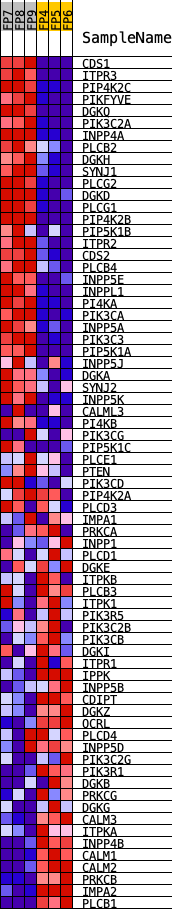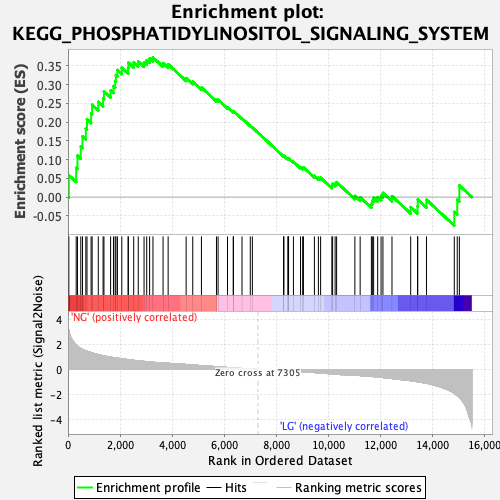
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
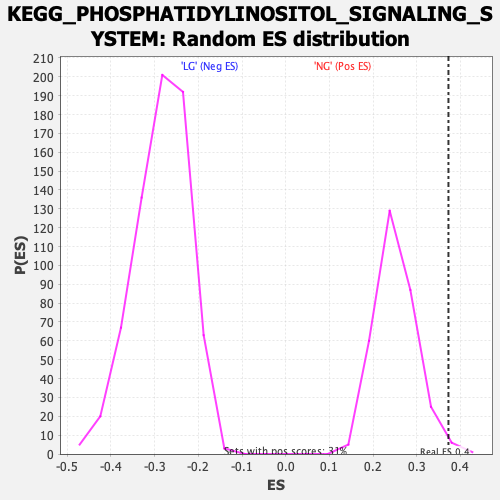
| GeneSet | KEGG_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM |
| Enrichment Score (ES) | 0.37271592 |
| Normalized Enrichment Score (NES) | 1.479455 |
| Nominal p-value | 0.0127795525 |
| FDR q-value | 0.22826222 |
| FWER p-Value | 0.818 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDS1 | Cds1 | 32 | 3.129 | 0.0583 | Yes | ||
| 2 | ITPR3 | Itpr3 | 315 | 1.962 | 0.0779 | Yes | ||
| 3 | PIP4K2C | Pip4k2c | 364 | 1.859 | 0.1107 | Yes | ||
| 4 | PIKFYVE | Pikfyve | 492 | 1.680 | 0.1349 | Yes | ||
| 5 | DGKQ | Dgkq | 557 | 1.616 | 0.1619 | Yes | ||
| 6 | PIK3C2A | Pik3c2a | 686 | 1.486 | 0.1824 | Yes | ||
| 7 | INPP4A | Inpp4a | 730 | 1.451 | 0.2076 | Yes | ||
| 8 | PLCB2 | Plcb2 | 888 | 1.342 | 0.2233 | Yes | ||
| 9 | DGKH | Dgkh | 927 | 1.322 | 0.2464 | Yes | ||
| 10 | SYNJ1 | Synj1 | 1164 | 1.181 | 0.2539 | Yes | ||
| 11 | PLCG2 | Plcg2 | 1349 | 1.099 | 0.2632 | Yes | ||
| 12 | DGKD | Dgkd | 1382 | 1.086 | 0.2821 | Yes | ||
| 13 | PLCG1 | Plcg1 | 1638 | 0.987 | 0.2846 | Yes | ||
| 14 | PIP4K2B | Pip4k2b | 1748 | 0.947 | 0.2959 | Yes | ||
| 15 | PIP5K1B | Pip5k1b | 1812 | 0.928 | 0.3097 | Yes | ||
| 16 | ITPR2 | Itpr2 | 1842 | 0.919 | 0.3255 | Yes | ||
| 17 | CDS2 | Cds2 | 1900 | 0.900 | 0.3392 | Yes | ||
| 18 | PLCB4 | Plcb4 | 2068 | 0.855 | 0.3449 | Yes | ||
| 19 | INPP5E | Inpp5e | 2311 | 0.783 | 0.3444 | Yes | ||
| 20 | INPPL1 | Inppl1 | 2320 | 0.781 | 0.3589 | Yes | ||
| 21 | PI4KA | Pi4ka | 2526 | 0.729 | 0.3597 | Yes | ||
| 22 | PIK3CA | Pik3ca | 2701 | 0.689 | 0.3618 | Yes | ||
| 23 | INPP5A | Inpp5a | 2924 | 0.644 | 0.3598 | Yes | ||
| 24 | PIK3C3 | Pik3c3 | 3026 | 0.624 | 0.3653 | Yes | ||
| 25 | PIP5K1A | Pip5k1a | 3137 | 0.604 | 0.3699 | Yes | ||
| 26 | INPP5J | Inpp5j | 3266 | 0.577 | 0.3727 | Yes | ||
| 27 | DGKA | Dgka | 3655 | 0.513 | 0.3575 | No | ||
| 28 | SYNJ2 | Synj2 | 3850 | 0.493 | 0.3545 | No | ||
| 29 | INPP5K | Inpp5k | 4542 | 0.399 | 0.3175 | No | ||
| 30 | CALML3 | Calml3 | 4796 | 0.359 | 0.3080 | No | ||
| 31 | PI4KB | Pi4kb | 5130 | 0.307 | 0.2924 | No | ||
| 32 | PIK3CG | Pik3cg | 5708 | 0.224 | 0.2594 | No | ||
| 33 | PIP5K1C | Pip5k1c | 5770 | 0.215 | 0.2596 | No | ||
| 34 | PLCE1 | Plce1 | 6133 | 0.159 | 0.2392 | No | ||
| 35 | PTEN | Pten | 6353 | 0.129 | 0.2276 | No | ||
| 36 | PIK3CD | Pik3cd | 6360 | 0.128 | 0.2296 | No | ||
| 37 | PIP4K2A | Pip4k2a | 6690 | 0.079 | 0.2099 | No | ||
| 38 | PLCD3 | Plcd3 | 7008 | 0.043 | 0.1902 | No | ||
| 39 | IMPA1 | Impa1 | 7088 | 0.031 | 0.1857 | No | ||
| 40 | PRKCA | Prkca | 8287 | -0.065 | 0.1094 | No | ||
| 41 | INPP1 | Inpp1 | 8301 | -0.067 | 0.1099 | No | ||
| 42 | PLCD1 | Plcd1 | 8453 | -0.088 | 0.1018 | No | ||
| 43 | DGKE | Dgke | 8474 | -0.091 | 0.1023 | No | ||
| 44 | ITPKB | Itpkb | 8480 | -0.092 | 0.1037 | No | ||
| 45 | PLCB3 | Plcb3 | 8667 | -0.117 | 0.0939 | No | ||
| 46 | ITPK1 | Itpk1 | 8940 | -0.158 | 0.0794 | No | ||
| 47 | PIK3R5 | Pik3r5 | 9023 | -0.172 | 0.0774 | No | ||
| 48 | PIK3C2B | Pik3c2b | 9051 | -0.175 | 0.0790 | No | ||
| 49 | PIK3CB | Pik3cb | 9473 | -0.238 | 0.0564 | No | ||
| 50 | DGKI | Dgki | 9626 | -0.262 | 0.0516 | No | ||
| 51 | ITPR1 | Itpr1 | 9712 | -0.276 | 0.0515 | No | ||
| 52 | IPPK | Ippk | 10145 | -0.347 | 0.0302 | No | ||
| 53 | INPP5B | Inpp5b | 10174 | -0.352 | 0.0352 | No | ||
| 54 | CDIPT | Cdipt | 10270 | -0.366 | 0.0361 | No | ||
| 55 | DGKZ | Dgkz | 10327 | -0.376 | 0.0397 | No | ||
| 56 | OCRL | Ocrl | 11029 | -0.447 | 0.0030 | No | ||
| 57 | PLCD4 | Plcd4 | 11232 | -0.483 | -0.0007 | No | ||
| 58 | INPP5D | Inpp5d | 11664 | -0.549 | -0.0180 | No | ||
| 59 | PIK3C2G | Pik3c2g | 11701 | -0.556 | -0.0096 | No | ||
| 60 | PIK3R1 | Pik3r1 | 11750 | -0.565 | -0.0019 | No | ||
| 61 | DGKB | Dgkb | 11906 | -0.592 | -0.0005 | No | ||
| 62 | PRKCG | Prkcg | 12036 | -0.622 | 0.0032 | No | ||
| 63 | DGKG | Dgkg | 12108 | -0.639 | 0.0110 | No | ||
| 64 | CALM3 | Calm3 | 12456 | -0.718 | 0.0024 | No | ||
| 65 | ITPKA | Itpka | 13175 | -0.892 | -0.0269 | No | ||
| 66 | INPP4B | Inpp4b | 13438 | -0.974 | -0.0250 | No | ||
| 67 | CALM1 | Calm1 | 13447 | -0.977 | -0.0067 | No | ||
| 68 | CALM2 | Calm2 | 13782 | -1.097 | -0.0071 | No | ||
| 69 | PRKCB | Prkcb | 14851 | -1.889 | -0.0398 | No | ||
| 70 | IMPA2 | Impa2 | 14965 | -2.078 | -0.0070 | No | ||
| 71 | PLCB1 | Plcb1 | 15046 | -2.237 | 0.0310 | No |