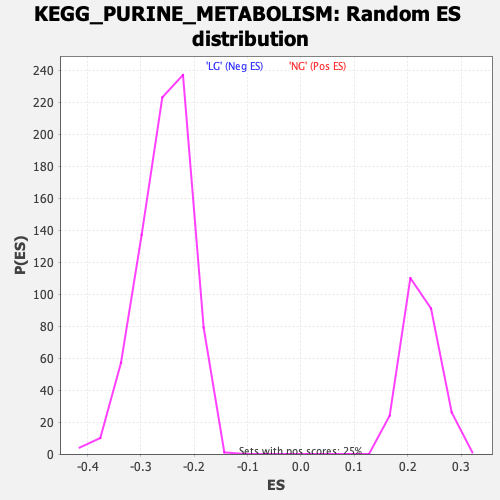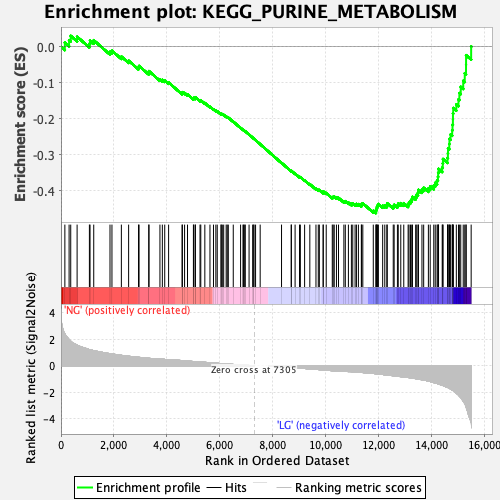
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_PURINE_METABOLISM |
| Enrichment Score (ES) | -0.46225506 |
| Normalized Enrichment Score (NES) | -1.8190451 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.007308117 |
| FWER p-Value | 0.084 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDE8A | Pde8a | 146 | 2.422 | 0.0114 | No | ||
| 2 | PDE3A | Pde3a | 306 | 1.983 | 0.0181 | No | ||
| 3 | GUCY1A2 | Gucy1a2 | 368 | 1.853 | 0.0301 | No | ||
| 4 | ADCY1 | Adcy1 | 609 | 1.555 | 0.0279 | No | ||
| 5 | NPR1 | Npr1 | 1074 | 1.223 | 0.0082 | No | ||
| 6 | PDE5A | Pde5a | 1098 | 1.216 | 0.0172 | No | ||
| 7 | AK4 | Ak4 | 1238 | 1.144 | 0.0180 | No | ||
| 8 | AMPD3 | Ampd3 | 1853 | 0.915 | -0.0140 | No | ||
| 9 | NPR2 | Npr2 | 1927 | 0.891 | -0.0111 | No | ||
| 10 | ADPRM | Adprm | 2283 | 0.790 | -0.0273 | No | ||
| 11 | POLR2A | Polr2a | 2558 | 0.722 | -0.0389 | No | ||
| 12 | POLR3A | Polr3a | 2932 | 0.643 | -0.0576 | No | ||
| 13 | PDE6B | Pde6b | 2952 | 0.638 | -0.0534 | No | ||
| 14 | ADCY2 | Adcy2 | 3321 | 0.565 | -0.0724 | No | ||
| 15 | PRUNE1 | Prune1 | 3326 | 0.565 | -0.0678 | No | ||
| 16 | PKLR | Pklr | 3742 | 0.499 | -0.0905 | No | ||
| 17 | POLR1C | Polr1c | 3833 | 0.496 | -0.0921 | No | ||
| 18 | CANT1 | Cant1 | 3923 | 0.482 | -0.0937 | No | ||
| 19 | NT5C | Nt5c | 4070 | 0.453 | -0.0993 | No | ||
| 20 | PRPS1L1 | Prps1l1 | 4579 | 0.393 | -0.1290 | No | ||
| 21 | POLR3F | Polr3f | 4591 | 0.391 | -0.1263 | No | ||
| 22 | PDE10A | Pde10a | 4674 | 0.379 | -0.1284 | No | ||
| 23 | ENTPD5 | Entpd5 | 4785 | 0.360 | -0.1324 | No | ||
| 24 | GUCY2F | Gucy2f | 5008 | 0.324 | -0.1441 | No | ||
| 25 | ENTPD2 | Entpd2 | 5020 | 0.323 | -0.1420 | No | ||
| 26 | PNP | Pnp | 5076 | 0.316 | -0.1429 | No | ||
| 27 | PKM | Pkm | 5082 | 0.315 | -0.1405 | No | ||
| 28 | ADCY6 | Adcy6 | 5258 | 0.288 | -0.1494 | No | ||
| 29 | PDE7A | Pde7a | 5287 | 0.283 | -0.1487 | No | ||
| 30 | ENTPD8 | Entpd8 | 5440 | 0.262 | -0.1564 | No | ||
| 31 | GUCY2D | Gucy2e | 5634 | 0.234 | -0.1669 | No | ||
| 32 | ADCY10 | Adcy10 | 5767 | 0.216 | -0.1736 | No | ||
| 33 | PDE9A | Pde9a | 5857 | 0.201 | -0.1777 | No | ||
| 34 | PDE4A | Pde4a | 5909 | 0.192 | -0.1793 | No | ||
| 35 | ADCY9 | Adcy9 | 6051 | 0.171 | -0.1870 | No | ||
| 36 | NUDT9 | Nudt9 | 6061 | 0.170 | -0.1861 | No | ||
| 37 | NME5 | Nme5 | 6101 | 0.163 | -0.1873 | No | ||
| 38 | AK1 | Ak1 | 6153 | 0.156 | -0.1893 | No | ||
| 39 | ADCY5 | Adcy5 | 6247 | 0.143 | -0.1941 | No | ||
| 40 | POLR3B | Polr3b | 6288 | 0.138 | -0.1955 | No | ||
| 41 | NT5C2 | Nt5c2 | 6328 | 0.133 | -0.1969 | No | ||
| 42 | AMPD2 | Ampd2 | 6515 | 0.103 | -0.2081 | No | ||
| 43 | POLR3C | Polr3c | 6792 | 0.070 | -0.2254 | No | ||
| 44 | PDE6G | Pde6g | 6883 | 0.057 | -0.2308 | No | ||
| 45 | NUDT2 | Nudt2 | 6905 | 0.054 | -0.2317 | No | ||
| 46 | NME2 | Nme2 | 6947 | 0.049 | -0.2339 | No | ||
| 47 | PRPS1 | Prps1 | 6969 | 0.046 | -0.2349 | No | ||
| 48 | RRM2B | Rrm2b | 7113 | 0.027 | -0.2439 | No | ||
| 49 | ATIC | Atic | 7249 | 0.009 | -0.2526 | No | ||
| 50 | GUCY1B1 | Gucy1b1 | 7290 | 0.003 | -0.2552 | No | ||
| 51 | POLR2E | Polr2e | 7297 | 0.002 | -0.2556 | No | ||
| 52 | ENTPD3 | Entpd3 | 7359 | 0.000 | -0.2596 | No | ||
| 53 | ALLC | Allc | 7533 | 0.000 | -0.2708 | No | ||
| 54 | POLR2K | Polr2k | 8341 | -0.073 | -0.3226 | No | ||
| 55 | DGUOK | Dguok | 8707 | -0.121 | -0.3453 | No | ||
| 56 | POLR1D | Polr1d | 8713 | -0.122 | -0.3446 | No | ||
| 57 | GUCY2C | Gucy2c | 8854 | -0.144 | -0.3525 | No | ||
| 58 | PRPS2 | Prps2 | 9021 | -0.172 | -0.3618 | No | ||
| 59 | POLR2B | Polr2b | 9053 | -0.176 | -0.3623 | No | ||
| 60 | POLR3D | Polr3d | 9214 | -0.198 | -0.3710 | No | ||
| 61 | POLR2H | Polr2h | 9411 | -0.228 | -0.3817 | No | ||
| 62 | POLR2L | Polr2l | 9643 | -0.264 | -0.3945 | No | ||
| 63 | NME7 | Nme7 | 9731 | -0.278 | -0.3977 | No | ||
| 64 | PAPSS2 | Papss2 | 9771 | -0.285 | -0.3978 | No | ||
| 65 | NT5C3A | Nt5c3 | 9916 | -0.313 | -0.4045 | No | ||
| 66 | ADCY3 | Adcy3 | 9917 | -0.313 | -0.4018 | No | ||
| 67 | PDE1B | Pde1b | 10031 | -0.330 | -0.4063 | No | ||
| 68 | POLE4 | Pole4 | 10260 | -0.365 | -0.4180 | No | ||
| 69 | POLR2F | Polr2f | 10293 | -0.371 | -0.4169 | No | ||
| 70 | ADA | Ada | 10334 | -0.376 | -0.4162 | No | ||
| 71 | POLR2G | Polr2g | 10418 | -0.392 | -0.4182 | No | ||
| 72 | NUDT5 | Nudt5 | 10496 | -0.405 | -0.4198 | No | ||
| 73 | PDE11A | Pde11a | 10699 | -0.421 | -0.4293 | No | ||
| 74 | PDE2A | Gm45837 | 10753 | -0.421 | -0.4291 | No | ||
| 75 | ENTPD4 | Entpd4b | 10870 | -0.432 | -0.4329 | No | ||
| 76 | NT5E | Nt5e | 10983 | -0.438 | -0.4364 | No | ||
| 77 | PDE1C | Pde1c | 11035 | -0.448 | -0.4359 | No | ||
| 78 | PAICS | Paics | 11123 | -0.465 | -0.4375 | No | ||
| 79 | HPRT1 | Hprt | 11174 | -0.476 | -0.4367 | No | ||
| 80 | PDE6D | Pde6d | 11248 | -0.486 | -0.4372 | No | ||
| 81 | GUCY1A1 | Gucy1a1 | 11359 | -0.494 | -0.4401 | No | ||
| 82 | ZNRD1 | Znrd1 | 11369 | -0.495 | -0.4365 | No | ||
| 83 | POLR3K | Polr3k | 11415 | -0.506 | -0.4350 | No | ||
| 84 | NME4 | Nme4 | 11811 | -0.575 | -0.4558 | No | ||
| 85 | AK5 | Ak5 | 11912 | -0.593 | -0.4571 | Yes | ||
| 86 | PDE6A | Pde6a | 11918 | -0.594 | -0.4524 | Yes | ||
| 87 | AK7 | Ak7 | 11936 | -0.598 | -0.4483 | Yes | ||
| 88 | GDA | Gda | 11953 | -0.601 | -0.4442 | Yes | ||
| 89 | POLR1A | Polr1a | 11974 | -0.606 | -0.4403 | Yes | ||
| 90 | POLR2C | Polr2c | 12004 | -0.615 | -0.4368 | Yes | ||
| 91 | GUK1 | Guk1 | 12166 | -0.652 | -0.4417 | Yes | ||
| 92 | POLR2I | Polr2i | 12246 | -0.671 | -0.4411 | Yes | ||
| 93 | NME1 | Nme1 | 12325 | -0.688 | -0.4402 | Yes | ||
| 94 | POLR1E | Polr1e | 12338 | -0.692 | -0.4350 | Yes | ||
| 95 | APRT | Aprt | 12566 | -0.745 | -0.4434 | Yes | ||
| 96 | PDE3B | Pde3b | 12600 | -0.753 | -0.4390 | Yes | ||
| 97 | POLR2D | Polr2d | 12724 | -0.786 | -0.4403 | Yes | ||
| 98 | PDE7B | Pde7b | 12755 | -0.791 | -0.4354 | Yes | ||
| 99 | ENTPD1 | Entpd1 | 12851 | -0.814 | -0.4346 | Yes | ||
| 100 | ADK | Adk | 12964 | -0.835 | -0.4346 | Yes | ||
| 101 | NME6 | Nme6 | 13129 | -0.878 | -0.4378 | Yes | ||
| 102 | NT5M | Nt5m | 13162 | -0.888 | -0.4322 | Yes | ||
| 103 | PFAS | Pfas | 13218 | -0.901 | -0.4280 | Yes | ||
| 104 | ENTPD6 | Entpd6 | 13266 | -0.917 | -0.4232 | Yes | ||
| 105 | IMPDH1 | Impdh1 | 13293 | -0.924 | -0.4169 | Yes | ||
| 106 | GMPR2 | Gmpr2 | 13414 | -0.964 | -0.4164 | Yes | ||
| 107 | ADSS2 | Adss | 13455 | -0.980 | -0.4105 | Yes | ||
| 108 | PNPT1 | Pnpt1 | 13503 | -0.996 | -0.4050 | Yes | ||
| 109 | ADCY4 | Adcy4 | 13520 | -1.004 | -0.3974 | Yes | ||
| 110 | PDE6H | Pde6h | 13652 | -1.049 | -0.3969 | Yes | ||
| 111 | ADCY8 | Adcy8 | 13717 | -1.075 | -0.3918 | Yes | ||
| 112 | PAPSS1 | Papss1 | 13895 | -1.146 | -0.3935 | Yes | ||
| 113 | GMPR | Gmpr | 13964 | -1.183 | -0.3877 | Yes | ||
| 114 | XDH | Xdh | 14099 | -1.261 | -0.3855 | Yes | ||
| 115 | PDE1A | Pde1a | 14154 | -1.294 | -0.3779 | Yes | ||
| 116 | POLD2 | Pold2 | 14224 | -1.334 | -0.3709 | Yes | ||
| 117 | FHIT | Fhit | 14257 | -1.350 | -0.3614 | Yes | ||
| 118 | GMPS | Gmps | 14266 | -1.357 | -0.3502 | Yes | ||
| 119 | POLR1B | Polr1b | 14279 | -1.367 | -0.3392 | Yes | ||
| 120 | ADSL | Adsl | 14418 | -1.466 | -0.3356 | Yes | ||
| 121 | GART | Gart | 14439 | -1.477 | -0.3241 | Yes | ||
| 122 | IMPDH2 | Impdh2 | 14447 | -1.483 | -0.3118 | Yes | ||
| 123 | AK2 | Ak2 | 14618 | -1.628 | -0.3089 | Yes | ||
| 124 | POLE2 | Pole2 | 14634 | -1.637 | -0.2957 | Yes | ||
| 125 | POLR3H | Polr3h | 14639 | -1.640 | -0.2819 | Yes | ||
| 126 | POLD1 | Pold1 | 14686 | -1.694 | -0.2703 | Yes | ||
| 127 | ADCY7 | Adcy7 | 14692 | -1.702 | -0.2560 | Yes | ||
| 128 | POLE3 | Pole3 | 14735 | -1.741 | -0.2437 | Yes | ||
| 129 | ENPP1 | Enpp1 | 14795 | -1.830 | -0.2318 | Yes | ||
| 130 | PDE4D | Pde4d | 14808 | -1.840 | -0.2167 | Yes | ||
| 131 | ITPA | Itpa | 14823 | -1.854 | -0.2017 | Yes | ||
| 132 | ADSS1 | Adssl1 | 14825 | -1.855 | -0.1857 | Yes | ||
| 133 | AMPD1 | Ampd1 | 14835 | -1.869 | -0.1702 | Yes | ||
| 134 | PDE4B | Pde4b | 14952 | -2.054 | -0.1601 | Yes | ||
| 135 | PPAT | Ppat | 15033 | -2.208 | -0.1463 | Yes | ||
| 136 | PRIM2 | Prim2 | 15067 | -2.285 | -0.1288 | Yes | ||
| 137 | ENPP3 | Enpp3 | 15119 | -2.389 | -0.1115 | Yes | ||
| 138 | DCK | Dck | 15213 | -2.656 | -0.0947 | Yes | ||
| 139 | POLA1 | Pola1 | 15271 | -2.864 | -0.0737 | Yes | ||
| 140 | RRM1 | Rrm1 | 15319 | -3.083 | -0.0502 | Yes | ||
| 141 | POLE | Pole | 15321 | -3.101 | -0.0236 | Yes | ||
| 142 | RRM2 | Rrm2 | 15513 | -4.272 | 0.0008 | Yes |