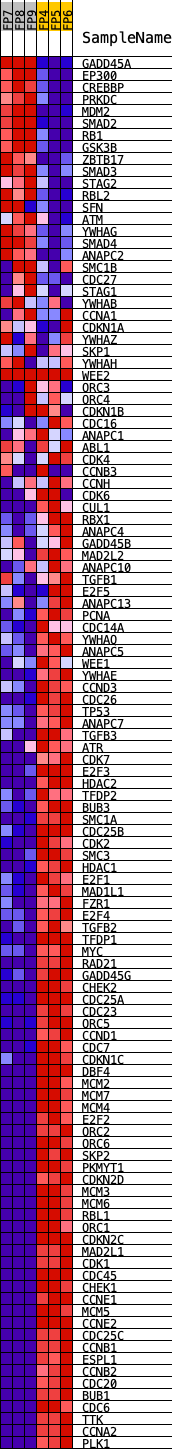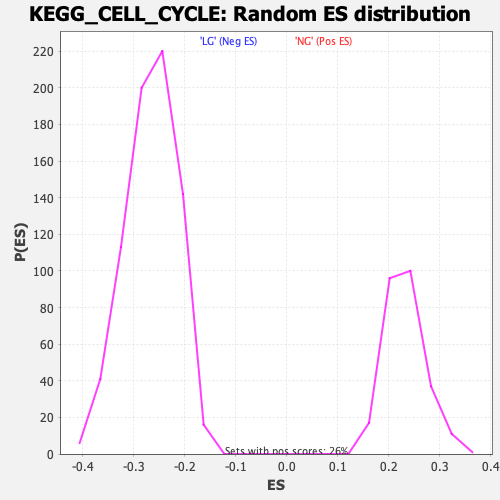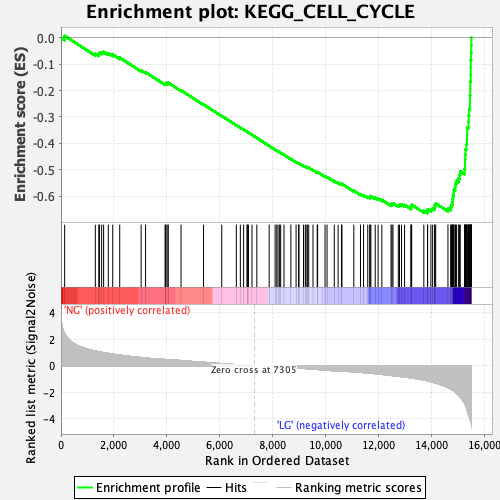
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.66441554 |
| Normalized Enrichment Score (NES) | -2.4965672 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GADD45A | Gadd45a | 138 | 2.445 | 0.0072 | No | ||
| 2 | EP300 | Ep300 | 1298 | 1.118 | -0.0606 | No | ||
| 3 | CREBBP | Crebbp | 1430 | 1.067 | -0.0621 | No | ||
| 4 | PRKDC | Prkdc | 1452 | 1.053 | -0.0565 | No | ||
| 5 | MDM2 | Mdm2 | 1537 | 1.023 | -0.0552 | No | ||
| 6 | SMAD2 | Smad2 | 1612 | 0.997 | -0.0534 | No | ||
| 7 | RB1 | Rb1 | 1789 | 0.937 | -0.0586 | No | ||
| 8 | GSK3B | Gsk3b | 1956 | 0.881 | -0.0636 | No | ||
| 9 | ZBTB17 | Zbtb17 | 2221 | 0.808 | -0.0754 | No | ||
| 10 | SMAD3 | Smad3 | 3030 | 0.624 | -0.1237 | No | ||
| 11 | STAG2 | Stag2 | 3198 | 0.591 | -0.1306 | No | ||
| 12 | RBL2 | Rbl2 | 3936 | 0.479 | -0.1753 | No | ||
| 13 | SFN | Sfn | 3965 | 0.473 | -0.1740 | No | ||
| 14 | ATM | Atm | 3970 | 0.472 | -0.1711 | No | ||
| 15 | YWHAG | Ywhag | 4047 | 0.457 | -0.1730 | No | ||
| 16 | SMAD4 | Smad4 | 4050 | 0.456 | -0.1701 | No | ||
| 17 | ANAPC2 | Anapc2 | 4539 | 0.399 | -0.1992 | No | ||
| 18 | SMC1B | Smc1b | 5391 | 0.269 | -0.2526 | No | ||
| 19 | CDC27 | Cdc27 | 6079 | 0.168 | -0.2961 | No | ||
| 20 | STAG1 | Stag1 | 6631 | 0.087 | -0.3313 | No | ||
| 21 | YWHAB | Ywhab | 6783 | 0.071 | -0.3406 | No | ||
| 22 | CCNA1 | Ccna1 | 6907 | 0.054 | -0.3482 | No | ||
| 23 | CDKN1A | Cdkn1a | 7037 | 0.037 | -0.3564 | No | ||
| 24 | YWHAZ | Ywhaz | 7048 | 0.035 | -0.3568 | No | ||
| 25 | SKP1 | Skp1a | 7090 | 0.030 | -0.3592 | No | ||
| 26 | YWHAH | Ywhah | 7223 | 0.013 | -0.3677 | No | ||
| 27 | WEE2 | Wee2 | 7407 | 0.000 | -0.3796 | No | ||
| 28 | ORC3 | Orc3 | 7874 | -0.004 | -0.4098 | No | ||
| 29 | ORC4 | Orc4 | 8098 | -0.037 | -0.4240 | No | ||
| 30 | CDKN1B | Cdkn1b | 8161 | -0.049 | -0.4277 | No | ||
| 31 | CDC16 | Cdc16 | 8209 | -0.055 | -0.4304 | No | ||
| 32 | ANAPC1 | Anapc1 | 8269 | -0.063 | -0.4338 | No | ||
| 33 | ABL1 | Abl1 | 8308 | -0.068 | -0.4358 | No | ||
| 34 | CDK4 | Cdk4 | 8434 | -0.086 | -0.4434 | No | ||
| 35 | CCNB3 | Ccnb3 | 8694 | -0.120 | -0.4594 | No | ||
| 36 | CCNH | Ccnh | 8891 | -0.150 | -0.4711 | No | ||
| 37 | CDK6 | Cdk6 | 8988 | -0.167 | -0.4763 | No | ||
| 38 | CUL1 | Cul1 | 9002 | -0.169 | -0.4760 | No | ||
| 39 | RBX1 | Rbx1 | 9172 | -0.192 | -0.4857 | No | ||
| 40 | ANAPC4 | Anapc4 | 9239 | -0.203 | -0.4886 | No | ||
| 41 | GADD45B | Gadd45b | 9269 | -0.208 | -0.4891 | No | ||
| 42 | MAD2L2 | Mad2l2 | 9318 | -0.215 | -0.4908 | No | ||
| 43 | ANAPC10 | Anapc10 | 9369 | -0.222 | -0.4926 | No | ||
| 44 | TGFB1 | Tgfb1 | 9531 | -0.248 | -0.5014 | No | ||
| 45 | E2F5 | E2f5 | 9690 | -0.273 | -0.5099 | No | ||
| 46 | ANAPC13 | Anapc13 | 9706 | -0.275 | -0.5090 | No | ||
| 47 | PCNA | Pcna | 9987 | -0.323 | -0.5251 | No | ||
| 48 | CDC14A | Cdc14a | 10062 | -0.335 | -0.5277 | No | ||
| 49 | YWHAQ | Ywhaq | 10336 | -0.378 | -0.5429 | No | ||
| 50 | ANAPC5 | Anapc5 | 10481 | -0.403 | -0.5496 | No | ||
| 51 | WEE1 | Wee1 | 10607 | -0.407 | -0.5550 | No | ||
| 52 | YWHAE | Ywhae | 10626 | -0.410 | -0.5535 | No | ||
| 53 | CCND3 | Ccnd3 | 11075 | -0.456 | -0.5795 | No | ||
| 54 | CDC26 | Cdc26 | 11327 | -0.492 | -0.5925 | No | ||
| 55 | TP53 | Trp53 | 11444 | -0.509 | -0.5967 | No | ||
| 56 | ANAPC7 | Anapc7 | 11602 | -0.536 | -0.6034 | No | ||
| 57 | TGFB3 | Tgfb3 | 11679 | -0.551 | -0.6047 | No | ||
| 58 | ATR | Atr | 11688 | -0.552 | -0.6015 | No | ||
| 59 | CDK7 | Cdk7 | 11720 | -0.558 | -0.5998 | No | ||
| 60 | E2F3 | E2f3 | 11884 | -0.589 | -0.6065 | No | ||
| 61 | HDAC2 | Hdac2 | 11993 | -0.613 | -0.6095 | No | ||
| 62 | TFDP2 | Tfdp2 | 12135 | -0.645 | -0.6144 | No | ||
| 63 | BUB3 | Bub3 | 12483 | -0.725 | -0.6321 | No | ||
| 64 | SMC1A | Smc1a | 12518 | -0.735 | -0.6295 | No | ||
| 65 | CDC25B | Cdc25b | 12556 | -0.742 | -0.6270 | No | ||
| 66 | CDK2 | Cdk2 | 12758 | -0.793 | -0.6348 | No | ||
| 67 | SMC3 | Smc3 | 12802 | -0.803 | -0.6322 | No | ||
| 68 | HDAC1 | Hdac1 | 12879 | -0.822 | -0.6317 | No | ||
| 69 | E2F1 | E2f1 | 12995 | -0.843 | -0.6336 | No | ||
| 70 | MAD1L1 | Mad1l1 | 13237 | -0.908 | -0.6433 | No | ||
| 71 | FZR1 | Fzr1 | 13239 | -0.909 | -0.6373 | No | ||
| 72 | E2F4 | E2f4 | 13264 | -0.916 | -0.6328 | No | ||
| 73 | TGFB2 | Tgfb2 | 13723 | -1.076 | -0.6555 | No | ||
| 74 | TFDP1 | Tfdp1 | 13862 | -1.129 | -0.6570 | Yes | ||
| 75 | MYC | Myc | 13868 | -1.132 | -0.6498 | Yes | ||
| 76 | RAD21 | Rad21 | 13990 | -1.198 | -0.6497 | Yes | ||
| 77 | GADD45G | Gadd45g | 14056 | -1.231 | -0.6458 | Yes | ||
| 78 | CHEK2 | Chek2 | 14135 | -1.288 | -0.6424 | Yes | ||
| 79 | CDC25A | Cdc25a | 14138 | -1.288 | -0.6340 | Yes | ||
| 80 | CDC23 | Cdc23 | 14171 | -1.301 | -0.6275 | Yes | ||
| 81 | ORC5 | Orc5 | 14635 | -1.638 | -0.6467 | Yes | ||
| 82 | CCND1 | Ccnd1 | 14737 | -1.745 | -0.6417 | Yes | ||
| 83 | CDC7 | Cdc7 | 14770 | -1.789 | -0.6320 | Yes | ||
| 84 | CDKN1C | Cdkn1c | 14804 | -1.837 | -0.6220 | Yes | ||
| 85 | DBF4 | Dbf4 | 14815 | -1.845 | -0.6105 | Yes | ||
| 86 | MCM2 | Mcm2 | 14826 | -1.855 | -0.5988 | Yes | ||
| 87 | MCM7 | Mcm7 | 14857 | -1.897 | -0.5883 | Yes | ||
| 88 | MCM4 | Mcm4 | 14859 | -1.899 | -0.5758 | Yes | ||
| 89 | E2F2 | E2f2 | 14911 | -1.981 | -0.5660 | Yes | ||
| 90 | ORC2 | Orc2 | 14916 | -1.993 | -0.5531 | Yes | ||
| 91 | ORC6 | Orc6 | 14956 | -2.061 | -0.5420 | Yes | ||
| 92 | SKP2 | Skp2 | 15030 | -2.197 | -0.5322 | Yes | ||
| 93 | PKMYT1 | Pkmyt1 | 15075 | -2.298 | -0.5199 | Yes | ||
| 94 | CDKN2D | Cdkn2d | 15100 | -2.346 | -0.5060 | Yes | ||
| 95 | MCM3 | Mcm3 | 15264 | -2.840 | -0.4978 | Yes | ||
| 96 | MCM6 | Mcm6 | 15281 | -2.888 | -0.4797 | Yes | ||
| 97 | RBL1 | Rbl1 | 15283 | -2.888 | -0.4607 | Yes | ||
| 98 | ORC1 | Orc1 | 15289 | -2.913 | -0.4418 | Yes | ||
| 99 | CDKN2C | Cdkn2c | 15298 | -2.981 | -0.4226 | Yes | ||
| 100 | MAD2L1 | Mad2l1 | 15332 | -3.170 | -0.4038 | Yes | ||
| 101 | CDK1 | Cdk1 | 15355 | -3.325 | -0.3833 | Yes | ||
| 102 | CDC45 | Cdc45 | 15358 | -3.330 | -0.3614 | Yes | ||
| 103 | CHEK1 | Chek1 | 15360 | -3.334 | -0.3395 | Yes | ||
| 104 | CCNE1 | Ccne1 | 15413 | -3.704 | -0.3184 | Yes | ||
| 105 | MCM5 | Mcm5 | 15418 | -3.742 | -0.2939 | Yes | ||
| 106 | CCNE2 | Ccne2 | 15435 | -3.837 | -0.2696 | Yes | ||
| 107 | CDC25C | Cdc25c | 15466 | -3.990 | -0.2452 | Yes | ||
| 108 | CCNB1 | Ccnb1 | 15467 | -3.995 | -0.2188 | Yes | ||
| 109 | ESPL1 | Espl1 | 15479 | -4.062 | -0.1927 | Yes | ||
| 110 | CCNB2 | Ccnb2 | 15480 | -4.070 | -0.1658 | Yes | ||
| 111 | CDC20 | Cdc20 | 15495 | -4.143 | -0.1393 | Yes | ||
| 112 | BUB1 | Bub1 | 15497 | -4.158 | -0.1119 | Yes | ||
| 113 | CDC6 | Cdc6 | 15498 | -4.172 | -0.0844 | Yes | ||
| 114 | TTK | Ttk | 15511 | -4.266 | -0.0570 | Yes | ||
| 115 | CCNA2 | Ccna2 | 15519 | -4.320 | -0.0289 | Yes | ||
| 116 | PLK1 | Plk1 | 15522 | -4.422 | 0.0002 | Yes |