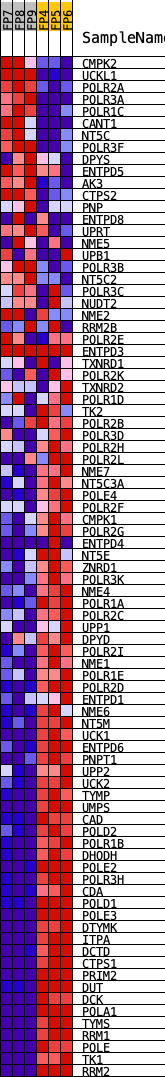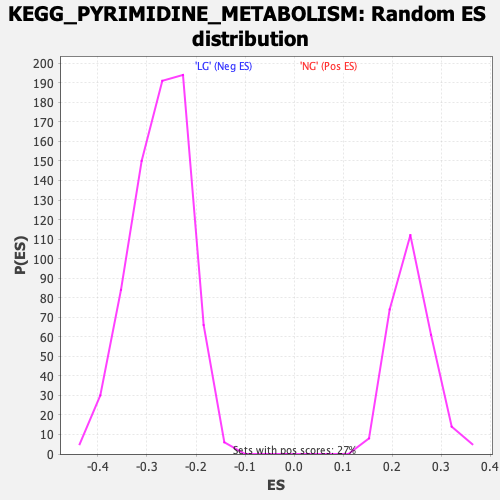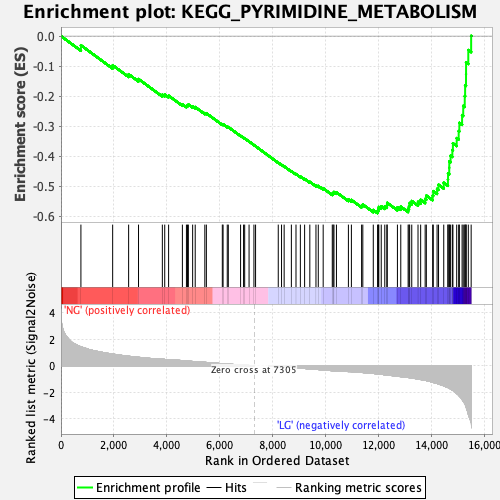
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_PYRIMIDINE_METABOLISM |
| Enrichment Score (ES) | -0.58943987 |
| Normalized Enrichment Score (NES) | -2.1475046 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CMPK2 | Cmpk2 | 753 | 1.436 | -0.0305 | No | ||
| 2 | UCKL1 | Uckl1 | 1954 | 0.881 | -0.0971 | No | ||
| 3 | POLR2A | Polr2a | 2558 | 0.722 | -0.1270 | No | ||
| 4 | POLR3A | Polr3a | 2932 | 0.643 | -0.1430 | No | ||
| 5 | POLR1C | Polr1c | 3833 | 0.496 | -0.1950 | No | ||
| 6 | CANT1 | Cant1 | 3923 | 0.482 | -0.1946 | No | ||
| 7 | NT5C | Nt5c | 4070 | 0.453 | -0.1983 | No | ||
| 8 | POLR3F | Polr3f | 4591 | 0.391 | -0.2270 | No | ||
| 9 | DPYS | Dpys | 4744 | 0.368 | -0.2322 | No | ||
| 10 | ENTPD5 | Entpd5 | 4785 | 0.360 | -0.2302 | No | ||
| 11 | AK3 | Ak3 | 4817 | 0.356 | -0.2277 | No | ||
| 12 | CTPS2 | Ctps2 | 4979 | 0.328 | -0.2340 | No | ||
| 13 | PNP | Pnp | 5076 | 0.316 | -0.2362 | No | ||
| 14 | ENTPD8 | Entpd8 | 5440 | 0.262 | -0.2564 | No | ||
| 15 | UPRT | Uprt | 5498 | 0.254 | -0.2569 | No | ||
| 16 | NME5 | Nme5 | 6101 | 0.163 | -0.2938 | No | ||
| 17 | UPB1 | Upb1 | 6130 | 0.159 | -0.2936 | No | ||
| 18 | POLR3B | Polr3b | 6288 | 0.138 | -0.3020 | No | ||
| 19 | NT5C2 | Nt5c2 | 6328 | 0.133 | -0.3028 | No | ||
| 20 | POLR3C | Polr3c | 6792 | 0.070 | -0.3319 | No | ||
| 21 | NUDT2 | Nudt2 | 6905 | 0.054 | -0.3385 | No | ||
| 22 | NME2 | Nme2 | 6947 | 0.049 | -0.3405 | No | ||
| 23 | RRM2B | Rrm2b | 7113 | 0.027 | -0.3509 | No | ||
| 24 | POLR2E | Polr2e | 7297 | 0.002 | -0.3627 | No | ||
| 25 | ENTPD3 | Entpd3 | 7359 | 0.000 | -0.3667 | No | ||
| 26 | TXNRD1 | Txnrd1 | 8217 | -0.056 | -0.4215 | No | ||
| 27 | POLR2K | Polr2k | 8341 | -0.073 | -0.4285 | No | ||
| 28 | TXNRD2 | Txnrd2 | 8440 | -0.086 | -0.4338 | No | ||
| 29 | POLR1D | Polr1d | 8713 | -0.122 | -0.4498 | No | ||
| 30 | TK2 | Tk2 | 8888 | -0.150 | -0.4592 | No | ||
| 31 | POLR2B | Polr2b | 9053 | -0.176 | -0.4676 | No | ||
| 32 | POLR3D | Polr3d | 9214 | -0.198 | -0.4754 | No | ||
| 33 | POLR2H | Polr2h | 9411 | -0.228 | -0.4852 | No | ||
| 34 | POLR2L | Polr2l | 9643 | -0.264 | -0.4969 | No | ||
| 35 | NME7 | Nme7 | 9731 | -0.278 | -0.4990 | No | ||
| 36 | NT5C3A | Nt5c3 | 9916 | -0.313 | -0.5069 | No | ||
| 37 | POLE4 | Pole4 | 10260 | -0.365 | -0.5245 | No | ||
| 38 | POLR2F | Polr2f | 10293 | -0.371 | -0.5219 | No | ||
| 39 | CMPK1 | Cmpk1 | 10324 | -0.375 | -0.5190 | No | ||
| 40 | POLR2G | Polr2g | 10418 | -0.392 | -0.5201 | No | ||
| 41 | ENTPD4 | Entpd4b | 10870 | -0.432 | -0.5438 | No | ||
| 42 | NT5E | Nt5e | 10983 | -0.438 | -0.5455 | No | ||
| 43 | ZNRD1 | Znrd1 | 11369 | -0.495 | -0.5642 | No | ||
| 44 | POLR3K | Polr3k | 11415 | -0.506 | -0.5607 | No | ||
| 45 | NME4 | Nme4 | 11811 | -0.575 | -0.5789 | No | ||
| 46 | POLR1A | Polr1a | 11974 | -0.606 | -0.5818 | Yes | ||
| 47 | POLR2C | Polr2c | 12004 | -0.615 | -0.5758 | Yes | ||
| 48 | UPP1 | Upp1 | 12020 | -0.618 | -0.5690 | Yes | ||
| 49 | DPYD | Dpyd | 12112 | -0.640 | -0.5667 | Yes | ||
| 50 | POLR2I | Polr2i | 12246 | -0.671 | -0.5668 | Yes | ||
| 51 | NME1 | Nme1 | 12325 | -0.688 | -0.5632 | Yes | ||
| 52 | POLR1E | Polr1e | 12338 | -0.692 | -0.5551 | Yes | ||
| 53 | POLR2D | Polr2d | 12724 | -0.786 | -0.5701 | Yes | ||
| 54 | ENTPD1 | Entpd1 | 12851 | -0.814 | -0.5679 | Yes | ||
| 55 | NME6 | Nme6 | 13129 | -0.878 | -0.5747 | Yes | ||
| 56 | NT5M | Nt5m | 13162 | -0.888 | -0.5655 | Yes | ||
| 57 | UCK1 | Uck1 | 13187 | -0.896 | -0.5557 | Yes | ||
| 58 | ENTPD6 | Entpd6 | 13266 | -0.917 | -0.5492 | Yes | ||
| 59 | PNPT1 | Pnpt1 | 13503 | -0.996 | -0.5518 | Yes | ||
| 60 | UPP2 | Upp2 | 13603 | -1.034 | -0.5451 | Yes | ||
| 61 | UCK2 | Uck2 | 13773 | -1.094 | -0.5422 | Yes | ||
| 62 | TYMP | Tymp | 13818 | -1.112 | -0.5309 | Yes | ||
| 63 | UMPS | Umps | 14057 | -1.233 | -0.5307 | Yes | ||
| 64 | CAD | Cad | 14080 | -1.249 | -0.5163 | Yes | ||
| 65 | POLD2 | Pold2 | 14224 | -1.334 | -0.5086 | Yes | ||
| 66 | POLR1B | Polr1b | 14279 | -1.367 | -0.4948 | Yes | ||
| 67 | DHODH | Dhodh | 14476 | -1.514 | -0.4882 | Yes | ||
| 68 | POLE2 | Pole2 | 14634 | -1.637 | -0.4776 | Yes | ||
| 69 | POLR3H | Polr3h | 14639 | -1.640 | -0.4571 | Yes | ||
| 70 | CDA | Cda | 14684 | -1.692 | -0.4385 | Yes | ||
| 71 | POLD1 | Pold1 | 14686 | -1.694 | -0.4170 | Yes | ||
| 72 | POLE3 | Pole3 | 14735 | -1.741 | -0.3981 | Yes | ||
| 73 | DTYMK | Dtymk | 14802 | -1.834 | -0.3791 | Yes | ||
| 74 | ITPA | Itpa | 14823 | -1.854 | -0.3568 | Yes | ||
| 75 | DCTD | Dctd | 14966 | -2.078 | -0.3397 | Yes | ||
| 76 | CTPS1 | Ctps | 15041 | -2.228 | -0.3162 | Yes | ||
| 77 | PRIM2 | Prim2 | 15067 | -2.285 | -0.2888 | Yes | ||
| 78 | DUT | Dut | 15170 | -2.533 | -0.2633 | Yes | ||
| 79 | DCK | Dck | 15213 | -2.656 | -0.2323 | Yes | ||
| 80 | POLA1 | Pola1 | 15271 | -2.864 | -0.1997 | Yes | ||
| 81 | TYMS | Tyms | 15285 | -2.895 | -0.1638 | Yes | ||
| 82 | RRM1 | Rrm1 | 15319 | -3.083 | -0.1268 | Yes | ||
| 83 | POLE | Pole | 15321 | -3.101 | -0.0875 | Yes | ||
| 84 | TK1 | Tk1 | 15405 | -3.658 | -0.0465 | Yes | ||
| 85 | RRM2 | Rrm2 | 15513 | -4.272 | 0.0008 | Yes |