127 | Yes | ||
| 131 | DCK | Dck | 14890 | -2.741 | -0.1992 | Yes | ||
| 132 | PRIM2 | Prim2 | 14956 | -2.884 | -0.1857 | Yes | ||
| 133 | GDA | Gda | 15035 | -3.050 | -0.1720 | Yes | ||
| 134 | POLA1 | Pola1 | 15115 | -3.254 | -0.1571 | Yes | ||
| 135 | ADCY8 | Adcy8 | 15131 | -3.320 | -0.1376 | Yes | ||
| 136 | POLE | Pole | 15155 | -3.370 | -0.1184 | Yes | ||
| 137 | RRM1 | Rrm1 | 15164 | -3.410 | -0.0979 | Yes | ||
| 138 | ENPP3 | Enpp3 | 15180 | -3.458 | -0.0776 | Yes | ||
| 139 | PDE7B | Pde7b | 15198 | -3.513 | -0.0571 | Yes | ||
| 140 | PDE1A | Pde1a | 15393 | -4.119 | -0.0444 | Yes | ||
| 141 | RRM2 | Rrm2 | 15407 | -4.177 | -0.0195 | Yes | ||
| 142 | ENPP1 | Enpp1 | 15473 | -4.411 | 0.0034 | Yes |