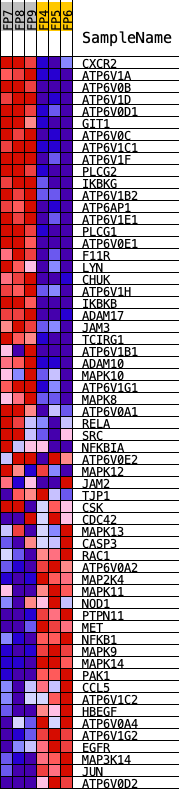
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_EPITHELIAL_CELL_SIGNALING_IN_HELICOBACTER_PYLORI_INFECTION |
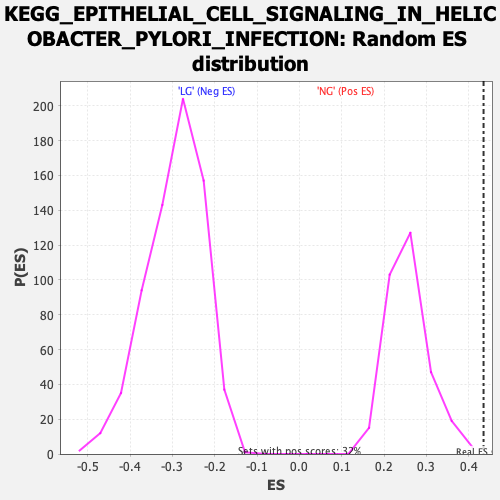
| Enrichment Score (ES) | 0.43525934 |
| Normalized Enrichment Score (NES) | 1.6933173 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08404922 |
| FWER p-Value | 0.263 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CXCR2 | Cxcr2 | 188 | 2.261 | 0.0346 | Yes | ||
| 2 | ATP6V1A | Atp6v1a | 441 | 1.737 | 0.0543 | Yes | ||
| 3 | ATP6V0B | Atp6v0b | 461 | 1.713 | 0.0885 | Yes | ||
| 4 | ATP6V1D | Atp6v1d | 697 | 1.480 | 0.1040 | Yes | ||
| 5 | ATP6V0D1 | Atp6v0d1 | 1008 | 1.261 | 0.1100 | Yes | ||
| 6 | GIT1 | Git1 | 1071 | 1.224 | 0.1313 | Yes | ||
| 7 | ATP6V0C | Atp6v0c | 1123 | 1.204 | 0.1530 | Yes | ||
| 8 | ATP6V1C1 | Atp6v1c1 | 1309 | 1.114 | 0.1641 | Yes | ||
| 9 | ATP6V1F | Atp6v1f | 1318 | 1.112 | 0.1866 | Yes | ||
| 10 | PLCG2 | Plcg2 | 1349 | 1.099 | 0.2074 | Yes | ||
| 11 | IKBKG | Ikbkg | 1409 | 1.077 | 0.2259 | Yes | ||
| 12 | ATP6V1B2 | Atp6v1b2 | 1502 | 1.037 | 0.2414 | Yes | ||
| 13 | ATP6AP1 | Atp6ap1 | 1516 | 1.032 | 0.2619 | Yes | ||
| 14 | ATP6V1E1 | Atp6v1e1 | 1608 | 0.998 | 0.2767 | Yes | ||
| 15 | PLCG1 | Plcg1 | 1638 | 0.987 | 0.2952 | Yes | ||
| 16 | ATP6V0E1 | Atp6v0e | 1654 | 0.978 | 0.3145 | Yes | ||
| 17 | F11R | F11r | 1790 | 0.936 | 0.3252 | Yes | ||
| 18 | LYN | Lyn | 2070 | 0.854 | 0.3248 | Yes | ||
| 19 | CHUK | Chuk | 2135 | 0.833 | 0.3379 | Yes | ||
| 20 | ATP6V1H | Atp6v1h | 2325 | 0.779 | 0.3418 | Yes | ||
| 21 | IKBKB | Ikbkb | 2340 | 0.774 | 0.3569 | Yes | ||
| 22 | ADAM17 | Adam17 | 2356 | 0.770 | 0.3719 | Yes | ||
| 23 | JAM3 | Jam3 | 2398 | 0.757 | 0.3849 | Yes | ||
| 24 | TCIRG1 | Tcirg1 | 2492 | 0.736 | 0.3941 | Yes | ||
| 25 | ATP6V1B1 | Atp6v1b1 | 2531 | 0.728 | 0.4067 | Yes | ||
| 26 | ADAM10 | Adam10 | 2719 | 0.686 | 0.4088 | Yes | ||
| 27 | MAPK10 | Mapk10 | 2739 | 0.681 | 0.4217 | Yes | ||
| 28 | ATP6V1G1 | Atp6v1g1 | 2748 | 0.679 | 0.4353 | Yes | ||
| 29 | MAPK8 | Mapk8 | 3986 | 0.468 | 0.3650 | No | ||
| 30 | ATP6V0A1 | Atp6v0a1 | 4347 | 0.429 | 0.3506 | No | ||
| 31 | RELA | Rela | 4444 | 0.415 | 0.3530 | No | ||
| 32 | SRC | Src | 5536 | 0.248 | 0.2876 | No | ||
| 33 | NFKBIA | Nfkbia | 5932 | 0.188 | 0.2659 | No | ||
| 34 | ATP6V0E2 | Atp6v0e2 | 6269 | 0.140 | 0.2471 | No | ||
| 35 | MAPK12 | Mapk12 | 6820 | 0.065 | 0.2128 | No | ||
| 36 | JAM2 | Jam2 | 6918 | 0.052 | 0.2076 | No | ||
| 37 | TJP1 | Tjp1 | 7982 | -0.020 | 0.1393 | No | ||
| 38 | CSK | Csk | 8130 | -0.043 | 0.1307 | No | ||
| 39 | CDC42 | Cdc42 | 8435 | -0.086 | 0.1128 | No | ||
| 40 | MAPK13 | Mapk13 | 9159 | -0.190 | 0.0700 | No | ||
| 41 | CASP3 | Casp3 | 9177 | -0.192 | 0.0729 | No | ||
| 42 | RAC1 | Rac1 | 9562 | -0.252 | 0.0533 | No | ||
| 43 | ATP6V0A2 | Atp6v0a2 | 9618 | -0.261 | 0.0551 | No | ||
| 44 | MAP2K4 | Map2k4 | 9737 | -0.280 | 0.0533 | No | ||
| 45 | MAPK11 | Mapk11 | 10363 | -0.382 | 0.0208 | No | ||
| 46 | NOD1 | Nod1 | 10384 | -0.386 | 0.0275 | No | ||
| 47 | PTPN11 | Ptpn11 | 10836 | -0.428 | 0.0072 | No | ||
| 48 | MET | Met | 10971 | -0.436 | 0.0075 | No | ||
| 49 | NFKB1 | Nfkb1 | 11113 | -0.463 | 0.0080 | No | ||
| 50 | MAPK9 | Mapk9 | 11878 | -0.588 | -0.0292 | No | ||
| 51 | MAPK14 | Mapk14 | 12063 | -0.629 | -0.0281 | No | ||
| 52 | PAK1 | Pak1 | 12137 | -0.646 | -0.0194 | No | ||
| 53 | CCL5 | Ccl5 | 13106 | -0.868 | -0.0641 | No | ||
| 54 | ATP6V1C2 | Atp6v1c2 | 13292 | -0.924 | -0.0569 | No | ||
| 55 | HBEGF | Hbegf | 13599 | -1.030 | -0.0554 | No | ||
| 56 | ATP6V0A4 | Atp6v0a4 | 13667 | -1.055 | -0.0379 | No | ||
| 57 | ATP6V1G2 | Atp6v1g2 | 13874 | -1.136 | -0.0277 | No | ||
| 58 | EGFR | Egfr | 13968 | -1.184 | -0.0092 | No | ||
| 59 | MAP3K14 | Map3k14 | 14142 | -1.290 | 0.0063 | No | ||
| 60 | JUN | Jun | 14761 | -1.775 | 0.0031 | No | ||
| 61 | ATP6V0D2 | Atp6v0d2 | 15045 | -2.233 | 0.0310 | No |