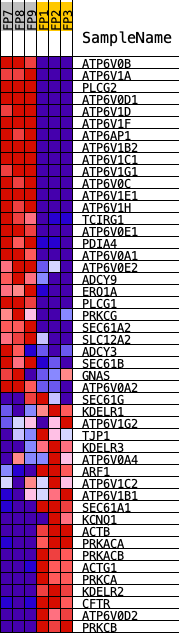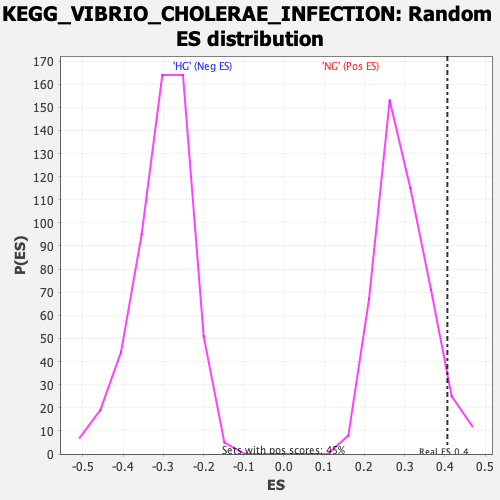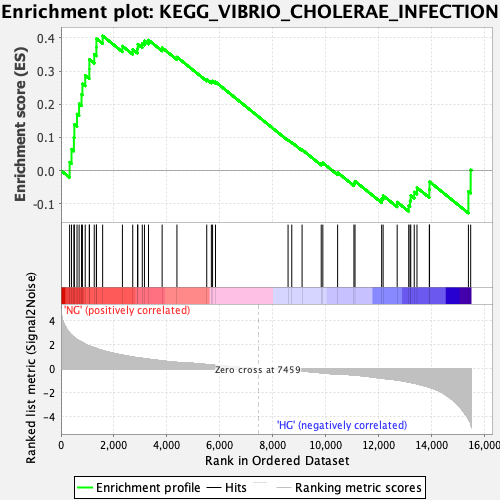
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_HG.phenotype.cls#NG_versus_HG_repos |
| Phenotype | phenotype.cls#NG_versus_HG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_VIBRIO_CHOLERAE_INFECTION |
| Enrichment Score (ES) | 0.40636224 |
| Normalized Enrichment Score (NES) | 1.3745201 |
| Nominal p-value | 0.05764967 |
| FDR q-value | 0.25979942 |
| FWER p-Value | 0.989 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP6V0B | Atp6v0b | 326 | 2.960 | 0.0252 | Yes | ||
| 2 | ATP6V1A | Atp6v1a | 398 | 2.800 | 0.0644 | Yes | ||
| 3 | PLCG2 | Plcg2 | 488 | 2.619 | 0.0996 | Yes | ||
| 4 | ATP6V0D1 | Atp6v0d1 | 505 | 2.590 | 0.1390 | Yes | ||
| 5 | ATP6V1D | Atp6v1d | 606 | 2.422 | 0.1704 | Yes | ||
| 6 | ATP6V1F | Atp6v1f | 685 | 2.295 | 0.2013 | Yes | ||
| 7 | ATP6AP1 | Atp6ap1 | 777 | 2.216 | 0.2300 | Yes | ||
| 8 | ATP6V1B2 | Atp6v1b2 | 810 | 2.172 | 0.2619 | Yes | ||
| 9 | ATP6V1C1 | Atp6v1c1 | 916 | 2.032 | 0.2869 | Yes | ||
| 10 | ATP6V1G1 | Atp6v1g1 | 1072 | 1.876 | 0.3062 | Yes | ||
| 11 | ATP6V0C | Atp6v0c | 1074 | 1.875 | 0.3355 | Yes | ||
| 12 | ATP6V1E1 | Atp6v1e1 | 1257 | 1.740 | 0.3509 | Yes | ||
| 13 | ATP6V1H | Atp6v1h | 1338 | 1.675 | 0.3719 | Yes | ||
| 14 | TCIRG1 | Tcirg1 | 1341 | 1.672 | 0.3979 | Yes | ||
| 15 | ATP6V0E1 | Atp6v0e | 1575 | 1.503 | 0.4064 | Yes | ||
| 16 | PDIA4 | Pdia4 | 2323 | 1.121 | 0.3756 | No | ||
| 17 | ATP6V0A1 | Atp6v0a1 | 2713 | 0.963 | 0.3656 | No | ||
| 18 | ATP6V0E2 | Atp6v0e2 | 2895 | 0.900 | 0.3679 | No | ||
| 19 | ADCY9 | Adcy9 | 2906 | 0.895 | 0.3813 | No | ||
| 20 | ERO1A | Ero1l | 3074 | 0.851 | 0.3838 | No | ||
| 21 | PLCG1 | Plcg1 | 3156 | 0.826 | 0.3915 | No | ||
| 22 | PRKCG | Prkcg | 3309 | 0.781 | 0.3939 | No | ||
| 23 | SEC61A2 | Sec61a2 | 3825 | 0.643 | 0.3706 | No | ||
| 24 | SLC12A2 | Slc12a2 | 4385 | 0.515 | 0.3426 | No | ||
| 25 | ADCY3 | Adcy3 | 5512 | 0.336 | 0.2751 | No | ||
| 26 | SEC61B | Gm10320 | 5687 | 0.302 | 0.2686 | No | ||
| 27 | GNAS | Gnas | 5733 | 0.294 | 0.2703 | No | ||
| 28 | ATP6V0A2 | Atp6v0a2 | 5843 | 0.277 | 0.2675 | No | ||
| 29 | SEC61G | Sec61g | 8588 | -0.106 | 0.0919 | No | ||
| 30 | KDELR1 | Kdelr1 | 8728 | -0.130 | 0.0850 | No | ||
| 31 | ATP6V1G2 | Atp6v1g2 | 9118 | -0.199 | 0.0629 | No | ||
| 32 | TJP1 | Tjp1 | 9842 | -0.340 | 0.0215 | No | ||
| 33 | KDELR3 | Kdelr3 | 9903 | -0.354 | 0.0232 | No | ||
| 34 | ATP6V0A4 | Atp6v0a4 | 10462 | -0.428 | -0.0062 | No | ||
| 35 | ARF1 | Arf1 | 11079 | -0.521 | -0.0378 | No | ||
| 36 | ATP6V1C2 | Atp6v1c2 | 11120 | -0.529 | -0.0321 | No | ||
| 37 | ATP6V1B1 | Atp6v1b1 | 12128 | -0.791 | -0.0848 | No | ||
| 38 | SEC61A1 | Sec61a1 | 12183 | -0.806 | -0.0757 | No | ||
| 39 | KCNQ1 | Kcnq1 | 12715 | -0.946 | -0.0952 | No | ||
| 40 | ACTB | Actb | 13151 | -1.112 | -0.1059 | No | ||
| 41 | PRKACA | Prkaca | 13202 | -1.133 | -0.0915 | No | ||
| 42 | PRKACB | Prkacb | 13228 | -1.145 | -0.0752 | No | ||
| 43 | ACTG1 | Actg1 | 13360 | -1.209 | -0.0647 | No | ||
| 44 | PRKCA | Prkca | 13464 | -1.260 | -0.0517 | No | ||
| 45 | KDELR2 | Kdelr2 | 13930 | -1.549 | -0.0575 | No | ||
| 46 | CFTR | Cftr | 13939 | -1.558 | -0.0337 | No | ||
| 47 | ATP6V0D2 | Atp6v0d2 | 15409 | -4.183 | -0.0632 | No | ||
| 48 | PRKCB | Prkcb | 15495 | -4.518 | 0.0019 | No |