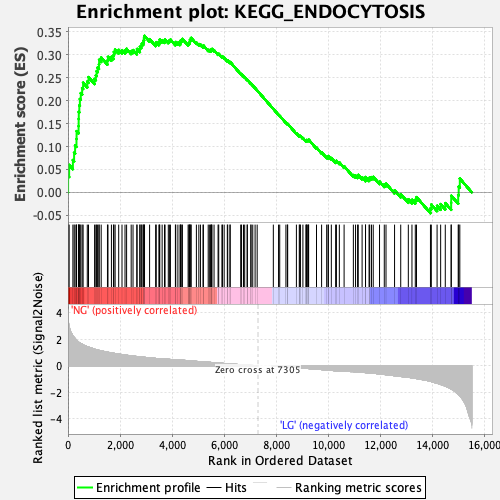
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_ENDOCYTOSIS |
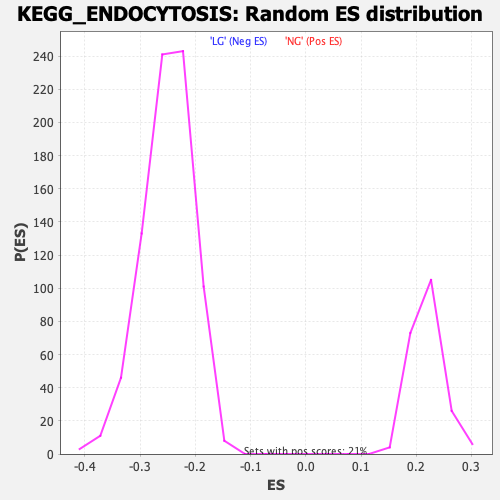
| Enrichment Score (ES) | 0.34135932 |
| Normalized Enrichment Score (NES) | 1.5615089 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2085835 |
| FWER p-Value | 0.6 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDR | Kdr | 4 | 4.063 | 0.0359 | Yes | ||
| 2 | CXCR4 | Cxcr4 | 44 | 2.984 | 0.0598 | Yes | ||
| 3 | CXCR2 | Cxcr2 | 188 | 2.261 | 0.0706 | Yes | ||
| 4 | VPS25 | Vps25 | 235 | 2.144 | 0.0867 | Yes | ||
| 5 | RAB11FIP4 | Rab11fip4 | 273 | 2.060 | 0.1026 | Yes | ||
| 6 | ASAP3 | Asap3 | 326 | 1.945 | 0.1165 | Yes | ||
| 7 | LDLR | Ldlr | 330 | 1.936 | 0.1335 | Yes | ||
| 8 | GRK4 | Grk4 | 406 | 1.785 | 0.1445 | Yes | ||
| 9 | ARAP2 | Arap2 | 410 | 1.779 | 0.1601 | Yes | ||
| 10 | CHMP4C | Chmp4c | 415 | 1.772 | 0.1756 | Yes | ||
| 11 | RET | Ret | 436 | 1.745 | 0.1898 | Yes | ||
| 12 | CBLB | Cblb | 457 | 1.718 | 0.2037 | Yes | ||
| 13 | PIKFYVE | Pikfyve | 492 | 1.680 | 0.2165 | Yes | ||
| 14 | SH3GL2 | Sh3gl2 | 544 | 1.631 | 0.2276 | Yes | ||
| 15 | HSPA8 | Hspa8 | 581 | 1.581 | 0.2393 | Yes | ||
| 16 | EHD3 | Ehd3 | 741 | 1.444 | 0.2418 | Yes | ||
| 17 | ARAP1 | Arap1 | 788 | 1.416 | 0.2514 | Yes | ||
| 18 | TRAF6 | Traf6 | 1022 | 1.254 | 0.2474 | Yes | ||
| 19 | GIT1 | Git1 | 1071 | 1.224 | 0.2551 | Yes | ||
| 20 | SH3GL3 | Sh3gl3 | 1099 | 1.215 | 0.2642 | Yes | ||
| 21 | STAM | Stam | 1138 | 1.193 | 0.2723 | Yes | ||
| 22 | CLTC | Cltc | 1190 | 1.168 | 0.2794 | Yes | ||
| 23 | PRKCZ | Prkcz | 1197 | 1.165 | 0.2893 | Yes | ||
| 24 | SMURF1 | Smurf1 | 1276 | 1.126 | 0.2943 | Yes | ||
| 25 | CHMP1B | Chmp1b | 1517 | 1.031 | 0.2878 | Yes | ||
| 26 | MDM2 | Mdm2 | 1537 | 1.023 | 0.2956 | Yes | ||
| 27 | CBL | Cbl | 1669 | 0.972 | 0.2958 | Yes | ||
| 28 | PIP4K2B | Pip4k2b | 1748 | 0.947 | 0.2991 | Yes | ||
| 29 | PSD2 | Psd2 | 1765 | 0.943 | 0.3064 | Yes | ||
| 30 | PIP5K1B | Pip5k1b | 1812 | 0.928 | 0.3117 | Yes | ||
| 31 | IGF1R | Igf1r | 1949 | 0.884 | 0.3107 | Yes | ||
| 32 | GIT2 | Git2 | 2076 | 0.853 | 0.3101 | Yes | ||
| 33 | RNF41 | Rnf41 | 2192 | 0.817 | 0.3098 | Yes | ||
| 34 | RAB31 | Rab31 | 2253 | 0.801 | 0.3131 | Yes | ||
| 35 | FLT1 | Flt1 | 2429 | 0.749 | 0.3083 | Yes | ||
| 36 | HGS | Hgs | 2500 | 0.734 | 0.3103 | Yes | ||
| 37 | PLD1 | Pld1 | 2647 | 0.700 | 0.3070 | Yes | ||
| 38 | PLD2 | Pld2 | 2654 | 0.698 | 0.3128 | Yes | ||
| 39 | IQSEC1 | Iqsec1 | 2763 | 0.677 | 0.3118 | Yes | ||
| 40 | RABEP1 | Rabep1 | 2766 | 0.676 | 0.3177 | Yes | ||
| 41 | STAM2 | Stam2 | 2816 | 0.667 | 0.3204 | Yes | ||
| 42 | AGAP2 | Agap2 | 2842 | 0.661 | 0.3247 | Yes | ||
| 43 | SH3GLB1 | Sh3glb1 | 2892 | 0.649 | 0.3273 | Yes | ||
| 44 | VPS37D | Vps37d | 2912 | 0.646 | 0.3318 | Yes | ||
| 45 | RAB11FIP1 | Rab11fip1 | 2925 | 0.644 | 0.3367 | Yes | ||
| 46 | VPS37A | Vps37a | 2942 | 0.641 | 0.3414 | Yes | ||
| 47 | PIP5K1A | Pip5k1a | 3137 | 0.604 | 0.3341 | No | ||
| 48 | ASAP2 | Asap2 | 3370 | 0.558 | 0.3240 | No | ||
| 49 | STAMBP | Stambp | 3392 | 0.553 | 0.3275 | No | ||
| 50 | KIT | Kit | 3495 | 0.536 | 0.3256 | No | ||
| 51 | RAB11FIP5 | Rab11fip5 | 3520 | 0.532 | 0.3288 | No | ||
| 52 | SMAP1 | Smap1 | 3527 | 0.531 | 0.3331 | No | ||
| 53 | ADRB2 | Adrb2 | 3619 | 0.518 | 0.3318 | No | ||
| 54 | ARFGAP3 | Arfgap3 | 3710 | 0.502 | 0.3304 | No | ||
| 55 | RAB11B | Rab11b | 3731 | 0.501 | 0.3336 | No | ||
| 56 | VPS4B | Vps4b | 3856 | 0.493 | 0.3299 | No | ||
| 57 | RAB22A | Rab22a | 3898 | 0.487 | 0.3315 | No | ||
| 58 | PARD6G | Pard6g | 3938 | 0.479 | 0.3332 | No | ||
| 59 | GRK3 | Grk3 | 4129 | 0.443 | 0.3248 | No | ||
| 60 | ERBB4 | Erbb4 | 4142 | 0.443 | 0.3280 | No | ||
| 61 | EEA1 | Eea1 | 4225 | 0.439 | 0.3265 | No | ||
| 62 | IQSEC3 | Iqsec3 | 4307 | 0.438 | 0.3251 | No | ||
| 63 | AP2A1 | Ap2a1 | 4325 | 0.433 | 0.3279 | No | ||
| 64 | VPS37B | Vps37b | 4332 | 0.432 | 0.3313 | No | ||
| 65 | PSD4 | Psd4 | 4375 | 0.425 | 0.3324 | No | ||
| 66 | TSG101 | Tsg101 | 4398 | 0.421 | 0.3347 | No | ||
| 67 | EGF | Egf | 4618 | 0.387 | 0.3239 | No | ||
| 68 | CHMP2B | Chmp2b | 4648 | 0.382 | 0.3254 | No | ||
| 69 | ASAP1 | Asap1 | 4673 | 0.379 | 0.3272 | No | ||
| 70 | HSPA2 | Hspa2 | 4685 | 0.377 | 0.3298 | No | ||
| 71 | DNAJC6 | Dnajc6 | 4691 | 0.377 | 0.3328 | No | ||
| 72 | RAB11A | Rab11a | 4725 | 0.372 | 0.3340 | No | ||
| 73 | RAB11FIP3 | Rab11fip3 | 4731 | 0.370 | 0.3370 | No | ||
| 74 | ACAP2 | Acap2 | 4936 | 0.334 | 0.3267 | No | ||
| 75 | SMURF2 | Smurf2 | 5030 | 0.321 | 0.3235 | No | ||
| 76 | USP8 | Usp8 | 5092 | 0.314 | 0.3223 | No | ||
| 77 | RAB5A | Rab5a | 5193 | 0.297 | 0.3184 | No | ||
| 78 | RAB11FIP2 | Rab11fip2 | 5204 | 0.295 | 0.3204 | No | ||
| 79 | CLTA | Clta | 5390 | 0.270 | 0.3107 | No | ||
| 80 | FGFR4 | Fgfr4 | 5434 | 0.264 | 0.3103 | No | ||
| 81 | NEDD4L | Nedd4l | 5482 | 0.257 | 0.3095 | No | ||
| 82 | RAB5B | Rab5b | 5510 | 0.253 | 0.3100 | No | ||
| 83 | PARD6A | Pard6a | 5512 | 0.252 | 0.3122 | No | ||
| 84 | SRC | Src | 5536 | 0.248 | 0.3129 | No | ||
| 85 | VPS37C | Vps37c | 5615 | 0.236 | 0.3099 | No | ||
| 86 | PIP5K1C | Pip5k1c | 5770 | 0.215 | 0.3018 | No | ||
| 87 | PSD | Psd | 5795 | 0.211 | 0.3021 | No | ||
| 88 | CHMP4B | Chmp4b | 5919 | 0.189 | 0.2958 | No | ||
| 89 | EPN2 | Epn2 | 5938 | 0.187 | 0.2963 | No | ||
| 90 | IQSEC2 | Iqsec2 | 6004 | 0.177 | 0.2936 | No | ||
| 91 | DAB2 | Dab2 | 6125 | 0.159 | 0.2872 | No | ||
| 92 | MVB12A | Mvb12a | 6141 | 0.158 | 0.2876 | No | ||
| 93 | AGAP1 | Agap1 | 6228 | 0.145 | 0.2833 | No | ||
| 94 | ARRB1 | Arrb1 | 6234 | 0.145 | 0.2843 | No | ||
| 95 | PARD3 | Pard3 | 6640 | 0.086 | 0.2587 | No | ||
| 96 | CBLC | Cblc | 6672 | 0.082 | 0.2574 | No | ||
| 97 | SMAP2 | Smap2 | 6747 | 0.076 | 0.2533 | No | ||
| 98 | MVB12B | Mvb12b | 6793 | 0.069 | 0.2510 | No | ||
| 99 | CHMP5 | Chmp5 | 6890 | 0.056 | 0.2452 | No | ||
| 100 | F2R | F2r | 6893 | 0.056 | 0.2456 | No | ||
| 101 | TFRC | Tfrc | 7030 | 0.038 | 0.2371 | No | ||
| 102 | AP2B1 | Ap2b1 | 7036 | 0.037 | 0.2371 | No | ||
| 103 | SH3GLB2 | Sh3glb2 | 7103 | 0.029 | 0.2330 | No | ||
| 104 | FGFR2 | Fgfr2 | 7193 | 0.016 | 0.2274 | No | ||
| 105 | ARFGAP1 | Arfgap1 | 7273 | 0.005 | 0.2223 | No | ||
| 106 | NEDD4 | Nedd4 | 7894 | -0.007 | 0.1820 | No | ||
| 107 | EPN1 | Epn1 | 8095 | -0.037 | 0.1693 | No | ||
| 108 | FGFR3 | Fgfr3 | 8140 | -0.045 | 0.1668 | No | ||
| 109 | GRK2 | Grk2 | 8378 | -0.079 | 0.1521 | No | ||
| 110 | CDC42 | Cdc42 | 8435 | -0.086 | 0.1492 | No | ||
| 111 | PARD6B | Pard6b | 8446 | -0.088 | 0.1493 | No | ||
| 112 | GRK5 | Grk5 | 8781 | -0.133 | 0.1288 | No | ||
| 113 | RBSN | Rbsn | 8892 | -0.150 | 0.1230 | No | ||
| 114 | VTA1 | Vta1 | 8913 | -0.153 | 0.1230 | No | ||
| 115 | PSD3 | Psd3 | 8936 | -0.157 | 0.1230 | No | ||
| 116 | SNF8 | Snf8 | 9031 | -0.173 | 0.1184 | No | ||
| 117 | ERBB3 | Erbb3 | 9147 | -0.188 | 0.1126 | No | ||
| 118 | DNM1L | Dnm1l | 9165 | -0.191 | 0.1132 | No | ||
| 119 | EPS15 | Eps15 | 9176 | -0.192 | 0.1142 | No | ||
| 120 | ITCH | Itch | 9216 | -0.198 | 0.1135 | No | ||
| 121 | RAB5C | Rab5c | 9243 | -0.203 | 0.1136 | No | ||
| 122 | ARFGAP2 | Arfgap2 | 9252 | -0.204 | 0.1149 | No | ||
| 123 | AP2S1 | Ap2s1 | 9554 | -0.252 | 0.0975 | No | ||
| 124 | ARRB2 | Arrb2 | 9750 | -0.281 | 0.0873 | No | ||
| 125 | DNM2 | Dnm2 | 9945 | -0.317 | 0.0775 | No | ||
| 126 | ACAP3 | Acap3 | 10001 | -0.325 | 0.0768 | No | ||
| 127 | VPS36 | Vps36 | 10009 | -0.326 | 0.0793 | No | ||
| 128 | SH3GL1 | Sh3gl1 | 10124 | -0.344 | 0.0749 | No | ||
| 129 | LDLRAP1 | Ldlrap1 | 10290 | -0.370 | 0.0674 | No | ||
| 130 | VPS45 | Vps45 | 10320 | -0.374 | 0.0689 | No | ||
| 131 | ARF6 | Arf6 | 10432 | -0.395 | 0.0652 | No | ||
| 132 | CHMP2A | Chmp2a | 10615 | -0.408 | 0.0570 | No | ||
| 133 | MET | Met | 10971 | -0.436 | 0.0377 | No | ||
| 134 | RAB4A | Rab4a | 11055 | -0.453 | 0.0363 | No | ||
| 135 | AP2M1 | Ap2m1 | 11134 | -0.466 | 0.0354 | No | ||
| 136 | VPS28 | Vps28 | 11158 | -0.472 | 0.0381 | No | ||
| 137 | AP2A2 | Ap2a2 | 11307 | -0.490 | 0.0328 | No | ||
| 138 | EHD2 | Ehd2 | 11438 | -0.509 | 0.0289 | No | ||
| 139 | ADRB1 | Adrb1 | 11439 | -0.509 | 0.0334 | No | ||
| 140 | RUFY1 | Rufy1 | 11575 | -0.531 | 0.0293 | No | ||
| 141 | EHD1 | Ehd1 | 11599 | -0.535 | 0.0326 | No | ||
| 142 | WWP1 | Wwp1 | 11667 | -0.549 | 0.0331 | No | ||
| 143 | CSF1R | Csf1r | 11735 | -0.562 | 0.0338 | No | ||
| 144 | SH3KBP1 | Sh3kbp1 | 11977 | -0.607 | 0.0235 | No | ||
| 145 | DNM1 | Dnm1 | 12160 | -0.651 | 0.0174 | No | ||
| 146 | HRAS | Hras | 12226 | -0.666 | 0.0191 | No | ||
| 147 | PRKCI | Prkci | 12555 | -0.742 | 0.0043 | No | ||
| 148 | CLTB | Cltb | 12792 | -0.801 | -0.0039 | No | ||
| 149 | IL2RA | Il2ra | 13084 | -0.863 | -0.0152 | No | ||
| 150 | CHMP6 | Chmp6 | 13224 | -0.904 | -0.0162 | No | ||
| 151 | PDGFRA | Pdgfra | 13352 | -0.947 | -0.0160 | No | ||
| 152 | GRK1 | Grk1 | 13398 | -0.960 | -0.0104 | No | ||
| 153 | GRK6 | Grk6 | 13935 | -1.167 | -0.0350 | No | ||
| 154 | EGFR | Egfr | 13968 | -1.184 | -0.0265 | No | ||
| 155 | ACAP1 | Acap1 | 14190 | -1.314 | -0.0292 | No | ||
| 156 | IL2RB | Il2rb | 14327 | -1.401 | -0.0256 | No | ||
| 157 | HSPA1L | Hspa1l | 14505 | -1.535 | -0.0235 | No | ||
| 158 | EHD4 | Ehd4 | 14731 | -1.740 | -0.0227 | No | ||
| 159 | DNM3 | Dnm3 | 14734 | -1.740 | -0.0074 | No | ||
| 160 | EPN3 | Epn3 | 15002 | -2.144 | -0.0057 | No | ||
| 161 | ARAP3 | Arap3 | 15021 | -2.181 | 0.0125 | No | ||
| 162 | NTRK1 | Ntrk1 | 15064 | -2.274 | 0.0300 | No |