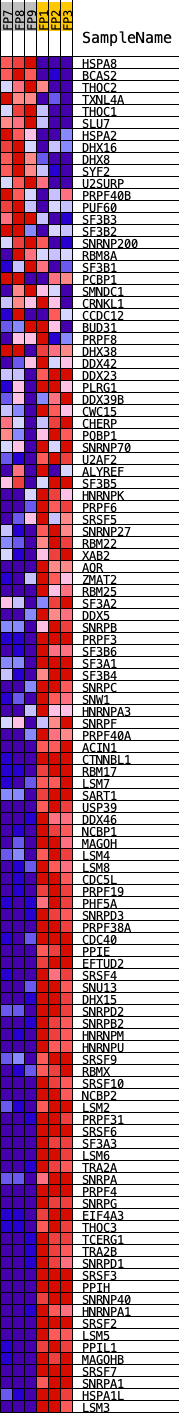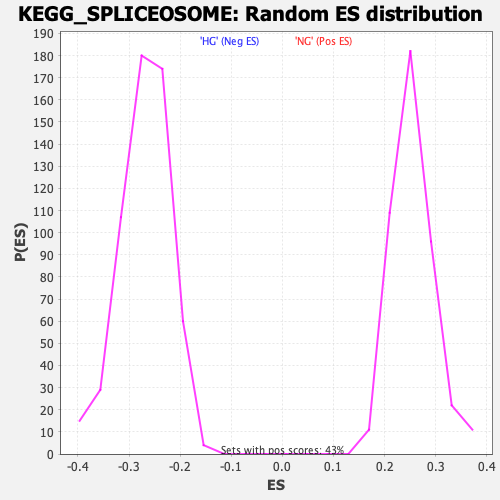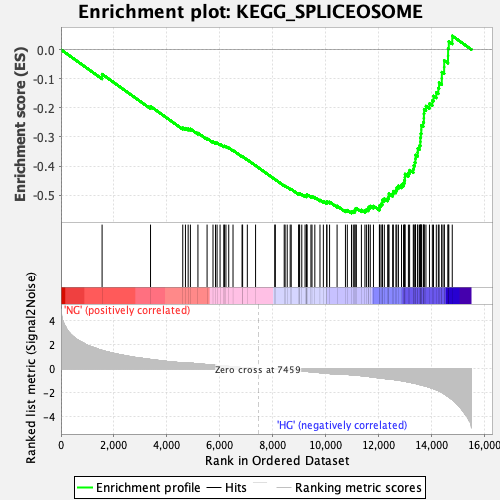
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_SPLICEOSOME |
| Enrichment Score (ES) | -0.5617211 |
| Normalized Enrichment Score (NES) | -2.091717 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HSPA8 | Hspa8 | 1554 | 1.517 | -0.0843 | No | ||
| 2 | BCAS2 | Bcas2 | 3386 | 0.757 | -0.1948 | No | ||
| 3 | THOC2 | Thoc2 | 4605 | 0.490 | -0.2685 | No | ||
| 4 | TXNL4A | Txnl4a | 4707 | 0.467 | -0.2700 | No | ||
| 5 | THOC1 | Thoc1 | 4809 | 0.443 | -0.2717 | No | ||
| 6 | SLU7 | Slu7 | 4894 | 0.440 | -0.2724 | No | ||
| 7 | HSPA2 | Hspa2 | 5179 | 0.397 | -0.2865 | No | ||
| 8 | DHX16 | Dhx16 | 5525 | 0.334 | -0.3052 | No | ||
| 9 | DHX8 | Dhx8 | 5749 | 0.291 | -0.3165 | No | ||
| 10 | SYF2 | Syf2 | 5844 | 0.277 | -0.3196 | No | ||
| 11 | U2SURP | U2surp | 5903 | 0.262 | -0.3205 | No | ||
| 12 | PRPF40B | Prpf40b | 6016 | 0.240 | -0.3251 | No | ||
| 13 | PUF60 | Puf60 | 6157 | 0.217 | -0.3318 | No | ||
| 14 | SF3B3 | Sf3b3 | 6181 | 0.214 | -0.3310 | No | ||
| 15 | SF3B2 | Sf3b2 | 6238 | 0.203 | -0.3324 | No | ||
| 16 | SNRNP200 | Snrnp200 | 6344 | 0.182 | -0.3373 | No | ||
| 17 | RBM8A | Rbm8a | 6506 | 0.150 | -0.3461 | No | ||
| 18 | SF3B1 | Sf3b1 | 6848 | 0.088 | -0.3672 | No | ||
| 19 | PCBP1 | Pcbp1 | 6869 | 0.085 | -0.3676 | No | ||
| 20 | SMNDC1 | Smndc1 | 7047 | 0.060 | -0.3784 | No | ||
| 21 | CRNKL1 | Crnkl1 | 7359 | 0.015 | -0.3984 | No | ||
| 22 | CCDC12 | Ccdc12 | 8084 | -0.010 | -0.4453 | No | ||
| 23 | BUD31 | Bud31 | 8106 | -0.014 | -0.4465 | No | ||
| 24 | PRPF8 | Prpf8 | 8442 | -0.078 | -0.4674 | No | ||
| 25 | DHX38 | Dhx38 | 8476 | -0.085 | -0.4686 | No | ||
| 26 | DDX42 | Ddx42 | 8556 | -0.099 | -0.4727 | No | ||
| 27 | DDX23 | Ddx23 | 8668 | -0.120 | -0.4786 | No | ||
| 28 | PLRG1 | Plrg1 | 8707 | -0.127 | -0.4797 | No | ||
| 29 | DDX39B | Ddx39b | 8987 | -0.173 | -0.4959 | No | ||
| 30 | CWC15 | Cwc15 | 9005 | -0.176 | -0.4950 | No | ||
| 31 | CHERP | Cherp | 9026 | -0.181 | -0.4944 | No | ||
| 32 | PQBP1 | Pqbp1 | 9107 | -0.198 | -0.4974 | No | ||
| 33 | SNRNP70 | Snrnp70 | 9231 | -0.217 | -0.5030 | No | ||
| 34 | U2AF2 | U2af2 | 9281 | -0.227 | -0.5037 | No | ||
| 35 | ALYREF | 4931428L18Rik | 9284 | -0.228 | -0.5014 | No | ||
| 36 | SF3B5 | Sf3b5 | 9294 | -0.230 | -0.4994 | No | ||
| 37 | HNRNPK | Hnrnpk | 9306 | -0.232 | -0.4976 | No | ||
| 38 | PRPF6 | Prpf6 | 9453 | -0.260 | -0.5043 | No | ||
| 39 | SRSF5 | Srsf5 | 9493 | -0.269 | -0.5039 | No | ||
| 40 | SNRNP27 | Snrnp27 | 9600 | -0.292 | -0.5076 | No | ||
| 41 | RBM22 | Rbm22 | 9795 | -0.329 | -0.5166 | No | ||
| 42 | XAB2 | Xab2 | 9923 | -0.358 | -0.5209 | No | ||
| 43 | AQR | Aqr | 10051 | -0.381 | -0.5250 | No | ||
| 44 | ZMAT2 | Zmat2 | 10055 | -0.382 | -0.5210 | No | ||
| 45 | RBM25 | Rbm25 | 10158 | -0.405 | -0.5232 | No | ||
| 46 | SF3A2 | Sf3a2 | 10442 | -0.425 | -0.5370 | No | ||
| 47 | DDX5 | Ddx5 | 10758 | -0.469 | -0.5523 | No | ||
| 48 | SNRPB | Snrpb | 10829 | -0.485 | -0.5515 | No | ||
| 49 | PRPF3 | Prpf3 | 10987 | -0.507 | -0.5562 | Yes | ||
| 50 | SF3B6 | Sf3b6 | 11055 | -0.518 | -0.5549 | Yes | ||
| 51 | SF3A1 | Sf3a1 | 11115 | -0.528 | -0.5530 | Yes | ||
| 52 | SF3B4 | Sf3b4 | 11122 | -0.530 | -0.5476 | Yes | ||
| 53 | SNRPC | Snrpc | 11171 | -0.539 | -0.5448 | Yes | ||
| 54 | SNW1 | Snw1 | 11362 | -0.581 | -0.5508 | Yes | ||
| 55 | HNRNPA3 | Gm17190 | 11495 | -0.613 | -0.5527 | Yes | ||
| 56 | SNRPF | Snrpf | 11551 | -0.630 | -0.5494 | Yes | ||
| 57 | PRPF40A | Prpf40a | 11625 | -0.648 | -0.5471 | Yes | ||
| 58 | ACIN1 | Acin1 | 11637 | -0.652 | -0.5407 | Yes | ||
| 59 | CTNNBL1 | Ctnnbl1 | 11698 | -0.670 | -0.5373 | Yes | ||
| 60 | RBM17 | Rbm17 | 11818 | -0.706 | -0.5373 | Yes | ||
| 61 | LSM7 | Lsm7 | 12042 | -0.767 | -0.5434 | Yes | ||
| 62 | SART1 | Sart1 | 12049 | -0.769 | -0.5354 | Yes | ||
| 63 | USP39 | Usp39 | 12104 | -0.785 | -0.5304 | Yes | ||
| 64 | DDX46 | Ddx46 | 12154 | -0.798 | -0.5248 | Yes | ||
| 65 | NCBP1 | Ncbp1 | 12156 | -0.799 | -0.5162 | Yes | ||
| 66 | MAGOH | Magoh | 12229 | -0.815 | -0.5120 | Yes | ||
| 67 | LSM4 | Lsm4 | 12356 | -0.850 | -0.5109 | Yes | ||
| 68 | LSM8 | Lsm8 | 12395 | -0.858 | -0.5040 | Yes | ||
| 69 | CDC5L | Cdc5l | 12402 | -0.860 | -0.4950 | Yes | ||
| 70 | PRPF19 | Prpf19 | 12544 | -0.891 | -0.4945 | Yes | ||
| 71 | PHF5A | Phf5a | 12570 | -0.899 | -0.4863 | Yes | ||
| 72 | SNRPD3 | Snrpd3 | 12668 | -0.931 | -0.4824 | Yes | ||
| 73 | PRPF38A | Prpf38a | 12700 | -0.941 | -0.4742 | Yes | ||
| 74 | CDC40 | Cdc40 | 12762 | -0.961 | -0.4677 | Yes | ||
| 75 | PPIE | Ppie | 12879 | -1.001 | -0.4643 | Yes | ||
| 76 | EFTUD2 | Eftud2 | 12956 | -1.029 | -0.4580 | Yes | ||
| 77 | SRSF4 | Srsf4 | 12990 | -1.038 | -0.4488 | Yes | ||
| 78 | SNU13 | Snu13 | 13006 | -1.047 | -0.4384 | Yes | ||
| 79 | DHX15 | Dhx15 | 13011 | -1.049 | -0.4272 | Yes | ||
| 80 | SNRPD2 | Snrpd2 | 13140 | -1.108 | -0.4234 | Yes | ||
| 81 | SNRPB2 | Snrpb2 | 13185 | -1.124 | -0.4140 | Yes | ||
| 82 | HNRNPM | Hnrnpm | 13324 | -1.190 | -0.4100 | Yes | ||
| 83 | HNRNPU | Hnrnpu | 13343 | -1.200 | -0.3981 | Yes | ||
| 84 | SRSF9 | Srsf9 | 13381 | -1.221 | -0.3872 | Yes | ||
| 85 | RBMX | Rbmx | 13406 | -1.234 | -0.3753 | Yes | ||
| 86 | SRSF10 | Srsf10 | 13410 | -1.236 | -0.3620 | Yes | ||
| 87 | NCBP2 | Ncbp2 | 13489 | -1.273 | -0.3532 | Yes | ||
| 88 | LSM2 | Lsm2 | 13494 | -1.276 | -0.3395 | Yes | ||
| 89 | PRPF31 | Prpf31 | 13561 | -1.321 | -0.3294 | Yes | ||
| 90 | SRSF6 | Srsf6 | 13590 | -1.336 | -0.3167 | Yes | ||
| 91 | SF3A3 | Sf3a3 | 13593 | -1.337 | -0.3022 | Yes | ||
| 92 | LSM6 | Lsm6 | 13610 | -1.347 | -0.2886 | Yes | ||
| 93 | TRA2A | Tra2a | 13627 | -1.353 | -0.2749 | Yes | ||
| 94 | SNRPA | Snrpa | 13628 | -1.354 | -0.2601 | Yes | ||
| 95 | PRPF4 | Prpf4 | 13702 | -1.394 | -0.2496 | Yes | ||
| 96 | SNRPG | Snrpg | 13724 | -1.411 | -0.2356 | Yes | ||
| 97 | EIF4A3 | Eif4a3 | 13727 | -1.413 | -0.2204 | Yes | ||
| 98 | THOC3 | Thoc3 | 13735 | -1.418 | -0.2054 | Yes | ||
| 99 | TCERG1 | Tcerg1 | 13803 | -1.452 | -0.1939 | Yes | ||
| 100 | TRA2B | Tra2b | 13936 | -1.555 | -0.1855 | Yes | ||
| 101 | SNRPD1 | Snrpd1 | 14045 | -1.631 | -0.1747 | Yes | ||
| 102 | SRSF3 | Srsf3 | 14088 | -1.666 | -0.1593 | Yes | ||
| 103 | PPIH | Gm7879 | 14195 | -1.753 | -0.1470 | Yes | ||
| 104 | SNRNP40 | Snrnp40 | 14271 | -1.824 | -0.1320 | Yes | ||
| 105 | HNRNPA1 | Hnrnpa1 | 14299 | -1.853 | -0.1136 | Yes | ||
| 106 | SRSF2 | Srsf2 | 14402 | -1.967 | -0.0988 | Yes | ||
| 107 | LSM5 | Lsm5 | 14405 | -1.970 | -0.0774 | Yes | ||
| 108 | PPIL1 | Ppil1 | 14491 | -2.090 | -0.0601 | Yes | ||
| 109 | MAGOHB | Magohb | 14494 | -2.091 | -0.0375 | Yes | ||
| 110 | SRSF7 | Srsf7 | 14638 | -2.310 | -0.0216 | Yes | ||
| 111 | SNRPA1 | Snrpa1 | 14641 | -2.317 | 0.0035 | Yes | ||
| 112 | HSPA1L | Hspa1l | 14668 | -2.365 | 0.0276 | Yes | ||
| 113 | LSM3 | Lsm3 | 14798 | -2.560 | 0.0472 | Yes |