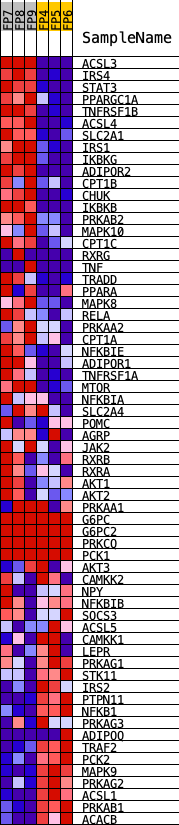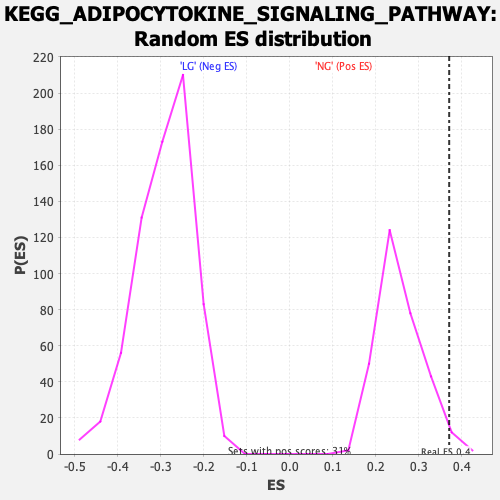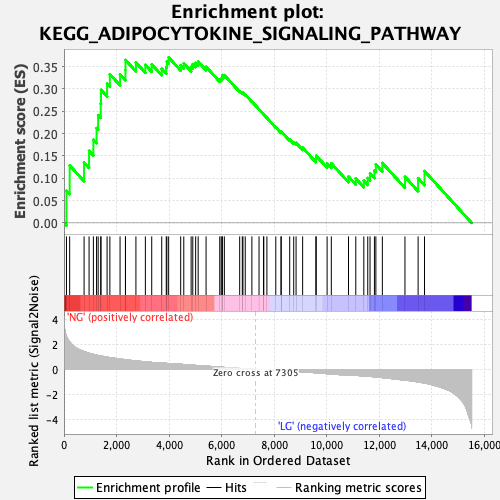
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_ADIPOCYTOKINE_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.3705259 |
| Normalized Enrichment Score (NES) | 1.4453499 |
| Nominal p-value | 0.025723472 |
| FDR q-value | 0.24103059 |
| FWER p-Value | 0.884 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ACSL3 | Acsl3 | 94 | 2.653 | 0.0720 | Yes | ||
| 2 | IRS4 | Irs4 | 216 | 2.191 | 0.1286 | Yes | ||
| 3 | STAT3 | Stat3 | 765 | 1.428 | 0.1352 | Yes | ||
| 4 | PPARGC1A | Ppargc1a | 955 | 1.305 | 0.1614 | Yes | ||
| 5 | TNFRSF1B | Tnfrsf1b | 1119 | 1.207 | 0.1864 | Yes | ||
| 6 | ACSL4 | Acsl4 | 1240 | 1.143 | 0.2122 | Yes | ||
| 7 | SLC2A1 | Slc2a1 | 1301 | 1.118 | 0.2413 | Yes | ||
| 8 | IRS1 | Irs1 | 1399 | 1.080 | 0.2667 | Yes | ||
| 9 | IKBKG | Ikbkg | 1409 | 1.077 | 0.2979 | Yes | ||
| 10 | ADIPOR2 | Adipor2 | 1641 | 0.984 | 0.3119 | Yes | ||
| 11 | CPT1B | Cpt1b | 1747 | 0.948 | 0.3330 | Yes | ||
| 12 | CHUK | Chuk | 2135 | 0.833 | 0.3325 | Yes | ||
| 13 | IKBKB | Ikbkb | 2340 | 0.774 | 0.3421 | Yes | ||
| 14 | PRKAB2 | Prkab2 | 2342 | 0.774 | 0.3648 | Yes | ||
| 15 | MAPK10 | Mapk10 | 2739 | 0.681 | 0.3592 | Yes | ||
| 16 | CPT1C | Cpt1c | 3099 | 0.610 | 0.3539 | Yes | ||
| 17 | RXRG | Rxrg | 3341 | 0.563 | 0.3549 | Yes | ||
| 18 | TNF | Tnf | 3722 | 0.501 | 0.3451 | Yes | ||
| 19 | TRADD | Tradd | 3894 | 0.487 | 0.3484 | Yes | ||
| 20 | PPARA | Ppara | 3918 | 0.483 | 0.3611 | Yes | ||
| 21 | MAPK8 | Mapk8 | 3986 | 0.468 | 0.3705 | Yes | ||
| 22 | RELA | Rela | 4444 | 0.415 | 0.3532 | No | ||
| 23 | PRKAA2 | Prkaa2 | 4562 | 0.396 | 0.3573 | No | ||
| 24 | CPT1A | Cpt1a | 4842 | 0.352 | 0.3496 | No | ||
| 25 | NFKBIE | Nfkbie | 4905 | 0.341 | 0.3556 | No | ||
| 26 | ADIPOR1 | Adipor1 | 5014 | 0.323 | 0.3581 | No | ||
| 27 | TNFRSF1A | Tnfrsf1a | 5110 | 0.311 | 0.3611 | No | ||
| 28 | MTOR | Mtor | 5411 | 0.266 | 0.3496 | No | ||
| 29 | NFKBIA | Nfkbia | 5932 | 0.188 | 0.3215 | No | ||
| 30 | SLC2A4 | Slc2a4 | 5987 | 0.179 | 0.3232 | No | ||
| 31 | POMC | Pomc | 6025 | 0.175 | 0.3260 | No | ||
| 32 | AGRP | Agrp | 6030 | 0.175 | 0.3309 | No | ||
| 33 | JAK2 | Jak2 | 6107 | 0.162 | 0.3307 | No | ||
| 34 | RXRB | Rxrb | 6691 | 0.079 | 0.2954 | No | ||
| 35 | RXRA | Rxra | 6791 | 0.070 | 0.2910 | No | ||
| 36 | AKT1 | Akt1 | 6815 | 0.066 | 0.2915 | No | ||
| 37 | AKT2 | Akt2 | 6903 | 0.054 | 0.2874 | No | ||
| 38 | PRKAA1 | Prkaa1 | 7156 | 0.021 | 0.2717 | No | ||
| 39 | G6PC | G6pc | 7428 | 0.000 | 0.2542 | No | ||
| 40 | G6PC2 | G6pc2 | 7598 | 0.000 | 0.2433 | No | ||
| 41 | PRKCQ | Prkcq | 7610 | 0.000 | 0.2426 | No | ||
| 42 | PCK1 | Pck1 | 7728 | 0.000 | 0.2350 | No | ||
| 43 | AKT3 | Akt3 | 8066 | -0.032 | 0.2142 | No | ||
| 44 | CAMKK2 | Camkk2 | 8267 | -0.063 | 0.2031 | No | ||
| 45 | NPY | Npy | 8271 | -0.063 | 0.2047 | No | ||
| 46 | NFKBIB | Nfkbib | 8596 | -0.107 | 0.1869 | No | ||
| 47 | SOCS3 | Socs3 | 8749 | -0.128 | 0.1808 | No | ||
| 48 | ACSL5 | Acsl5 | 8839 | -0.141 | 0.1792 | No | ||
| 49 | CAMKK1 | Camkk1 | 9089 | -0.181 | 0.1685 | No | ||
| 50 | LEPR | Lepr | 9593 | -0.257 | 0.1435 | No | ||
| 51 | PRKAG1 | Prkag1 | 9606 | -0.259 | 0.1504 | No | ||
| 52 | STK11 | Stk11 | 10021 | -0.328 | 0.1332 | No | ||
| 53 | IRS2 | Irs2 | 10181 | -0.352 | 0.1333 | No | ||
| 54 | PTPN11 | Ptpn11 | 10836 | -0.428 | 0.1036 | No | ||
| 55 | NFKB1 | Nfkb1 | 11113 | -0.463 | 0.0994 | No | ||
| 56 | PRKAG3 | Prkag3 | 11419 | -0.506 | 0.0946 | No | ||
| 57 | ADIPOQ | Adipoq | 11569 | -0.528 | 0.1005 | No | ||
| 58 | TRAF2 | Traf2 | 11656 | -0.547 | 0.1110 | No | ||
| 59 | PCK2 | Pck2 | 11823 | -0.578 | 0.1173 | No | ||
| 60 | MAPK9 | Mapk9 | 11878 | -0.588 | 0.1311 | No | ||
| 61 | PRKAG2 | Prkag2 | 12125 | -0.643 | 0.1341 | No | ||
| 62 | ACSL1 | Acsl1 | 12986 | -0.840 | 0.1032 | No | ||
| 63 | PRKAB1 | Prkab1 | 13487 | -0.990 | 0.1000 | No | ||
| 64 | ACACB | Acacb | 13730 | -1.078 | 0.1161 | No |