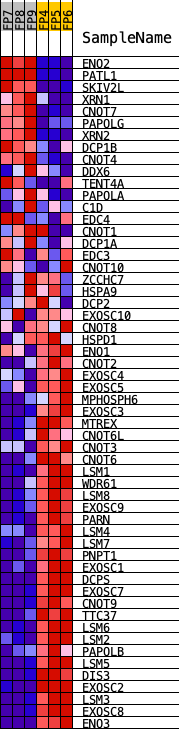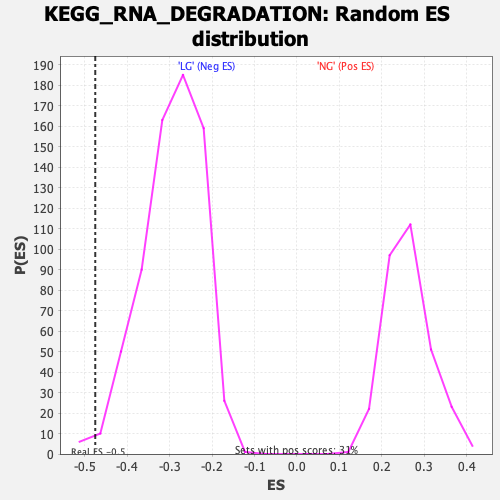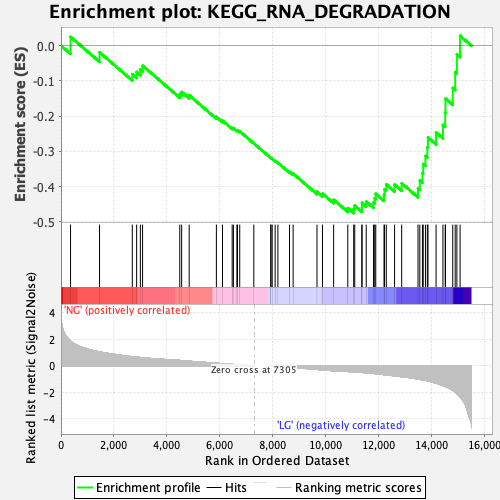
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_RNA_DEGRADATION |
| Enrichment Score (ES) | -0.4750402 |
| Normalized Enrichment Score (NES) | -1.6192281 |
| Nominal p-value | 0.010144928 |
| FDR q-value | 0.047082596 |
| FWER p-Value | 0.621 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ENO2 | Eno2 | 361 | 1.866 | 0.0249 | No | ||
| 2 | PATL1 | Patl1 | 1457 | 1.051 | -0.0187 | No | ||
| 3 | SKIV2L | Skiv2l | 2696 | 0.689 | -0.0810 | No | ||
| 4 | XRN1 | Xrn1 | 2861 | 0.656 | -0.0746 | No | ||
| 5 | CNOT7 | Cnot7 | 3002 | 0.629 | -0.0674 | No | ||
| 6 | PAPOLG | Papolg | 3085 | 0.613 | -0.0568 | No | ||
| 7 | XRN2 | Xrn2 | 4492 | 0.407 | -0.1372 | No | ||
| 8 | DCP1B | Dcp1b | 4563 | 0.395 | -0.1315 | No | ||
| 9 | CNOT4 | Cnot4 | 4848 | 0.351 | -0.1408 | No | ||
| 10 | DDX6 | Ddx6 | 5877 | 0.197 | -0.2022 | No | ||
| 11 | TENT4A | Tent4a | 6108 | 0.162 | -0.2129 | No | ||
| 12 | PAPOLA | Papola | 6474 | 0.111 | -0.2336 | No | ||
| 13 | C1D | C1d | 6521 | 0.102 | -0.2339 | No | ||
| 14 | EDC4 | Edc4 | 6653 | 0.084 | -0.2402 | No | ||
| 15 | CNOT1 | Cnot1 | 6675 | 0.082 | -0.2394 | No | ||
| 16 | DCP1A | Dcp1a | 6760 | 0.074 | -0.2430 | No | ||
| 17 | EDC3 | Edc3 | 7293 | 0.003 | -0.2773 | No | ||
| 18 | CNOT10 | Cnot10 | 7934 | -0.012 | -0.3183 | No | ||
| 19 | ZCCHC7 | Zcchc7 | 7940 | -0.013 | -0.3183 | No | ||
| 20 | HSPA9 | Hspa9 | 7988 | -0.020 | -0.3208 | No | ||
| 21 | DCP2 | Dcp2 | 8099 | -0.037 | -0.3270 | No | ||
| 22 | EXOSC10 | Exosc10 | 8204 | -0.054 | -0.3323 | No | ||
| 23 | CNOT8 | Cnot8 | 8643 | -0.114 | -0.3577 | No | ||
| 24 | HSPD1 | Hspd1 | 8780 | -0.133 | -0.3630 | No | ||
| 25 | ENO1 | Eno1 | 9682 | -0.271 | -0.4142 | No | ||
| 26 | CNOT2 | Cnot2 | 9889 | -0.306 | -0.4196 | No | ||
| 27 | EXOSC4 | Exosc4 | 10312 | -0.374 | -0.4373 | No | ||
| 28 | EXOSC5 | Exosc5 | 10847 | -0.429 | -0.4607 | No | ||
| 29 | MPHOSPH6 | Mphosph6 | 11070 | -0.455 | -0.4633 | Yes | ||
| 30 | EXOSC3 | Exosc3 | 11110 | -0.462 | -0.4539 | Yes | ||
| 31 | MTREX | Mtrex | 11381 | -0.497 | -0.4585 | Yes | ||
| 32 | CNOT6L | Cnot6l | 11384 | -0.498 | -0.4457 | Yes | ||
| 33 | CNOT3 | Cnot3 | 11544 | -0.525 | -0.4425 | Yes | ||
| 34 | CNOT6 | Cnot6 | 11818 | -0.576 | -0.4452 | Yes | ||
| 35 | LSM1 | Lsm1 | 11865 | -0.584 | -0.4331 | Yes | ||
| 36 | WDR61 | Wdr61 | 11905 | -0.592 | -0.4203 | Yes | ||
| 37 | LSM8 | Lsm8 | 12217 | -0.664 | -0.4233 | Yes | ||
| 38 | EXOSC9 | Exosc9 | 12237 | -0.669 | -0.4072 | Yes | ||
| 39 | PARN | Parn | 12303 | -0.683 | -0.3938 | Yes | ||
| 40 | LSM4 | Lsm4 | 12616 | -0.760 | -0.3943 | Yes | ||
| 41 | LSM7 | Lsm7 | 12885 | -0.823 | -0.3904 | Yes | ||
| 42 | PNPT1 | Pnpt1 | 13503 | -0.996 | -0.4045 | Yes | ||
| 43 | EXOSC1 | Exosc1 | 13571 | -1.019 | -0.3825 | Yes | ||
| 44 | DCPS | Dcps | 13671 | -1.057 | -0.3616 | Yes | ||
| 45 | EXOSC7 | Exosc7 | 13699 | -1.069 | -0.3357 | Yes | ||
| 46 | CNOT9 | Cnot9 | 13784 | -1.098 | -0.3128 | Yes | ||
| 47 | TTC37 | Ttc37 | 13855 | -1.127 | -0.2882 | Yes | ||
| 48 | LSM6 | Lsm6 | 13880 | -1.138 | -0.2603 | Yes | ||
| 49 | LSM2 | Lsm2 | 14186 | -1.310 | -0.2462 | Yes | ||
| 50 | PAPOLB | Papolb | 14446 | -1.482 | -0.2247 | Yes | ||
| 51 | LSM5 | Lsm5 | 14532 | -1.557 | -0.1899 | Yes | ||
| 52 | DIS3 | Dis3 | 14542 | -1.565 | -0.1501 | Yes | ||
| 53 | EXOSC2 | Exosc2 | 14817 | -1.846 | -0.1201 | Yes | ||
| 54 | LSM3 | Lsm3 | 14909 | -1.980 | -0.0748 | Yes | ||
| 55 | EXOSC8 | Exosc8 | 14973 | -2.093 | -0.0248 | Yes | ||
| 56 | ENO3 | Eno3 | 15094 | -2.337 | 0.0279 | Yes |