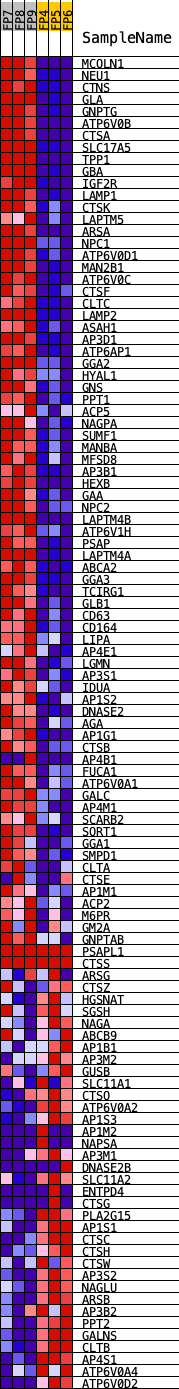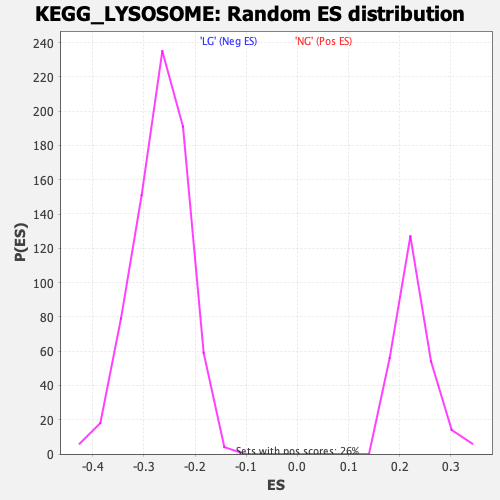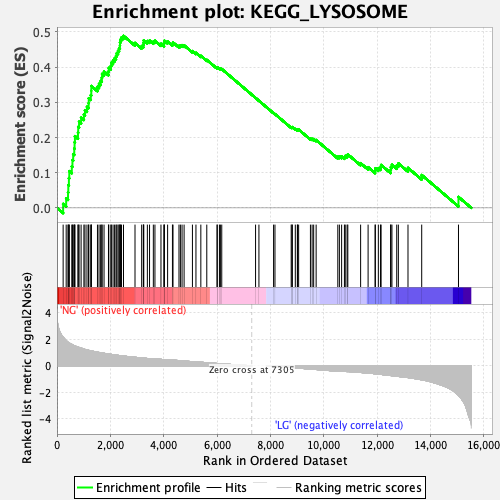
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_LYSOSOME |
| Enrichment Score (ES) | 0.48889416 |
| Normalized Enrichment Score (NES) | 2.1438372 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MCOLN1 | Mcoln1 | 230 | 2.159 | 0.0113 | Yes | ||
| 2 | NEU1 | Neu1 | 344 | 1.904 | 0.0271 | Yes | ||
| 3 | CTNS | Ctns | 414 | 1.772 | 0.0441 | Yes | ||
| 4 | GLA | Gla | 424 | 1.761 | 0.0649 | Yes | ||
| 5 | GNPTG | Gnptg | 446 | 1.728 | 0.0845 | Yes | ||
| 6 | ATP6V0B | Atp6v0b | 461 | 1.713 | 0.1044 | Yes | ||
| 7 | CTSA | Ctsa | 555 | 1.617 | 0.1180 | Yes | ||
| 8 | SLC17A5 | Slc17a5 | 579 | 1.586 | 0.1358 | Yes | ||
| 9 | TPP1 | Tpp1 | 607 | 1.556 | 0.1529 | Yes | ||
| 10 | GBA | Gba | 647 | 1.520 | 0.1688 | Yes | ||
| 11 | IGF2R | Igf2r | 659 | 1.509 | 0.1864 | Yes | ||
| 12 | LAMP1 | Lamp1 | 672 | 1.494 | 0.2038 | Yes | ||
| 13 | CTSK | Ctsk | 787 | 1.417 | 0.2136 | Yes | ||
| 14 | LAPTM5 | Laptm5 | 795 | 1.412 | 0.2303 | Yes | ||
| 15 | ARSA | Arsa | 827 | 1.387 | 0.2451 | Yes | ||
| 16 | NPC1 | Npc1 | 903 | 1.334 | 0.2565 | Yes | ||
| 17 | ATP6V0D1 | Atp6v0d1 | 1008 | 1.261 | 0.2650 | Yes | ||
| 18 | MAN2B1 | Man2b1 | 1048 | 1.239 | 0.2775 | Yes | ||
| 19 | ATP6V0C | Atp6v0c | 1123 | 1.204 | 0.2874 | Yes | ||
| 20 | CTSF | Ctsf | 1185 | 1.171 | 0.2976 | Yes | ||
| 21 | CLTC | Cltc | 1190 | 1.168 | 0.3115 | Yes | ||
| 22 | LAMP2 | Lamp2 | 1266 | 1.130 | 0.3204 | Yes | ||
| 23 | ASAH1 | Asah1 | 1285 | 1.123 | 0.3329 | Yes | ||
| 24 | AP3D1 | Ap3d1 | 1287 | 1.123 | 0.3464 | Yes | ||
| 25 | ATP6AP1 | Atp6ap1 | 1516 | 1.032 | 0.3442 | Yes | ||
| 26 | GGA2 | Gga2 | 1575 | 1.010 | 0.3527 | Yes | ||
| 27 | HYAL1 | Hyal1 | 1635 | 0.988 | 0.3608 | Yes | ||
| 28 | GNS | Gns | 1679 | 0.968 | 0.3698 | Yes | ||
| 29 | PPT1 | Ppt1 | 1690 | 0.965 | 0.3809 | Yes | ||
| 30 | ACP5 | Acp5 | 1763 | 0.943 | 0.3876 | Yes | ||
| 31 | NAGPA | Nagpa | 1929 | 0.891 | 0.3878 | Yes | ||
| 32 | SUMF1 | Sumf1 | 1941 | 0.887 | 0.3978 | Yes | ||
| 33 | MANBA | Manba | 2017 | 0.869 | 0.4035 | Yes | ||
| 34 | MFSD8 | Mfsd8 | 2037 | 0.862 | 0.4127 | Yes | ||
| 35 | AP3B1 | Ap3b1 | 2100 | 0.843 | 0.4189 | Yes | ||
| 36 | HEXB | Hexb | 2158 | 0.828 | 0.4253 | Yes | ||
| 37 | GAA | Gaa | 2210 | 0.813 | 0.4319 | Yes | ||
| 38 | NPC2 | Npc2 | 2240 | 0.804 | 0.4397 | Yes | ||
| 39 | LAPTM4B | Laptm4b | 2287 | 0.789 | 0.4463 | Yes | ||
| 40 | ATP6V1H | Atp6v1h | 2325 | 0.779 | 0.4534 | Yes | ||
| 41 | PSAP | Psap | 2357 | 0.769 | 0.4607 | Yes | ||
| 42 | LAPTM4A | Laptm4a | 2361 | 0.769 | 0.4699 | Yes | ||
| 43 | ABCA2 | Abca2 | 2376 | 0.763 | 0.4782 | Yes | ||
| 44 | GGA3 | Gga3 | 2418 | 0.752 | 0.4847 | Yes | ||
| 45 | TCIRG1 | Tcirg1 | 2492 | 0.736 | 0.4889 | Yes | ||
| 46 | GLB1 | Glb1 | 2923 | 0.644 | 0.4688 | No | ||
| 47 | CD63 | Cd63 | 3173 | 0.595 | 0.4599 | No | ||
| 48 | CD164 | Cd164 | 3235 | 0.584 | 0.4630 | No | ||
| 49 | LIPA | Lipa | 3243 | 0.582 | 0.4696 | No | ||
| 50 | AP4E1 | Ap4e1 | 3251 | 0.581 | 0.4762 | No | ||
| 51 | LGMN | Lgmn | 3386 | 0.554 | 0.4743 | No | ||
| 52 | AP3S1 | Ap3s1 | 3468 | 0.540 | 0.4756 | No | ||
| 53 | IDUA | Idua | 3610 | 0.519 | 0.4727 | No | ||
| 54 | AP1S2 | Ap1s2 | 3670 | 0.509 | 0.4751 | No | ||
| 55 | DNASE2 | Dnase2a | 3896 | 0.487 | 0.4664 | No | ||
| 56 | AGA | Aga | 4011 | 0.463 | 0.4646 | No | ||
| 57 | AP1G1 | Ap1g1 | 4013 | 0.463 | 0.4702 | No | ||
| 58 | CTSB | Ctsb | 4024 | 0.461 | 0.4751 | No | ||
| 59 | AP4B1 | Ap4b1 | 4145 | 0.443 | 0.4727 | No | ||
| 60 | FUCA1 | Fuca1 | 4326 | 0.433 | 0.4663 | No | ||
| 61 | ATP6V0A1 | Atp6v0a1 | 4347 | 0.429 | 0.4702 | No | ||
| 62 | GALC | Galc | 4571 | 0.394 | 0.4606 | No | ||
| 63 | AP4M1 | Ap4m1 | 4624 | 0.386 | 0.4619 | No | ||
| 64 | SCARB2 | Scarb2 | 4686 | 0.377 | 0.4625 | No | ||
| 65 | SORT1 | Sort1 | 4763 | 0.365 | 0.4620 | No | ||
| 66 | GGA1 | Gga1 | 5075 | 0.316 | 0.4456 | No | ||
| 67 | SMPD1 | Smpd1 | 5207 | 0.294 | 0.4407 | No | ||
| 68 | CLTA | Clta | 5390 | 0.270 | 0.4322 | No | ||
| 69 | CTSE | Ctse | 5613 | 0.236 | 0.4206 | No | ||
| 70 | AP1M1 | Ap1m1 | 5998 | 0.178 | 0.3979 | No | ||
| 71 | ACP2 | Acp2 | 6005 | 0.177 | 0.3997 | No | ||
| 72 | M6PR | M6pr | 6087 | 0.166 | 0.3964 | No | ||
| 73 | GM2A | Gm2a | 6123 | 0.159 | 0.3961 | No | ||
| 74 | GNPTAB | Gnptab | 6174 | 0.153 | 0.3947 | No | ||
| 75 | PSAPL1 | Psapl1 | 7441 | 0.000 | 0.3126 | No | ||
| 76 | CTSS | Ctss | 7567 | 0.000 | 0.3045 | No | ||
| 77 | ARSG | Arsg | 8116 | -0.040 | 0.2694 | No | ||
| 78 | CTSZ | Ctsz | 8168 | -0.050 | 0.2667 | No | ||
| 79 | HGSNAT | Hgsnat | 8776 | -0.132 | 0.2289 | No | ||
| 80 | SGSH | Sgsh | 8807 | -0.137 | 0.2287 | No | ||
| 81 | NAGA | Naga | 8818 | -0.138 | 0.2297 | No | ||
| 82 | ABCB9 | Abcb9 | 8930 | -0.156 | 0.2244 | No | ||
| 83 | AP1B1 | Ap1b1 | 9007 | -0.169 | 0.2215 | No | ||
| 84 | AP3M2 | Ap3m2 | 9012 | -0.171 | 0.2233 | No | ||
| 85 | GUSB | Gusb | 9055 | -0.176 | 0.2227 | No | ||
| 86 | SLC11A1 | Slc11a1 | 9497 | -0.242 | 0.1971 | No | ||
| 87 | CTSO | Ctso | 9560 | -0.252 | 0.1961 | No | ||
| 88 | ATP6V0A2 | Atp6v0a2 | 9618 | -0.261 | 0.1956 | No | ||
| 89 | AP1S3 | Ap1s3 | 9711 | -0.276 | 0.1930 | No | ||
| 90 | AP1M2 | Ap1m2 | 10523 | -0.407 | 0.1453 | No | ||
| 91 | NAPSA | Napsa | 10582 | -0.407 | 0.1465 | No | ||
| 92 | AP3M1 | Ap3m1 | 10666 | -0.417 | 0.1461 | No | ||
| 93 | DNASE2B | Dnase2b | 10771 | -0.421 | 0.1445 | No | ||
| 94 | SLC11A2 | Slc11a2 | 10801 | -0.423 | 0.1478 | No | ||
| 95 | ENTPD4 | Entpd4b | 10870 | -0.432 | 0.1486 | No | ||
| 96 | CTSG | Ctsg | 10898 | -0.432 | 0.1521 | No | ||
| 97 | PLA2G15 | Pla2g15 | 11378 | -0.496 | 0.1270 | No | ||
| 98 | AP1S1 | Ap1s1 | 11658 | -0.547 | 0.1156 | No | ||
| 99 | CTSC | Ctsc | 11920 | -0.595 | 0.1059 | No | ||
| 100 | CTSH | Ctsh | 11930 | -0.597 | 0.1125 | No | ||
| 101 | CTSW | Ctsw | 12042 | -0.625 | 0.1129 | No | ||
| 102 | AP3S2 | Ap3s2 | 12122 | -0.642 | 0.1156 | No | ||
| 103 | NAGLU | Naglu | 12145 | -0.646 | 0.1220 | No | ||
| 104 | ARSB | Arsb | 12501 | -0.730 | 0.1078 | No | ||
| 105 | AP3B2 | Ap3b2 | 12511 | -0.733 | 0.1162 | No | ||
| 106 | PPT2 | Ppt2 | 12551 | -0.741 | 0.1226 | No | ||
| 107 | GALNS | Galns | 12731 | -0.789 | 0.1206 | No | ||
| 108 | CLTB | Cltb | 12792 | -0.801 | 0.1264 | No | ||
| 109 | AP4S1 | Ap4s1 | 13153 | -0.886 | 0.1138 | No | ||
| 110 | ATP6V0A4 | Atp6v0a4 | 13667 | -1.055 | 0.0934 | No | ||
| 111 | ATP6V0D2 | Atp6v0d2 | 15045 | -2.233 | 0.0311 | No |