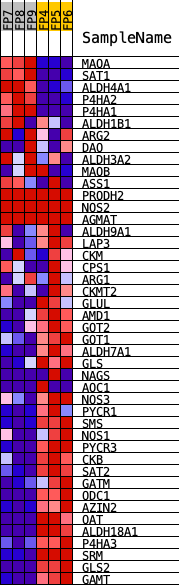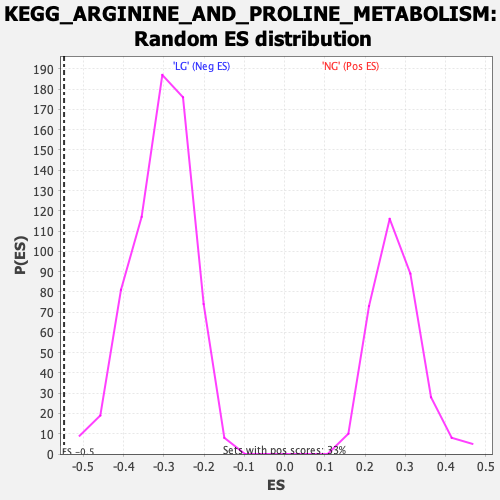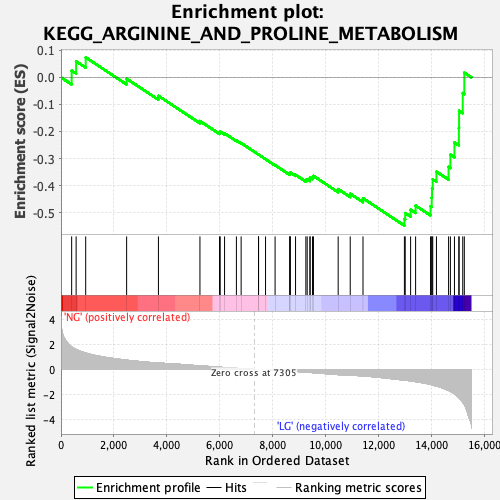
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_ARGININE_AND_PROLINE_METABOLISM |
| Enrichment Score (ES) | -0.54814935 |
| Normalized Enrichment Score (NES) | -1.7915545 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.00907249 |
| FWER p-Value | 0.122 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MAOA | Maoa | 403 | 1.794 | 0.0251 | No | ||
| 2 | SAT1 | Sat1 | 572 | 1.593 | 0.0596 | No | ||
| 3 | ALDH4A1 | Aldh4a1 | 934 | 1.318 | 0.0738 | No | ||
| 4 | P4HA2 | P4ha2 | 2482 | 0.737 | -0.0051 | No | ||
| 5 | P4HA1 | P4ha1 | 3685 | 0.506 | -0.0683 | No | ||
| 6 | ALDH1B1 | Aldh1b1 | 5255 | 0.288 | -0.1614 | No | ||
| 7 | ARG2 | Arg2 | 5996 | 0.178 | -0.2041 | No | ||
| 8 | DAO | Dao | 6021 | 0.176 | -0.2007 | No | ||
| 9 | ALDH3A2 | Aldh3a2 | 6186 | 0.151 | -0.2070 | No | ||
| 10 | MAOB | Maob | 6634 | 0.087 | -0.2334 | No | ||
| 11 | ASS1 | Ass1 | 6813 | 0.066 | -0.2430 | No | ||
| 12 | PRODH2 | Prodh2 | 7471 | 0.000 | -0.2854 | No | ||
| 13 | NOS2 | Nos2 | 7472 | 0.000 | -0.2854 | No | ||
| 14 | AGMAT | Agmat | 7736 | 0.000 | -0.3024 | No | ||
| 15 | ALDH9A1 | Aldh9a1 | 8097 | -0.037 | -0.3246 | No | ||
| 16 | LAP3 | Lap3 | 8652 | -0.115 | -0.3571 | No | ||
| 17 | CKM | Ckm | 8657 | -0.115 | -0.3541 | No | ||
| 18 | CPS1 | Cps1 | 8673 | -0.118 | -0.3517 | No | ||
| 19 | ARG1 | Arg1 | 8871 | -0.147 | -0.3603 | No | ||
| 20 | CKMT2 | Ckmt2 | 9261 | -0.206 | -0.3795 | No | ||
| 21 | GLUL | Glul | 9311 | -0.214 | -0.3766 | No | ||
| 22 | AMD1 | Amd1 | 9418 | -0.229 | -0.3769 | No | ||
| 23 | GOT2 | Got2 | 9419 | -0.230 | -0.3703 | No | ||
| 24 | GOT1 | Got1 | 9513 | -0.245 | -0.3694 | No | ||
| 25 | ALDH7A1 | Aldh7a1 | 9548 | -0.250 | -0.3644 | No | ||
| 26 | GLS | Gls | 10483 | -0.404 | -0.4133 | No | ||
| 27 | NAGS | Nags | 10939 | -0.432 | -0.4304 | No | ||
| 28 | AOC1 | Aoc1 | 11422 | -0.506 | -0.4471 | No | ||
| 29 | NOS3 | Nos3 | 12988 | -0.841 | -0.5242 | Yes | ||
| 30 | PYCR1 | Pycr1 | 13019 | -0.849 | -0.5019 | Yes | ||
| 31 | SMS | Sms | 13223 | -0.903 | -0.4893 | Yes | ||
| 32 | NOS1 | Nos1 | 13412 | -0.963 | -0.4740 | Yes | ||
| 33 | PYCR3 | Pycrl | 13977 | -1.190 | -0.4766 | Yes | ||
| 34 | CKB | Ckb | 14021 | -1.211 | -0.4448 | Yes | ||
| 35 | SAT2 | Sat2 | 14033 | -1.221 | -0.4107 | Yes | ||
| 36 | GATM | Gatm | 14060 | -1.235 | -0.3772 | Yes | ||
| 37 | ODC1 | Odc1 | 14200 | -1.319 | -0.3486 | Yes | ||
| 38 | AZIN2 | Azin2 | 14659 | -1.663 | -0.3308 | Yes | ||
| 39 | OAT | Oat | 14728 | -1.737 | -0.2857 | Yes | ||
| 40 | ALDH18A1 | Aldh18a1 | 14881 | -1.931 | -0.2405 | Yes | ||
| 41 | P4HA3 | P4ha3 | 15048 | -2.242 | -0.1874 | Yes | ||
| 42 | SRM | Srm | 15056 | -2.257 | -0.1235 | Yes | ||
| 43 | GLS2 | Gls2 | 15195 | -2.610 | -0.0580 | Yes | ||
| 44 | GAMT | Gamt | 15254 | -2.783 | 0.0175 | Yes |