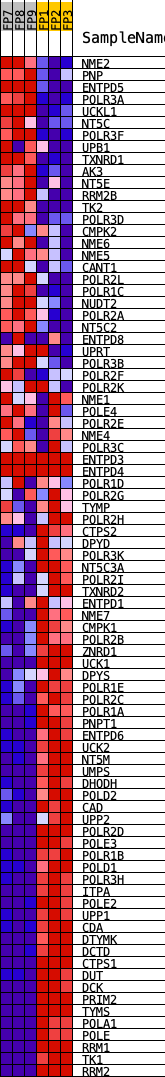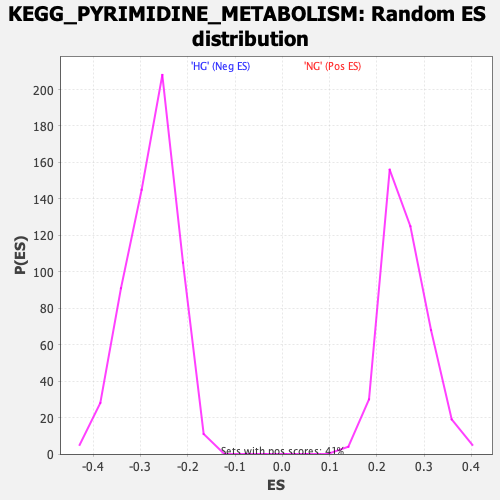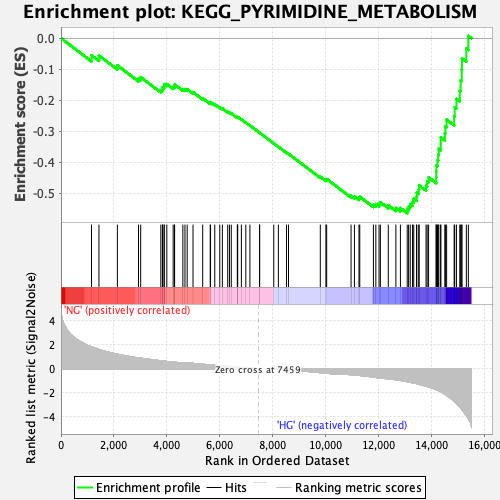
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_PYRIMIDINE_METABOLISM |
| Enrichment Score (ES) | -0.56445205 |
| Normalized Enrichment Score (NES) | -2.04666 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NME2 | Nme2 | 1154 | 1.809 | -0.0550 | No | ||
| 2 | PNP | Pnp | 1435 | 1.591 | -0.0558 | No | ||
| 3 | ENTPD5 | Entpd5 | 2132 | 1.201 | -0.0877 | No | ||
| 4 | POLR3A | Polr3a | 2931 | 0.886 | -0.1297 | No | ||
| 5 | UCKL1 | Uckl1 | 3014 | 0.865 | -0.1256 | No | ||
| 6 | NT5C | Nt5c | 3776 | 0.657 | -0.1677 | No | ||
| 7 | POLR3F | Polr3f | 3837 | 0.640 | -0.1646 | No | ||
| 8 | UPB1 | Upb1 | 3841 | 0.638 | -0.1578 | No | ||
| 9 | TXNRD1 | Txnrd1 | 3900 | 0.624 | -0.1548 | No | ||
| 10 | AK3 | Ak3 | 3903 | 0.624 | -0.1481 | No | ||
| 11 | NT5E | Nt5e | 4002 | 0.598 | -0.1479 | No | ||
| 12 | RRM2B | Rrm2b | 4242 | 0.540 | -0.1575 | No | ||
| 13 | TK2 | Tk2 | 4293 | 0.532 | -0.1549 | No | ||
| 14 | POLR3D | Polr3d | 4297 | 0.532 | -0.1493 | No | ||
| 15 | CMPK2 | Cmpk2 | 4609 | 0.488 | -0.1641 | No | ||
| 16 | NME6 | Nme6 | 4688 | 0.472 | -0.1640 | No | ||
| 17 | NME5 | Nme5 | 4772 | 0.451 | -0.1645 | No | ||
| 18 | CANT1 | Cant1 | 4993 | 0.436 | -0.1740 | No | ||
| 19 | POLR2L | Polr2l | 5359 | 0.362 | -0.1937 | No | ||
| 20 | POLR1C | Polr1c | 5644 | 0.311 | -0.2087 | No | ||
| 21 | NUDT2 | Nudt2 | 5656 | 0.308 | -0.2060 | No | ||
| 22 | POLR2A | Polr2a | 5816 | 0.281 | -0.2132 | No | ||
| 23 | NT5C2 | Nt5c2 | 6005 | 0.243 | -0.2228 | No | ||
| 24 | ENTPD8 | Entpd8 | 6103 | 0.227 | -0.2266 | No | ||
| 25 | UPRT | Uprt | 6305 | 0.189 | -0.2375 | No | ||
| 26 | POLR3B | Polr3b | 6371 | 0.177 | -0.2398 | No | ||
| 27 | POLR2F | Polr2f | 6440 | 0.164 | -0.2424 | No | ||
| 28 | POLR2K | Polr2k | 6668 | 0.122 | -0.2558 | No | ||
| 29 | NME1 | Nme1 | 6678 | 0.121 | -0.2550 | No | ||
| 30 | POLE4 | Pole4 | 6682 | 0.120 | -0.2539 | No | ||
| 31 | POLR2E | Polr2e | 6822 | 0.093 | -0.2619 | No | ||
| 32 | NME4 | Nme4 | 6990 | 0.068 | -0.2720 | No | ||
| 33 | POLR3C | Polr3c | 7145 | 0.050 | -0.2814 | No | ||
| 34 | ENTPD3 | Entpd3 | 7511 | 0.000 | -0.3050 | No | ||
| 35 | ENTPD4 | Entpd4b | 7512 | 0.000 | -0.3050 | No | ||
| 36 | POLR1D | Polr1d | 8047 | -0.003 | -0.3396 | No | ||
| 37 | POLR2G | Polr2g | 8221 | -0.037 | -0.3504 | No | ||
| 38 | TYMP | Tymp | 8528 | -0.094 | -0.3692 | No | ||
| 39 | POLR2H | Polr2h | 8604 | -0.108 | -0.3729 | No | ||
| 40 | CTPS2 | Ctps2 | 9807 | -0.333 | -0.4471 | No | ||
| 41 | DPYD | Dpyd | 10019 | -0.375 | -0.4566 | No | ||
| 42 | POLR3K | Polr3k | 10048 | -0.380 | -0.4543 | No | ||
| 43 | NT5C3A | Nt5c3 | 10970 | -0.503 | -0.5085 | No | ||
| 44 | POLR2I | Polr2i | 11098 | -0.525 | -0.5110 | No | ||
| 45 | TXNRD2 | Txnrd2 | 11270 | -0.562 | -0.5159 | No | ||
| 46 | ENTPD1 | Entpd1 | 11300 | -0.569 | -0.5116 | No | ||
| 47 | NME7 | Nme7 | 11816 | -0.705 | -0.5372 | No | ||
| 48 | CMPK1 | Cmpk1 | 11908 | -0.732 | -0.5351 | No | ||
| 49 | POLR2B | Polr2b | 12029 | -0.766 | -0.5345 | No | ||
| 50 | ZNRD1 | Znrd1 | 12079 | -0.779 | -0.5292 | No | ||
| 51 | UCK1 | Uck1 | 12378 | -0.852 | -0.5392 | No | ||
| 52 | DPYS | Dpys | 12666 | -0.930 | -0.5476 | No | ||
| 53 | POLR1E | Polr1e | 12837 | -0.986 | -0.5479 | No | ||
| 54 | POLR2C | Polr2c | 13094 | -1.090 | -0.5525 | Yes | ||
| 55 | POLR1A | Polr1a | 13147 | -1.110 | -0.5438 | Yes | ||
| 56 | PNPT1 | Pnpt1 | 13209 | -1.135 | -0.5354 | Yes | ||
| 57 | ENTPD6 | Entpd6 | 13296 | -1.179 | -0.5281 | Yes | ||
| 58 | UCK2 | Uck2 | 13341 | -1.199 | -0.5178 | Yes | ||
| 59 | NT5M | Nt5m | 13450 | -1.252 | -0.5111 | Yes | ||
| 60 | UMPS | Umps | 13455 | -1.254 | -0.4977 | Yes | ||
| 61 | DHODH | Dhodh | 13532 | -1.302 | -0.4884 | Yes | ||
| 62 | POLD2 | Pold2 | 13544 | -1.311 | -0.4748 | Yes | ||
| 63 | CAD | Cad | 13807 | -1.455 | -0.4759 | Yes | ||
| 64 | UPP2 | Upp2 | 13849 | -1.487 | -0.4623 | Yes | ||
| 65 | POLR2D | Polr2d | 13903 | -1.524 | -0.4491 | Yes | ||
| 66 | POLE3 | Pole3 | 14191 | -1.751 | -0.4486 | Yes | ||
| 67 | POLR1B | Polr1b | 14194 | -1.751 | -0.4296 | Yes | ||
| 68 | POLD1 | Pold1 | 14198 | -1.754 | -0.4106 | Yes | ||
| 69 | POLR3H | Polr3h | 14239 | -1.790 | -0.3937 | Yes | ||
| 70 | ITPA | Itpa | 14268 | -1.822 | -0.3756 | Yes | ||
| 71 | POLE2 | Pole2 | 14290 | -1.839 | -0.3569 | Yes | ||
| 72 | UPP1 | Upp1 | 14362 | -1.912 | -0.3406 | Yes | ||
| 73 | CDA | Cda | 14363 | -1.912 | -0.3197 | Yes | ||
| 74 | DTYMK | Dtymk | 14518 | -2.128 | -0.3065 | Yes | ||
| 75 | DCTD | Dctd | 14538 | -2.156 | -0.2842 | Yes | ||
| 76 | CTPS1 | Ctps | 14585 | -2.233 | -0.2628 | Yes | ||
| 77 | DUT | Dut | 14869 | -2.697 | -0.2517 | Yes | ||
| 78 | DCK | Dck | 14890 | -2.741 | -0.2230 | Yes | ||
| 79 | PRIM2 | Prim2 | 14956 | -2.884 | -0.1958 | Yes | ||
| 80 | TYMS | Tyms | 15083 | -3.168 | -0.1693 | Yes | ||
| 81 | POLA1 | Pola1 | 15115 | -3.254 | -0.1358 | Yes | ||
| 82 | POLE | Pole | 15155 | -3.370 | -0.1016 | Yes | ||
| 83 | RRM1 | Rrm1 | 15164 | -3.410 | -0.0649 | Yes | ||
| 84 | TK1 | Tk1 | 15331 | -3.893 | -0.0331 | Yes | ||
| 85 | RRM2 | Rrm2 | 15407 | -4.177 | 0.0076 | Yes |