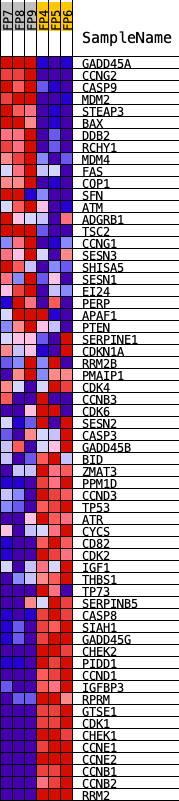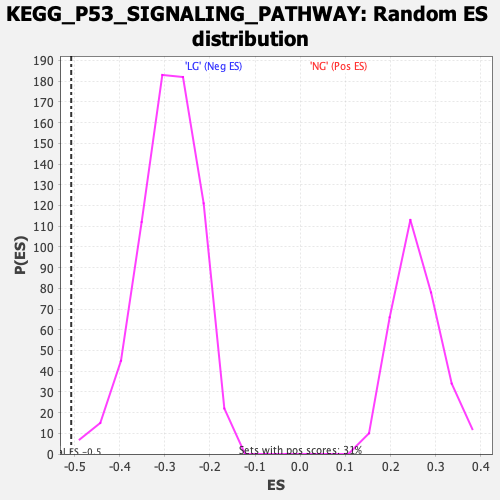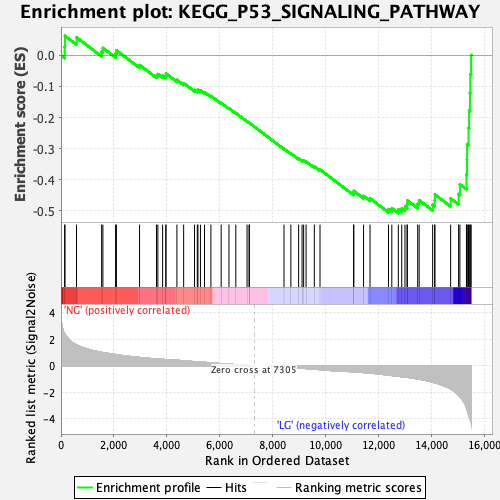
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_P53_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.50739247 |
| Normalized Enrichment Score (NES) | -1.7370912 |
| Nominal p-value | 0.0014556041 |
| FDR q-value | 0.016836757 |
| FWER p-Value | 0.225 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GADD45A | Gadd45a | 138 | 2.445 | 0.0275 | No | ||
| 2 | CCNG2 | Ccng2 | 150 | 2.406 | 0.0626 | No | ||
| 3 | CASP9 | Casp9 | 587 | 1.576 | 0.0578 | No | ||
| 4 | MDM2 | Mdm2 | 1537 | 1.023 | 0.0117 | No | ||
| 5 | STEAP3 | Steap3 | 1584 | 1.006 | 0.0237 | No | ||
| 6 | BAX | Bax | 2064 | 0.856 | 0.0055 | No | ||
| 7 | DDB2 | Ddb2 | 2095 | 0.844 | 0.0161 | No | ||
| 8 | RCHY1 | Rchy1 | 2971 | 0.635 | -0.0310 | No | ||
| 9 | MDM4 | Mdm4 | 3611 | 0.519 | -0.0646 | No | ||
| 10 | FAS | Fas | 3663 | 0.510 | -0.0603 | No | ||
| 11 | COP1 | Cop1 | 3843 | 0.494 | -0.0646 | No | ||
| 12 | SFN | Sfn | 3965 | 0.473 | -0.0653 | No | ||
| 13 | ATM | Atm | 3970 | 0.472 | -0.0586 | No | ||
| 14 | ADGRB1 | Adgrb1 | 4384 | 0.423 | -0.0790 | No | ||
| 15 | TSC2 | Tsc2 | 4639 | 0.384 | -0.0897 | No | ||
| 16 | CCNG1 | Ccng1 | 5050 | 0.319 | -0.1115 | No | ||
| 17 | SESN3 | Sesn3 | 5155 | 0.303 | -0.1137 | No | ||
| 18 | SHISA5 | Shisa5 | 5179 | 0.299 | -0.1107 | No | ||
| 19 | SESN1 | Sesn1 | 5274 | 0.286 | -0.1125 | No | ||
| 20 | EI24 | Ei24 | 5428 | 0.265 | -0.1185 | No | ||
| 21 | PERP | Perp | 5670 | 0.231 | -0.1306 | No | ||
| 22 | APAF1 | Apaf1 | 6058 | 0.170 | -0.1531 | No | ||
| 23 | PTEN | Pten | 6353 | 0.129 | -0.1702 | No | ||
| 24 | SERPINE1 | Serpine1 | 6612 | 0.090 | -0.1856 | No | ||
| 25 | CDKN1A | Cdkn1a | 7037 | 0.037 | -0.2124 | No | ||
| 26 | RRM2B | Rrm2b | 7113 | 0.027 | -0.2169 | No | ||
| 27 | PMAIP1 | Pmaip1 | 7125 | 0.026 | -0.2172 | No | ||
| 28 | CDK4 | Cdk4 | 8434 | -0.086 | -0.3005 | No | ||
| 29 | CCNB3 | Ccnb3 | 8694 | -0.120 | -0.3155 | No | ||
| 30 | CDK6 | Cdk6 | 8988 | -0.167 | -0.3319 | No | ||
| 31 | SESN2 | Sesn2 | 9118 | -0.184 | -0.3375 | No | ||
| 32 | CASP3 | Casp3 | 9177 | -0.192 | -0.3384 | No | ||
| 33 | GADD45B | Gadd45b | 9269 | -0.208 | -0.3412 | No | ||
| 34 | BID | Bid | 9581 | -0.255 | -0.3575 | No | ||
| 35 | ZMAT3 | Zmat3 | 9796 | -0.289 | -0.3671 | No | ||
| 36 | PPM1D | Ppm1d | 11066 | -0.454 | -0.4424 | No | ||
| 37 | CCND3 | Ccnd3 | 11075 | -0.456 | -0.4361 | No | ||
| 38 | TP53 | Trp53 | 11444 | -0.509 | -0.4523 | No | ||
| 39 | ATR | Atr | 11688 | -0.552 | -0.4598 | No | ||
| 40 | CYCS | Cycs | 12387 | -0.705 | -0.4944 | No | ||
| 41 | CD82 | Cd82 | 12512 | -0.733 | -0.4915 | No | ||
| 42 | CDK2 | Cdk2 | 12758 | -0.793 | -0.4956 | Yes | ||
| 43 | IGF1 | Igf1 | 12887 | -0.824 | -0.4916 | Yes | ||
| 44 | THBS1 | Thbs1 | 13010 | -0.847 | -0.4869 | Yes | ||
| 45 | TP73 | Trp73 | 13089 | -0.863 | -0.4791 | Yes | ||
| 46 | SERPINB5 | Serpinb5 | 13099 | -0.866 | -0.4668 | Yes | ||
| 47 | CASP8 | Casp8 | 13491 | -0.990 | -0.4773 | Yes | ||
| 48 | SIAH1 | Siah1a | 13546 | -1.011 | -0.4658 | Yes | ||
| 49 | GADD45G | Gadd45g | 14056 | -1.231 | -0.4803 | Yes | ||
| 50 | CHEK2 | Chek2 | 14135 | -1.288 | -0.4662 | Yes | ||
| 51 | PIDD1 | Pidd1 | 14140 | -1.289 | -0.4473 | Yes | ||
| 52 | CCND1 | Ccnd1 | 14737 | -1.745 | -0.4598 | Yes | ||
| 53 | IGFBP3 | Igfbp3 | 15038 | -2.223 | -0.4461 | Yes | ||
| 54 | RPRM | Rprm | 15087 | -2.322 | -0.4147 | Yes | ||
| 55 | GTSE1 | Gtse1 | 15334 | -3.185 | -0.3832 | Yes | ||
| 56 | CDK1 | Cdk1 | 15355 | -3.325 | -0.3350 | Yes | ||
| 57 | CHEK1 | Chek1 | 15360 | -3.334 | -0.2856 | Yes | ||
| 58 | CCNE1 | Ccne1 | 15413 | -3.704 | -0.2338 | Yes | ||
| 59 | CCNE2 | Ccne2 | 15435 | -3.837 | -0.1780 | Yes | ||
| 60 | CCNB1 | Ccnb1 | 15467 | -3.995 | -0.1206 | Yes | ||
| 61 | CCNB2 | Ccnb2 | 15480 | -4.070 | -0.0608 | Yes | ||
| 62 | RRM2 | Rrm2 | 15513 | -4.272 | 0.0008 | Yes |