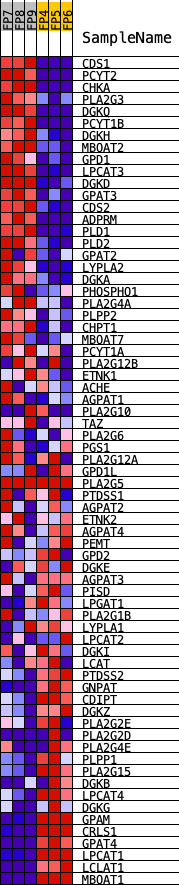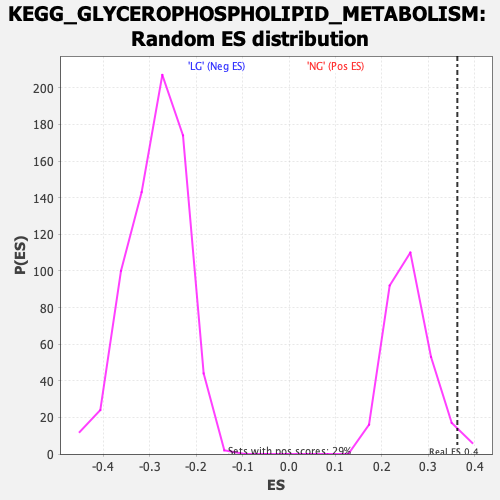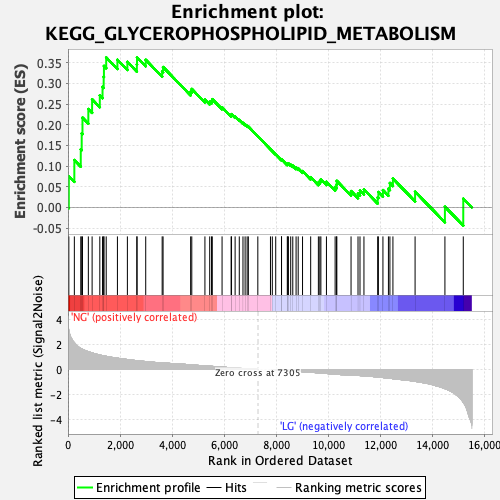
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_GLYCEROPHOSPHOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.36259297 |
| Normalized Enrichment Score (NES) | 1.4063255 |
| Nominal p-value | 0.020408163 |
| FDR q-value | 0.26614383 |
| FWER p-Value | 0.929 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDS1 | Cds1 | 32 | 3.129 | 0.0755 | Yes | ||
| 2 | PCYT2 | Pcyt2 | 244 | 2.125 | 0.1146 | Yes | ||
| 3 | CHKA | Chka | 491 | 1.680 | 0.1404 | Yes | ||
| 4 | PLA2G3 | Pla2g3 | 527 | 1.644 | 0.1789 | Yes | ||
| 5 | DGKQ | Dgkq | 557 | 1.616 | 0.2171 | Yes | ||
| 6 | PCYT1B | Pcyt1b | 782 | 1.422 | 0.2378 | Yes | ||
| 7 | DGKH | Dgkh | 927 | 1.322 | 0.2613 | Yes | ||
| 8 | MBOAT2 | Mboat2 | 1223 | 1.150 | 0.2707 | Yes | ||
| 9 | GPD1 | Gpd1 | 1328 | 1.106 | 0.2915 | Yes | ||
| 10 | LPCAT3 | Lpcat3 | 1372 | 1.088 | 0.3157 | Yes | ||
| 11 | DGKD | Dgkd | 1382 | 1.086 | 0.3420 | Yes | ||
| 12 | GPAT3 | Gpat3 | 1469 | 1.047 | 0.3624 | Yes | ||
| 13 | CDS2 | Cds2 | 1900 | 0.900 | 0.3569 | Yes | ||
| 14 | ADPRM | Adprm | 2283 | 0.790 | 0.3518 | Yes | ||
| 15 | PLD1 | Pld1 | 2647 | 0.700 | 0.3457 | Yes | ||
| 16 | PLD2 | Pld2 | 2654 | 0.698 | 0.3626 | Yes | ||
| 17 | GPAT2 | Gpat2 | 2988 | 0.632 | 0.3567 | No | ||
| 18 | LYPLA2 | Lypla2 | 3620 | 0.518 | 0.3288 | No | ||
| 19 | DGKA | Dgka | 3655 | 0.513 | 0.3393 | No | ||
| 20 | PHOSPHO1 | Phospho1 | 4715 | 0.373 | 0.2800 | No | ||
| 21 | PLA2G4A | Pla2g4a | 4759 | 0.366 | 0.2863 | No | ||
| 22 | PLPP2 | Plpp2 | 5264 | 0.287 | 0.2608 | No | ||
| 23 | CHPT1 | Chpt1 | 5448 | 0.262 | 0.2555 | No | ||
| 24 | MBOAT7 | Mboat7 | 5518 | 0.252 | 0.2573 | No | ||
| 25 | PCYT1A | Pcyt1a | 5547 | 0.246 | 0.2616 | No | ||
| 26 | PLA2G12B | Pla2g12b | 5923 | 0.189 | 0.2420 | No | ||
| 27 | ETNK1 | Etnk1 | 6271 | 0.139 | 0.2230 | No | ||
| 28 | ACHE | Ache | 6287 | 0.138 | 0.2254 | No | ||
| 29 | AGPAT1 | Agpat1 | 6425 | 0.117 | 0.2195 | No | ||
| 30 | PLA2G10 | Pla2g10 | 6585 | 0.093 | 0.2115 | No | ||
| 31 | TAZ | Taz | 6727 | 0.078 | 0.2043 | No | ||
| 32 | PLA2G6 | Pla2g6 | 6809 | 0.067 | 0.2007 | No | ||
| 33 | PGS1 | Pgs1 | 6887 | 0.057 | 0.1971 | No | ||
| 34 | PLA2G12A | Pla2g12a | 6934 | 0.051 | 0.1954 | No | ||
| 35 | GPD1L | Gpd1l | 7298 | 0.001 | 0.1720 | No | ||
| 36 | PLA2G5 | Pla2g5 | 7784 | 0.000 | 0.1406 | No | ||
| 37 | PTDSS1 | Ptdss1 | 7855 | -0.001 | 0.1361 | No | ||
| 38 | AGPAT2 | Agpat2 | 7986 | -0.020 | 0.1282 | No | ||
| 39 | ETNK2 | Etnk2 | 8208 | -0.055 | 0.1153 | No | ||
| 40 | AGPAT4 | Agpat4 | 8210 | -0.055 | 0.1166 | No | ||
| 41 | PEMT | Pemt | 8431 | -0.085 | 0.1044 | No | ||
| 42 | GPD2 | Gpd2 | 8448 | -0.088 | 0.1056 | No | ||
| 43 | DGKE | Dgke | 8474 | -0.091 | 0.1062 | No | ||
| 44 | AGPAT3 | Agpat3 | 8562 | -0.103 | 0.1031 | No | ||
| 45 | PISD | Pisd | 8642 | -0.114 | 0.1009 | No | ||
| 46 | LPGAT1 | Lpgat1 | 8769 | -0.131 | 0.0960 | No | ||
| 47 | PLA2G1B | Pla2g1b | 8850 | -0.143 | 0.0943 | No | ||
| 48 | LYPLA1 | Lypla1 | 9017 | -0.172 | 0.0879 | No | ||
| 49 | LPCAT2 | Lpcat2 | 9330 | -0.217 | 0.0730 | No | ||
| 50 | DGKI | Dgki | 9626 | -0.262 | 0.0605 | No | ||
| 51 | LCAT | Lcat | 9681 | -0.271 | 0.0637 | No | ||
| 52 | PTDSS2 | Ptdss2 | 9722 | -0.278 | 0.0680 | No | ||
| 53 | GNPAT | Gnpat | 9936 | -0.315 | 0.0620 | No | ||
| 54 | CDIPT | Cdipt | 10270 | -0.366 | 0.0496 | No | ||
| 55 | DGKZ | Dgkz | 10327 | -0.376 | 0.0553 | No | ||
| 56 | PLA2G2E | Pla2g2e | 10333 | -0.376 | 0.0643 | No | ||
| 57 | PLA2G2D | Pla2g2d | 10882 | -0.432 | 0.0395 | No | ||
| 58 | PLA2G4E | Pla2g4e | 11150 | -0.470 | 0.0339 | No | ||
| 59 | PLPP1 | Plpp1 | 11226 | -0.482 | 0.0410 | No | ||
| 60 | PLA2G15 | Pla2g15 | 11378 | -0.496 | 0.0435 | No | ||
| 61 | DGKB | Dgkb | 11906 | -0.592 | 0.0241 | No | ||
| 62 | LPCAT4 | Lpcat4 | 11933 | -0.598 | 0.0373 | No | ||
| 63 | DGKG | Dgkg | 12108 | -0.639 | 0.0419 | No | ||
| 64 | GPAM | Gpam | 12319 | -0.686 | 0.0453 | No | ||
| 65 | CRLS1 | Crls1 | 12376 | -0.703 | 0.0591 | No | ||
| 66 | GPAT4 | Gpat4 | 12492 | -0.727 | 0.0697 | No | ||
| 67 | LPCAT1 | Lpcat1 | 13344 | -0.945 | 0.0381 | No | ||
| 68 | LCLAT1 | Lclat1 | 14491 | -1.524 | 0.0018 | No | ||
| 69 | MBOAT1 | Mboat1 | 15197 | -2.623 | 0.0212 | No |