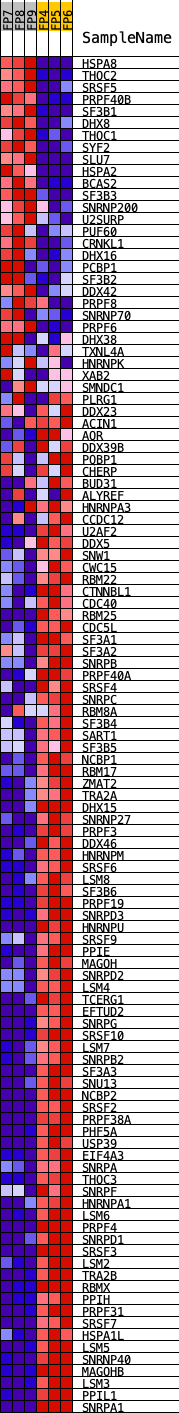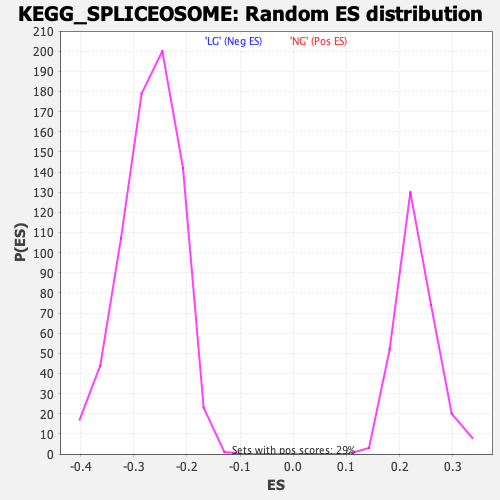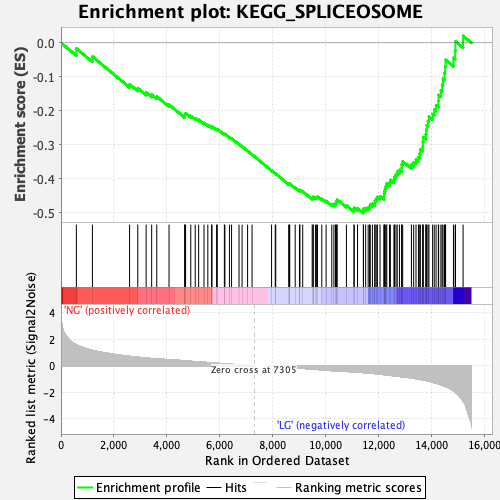
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | LG |
| GeneSet | KEGG_SPLICEOSOME |
| Enrichment Score (ES) | -0.50180817 |
| Normalized Enrichment Score (NES) | -1.8729767 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.004560605 |
| FWER p-Value | 0.043 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HSPA8 | Hspa8 | 581 | 1.581 | -0.0167 | No | ||
| 2 | THOC2 | Thoc2 | 1188 | 1.169 | -0.0404 | No | ||
| 3 | SRSF5 | Srsf5 | 2594 | 0.715 | -0.1221 | No | ||
| 4 | PRPF40B | Prpf40b | 2903 | 0.648 | -0.1334 | No | ||
| 5 | SF3B1 | Sf3b1 | 3220 | 0.586 | -0.1461 | No | ||
| 6 | DHX8 | Dhx8 | 3429 | 0.547 | -0.1523 | No | ||
| 7 | THOC1 | Thoc1 | 3622 | 0.518 | -0.1579 | No | ||
| 8 | SYF2 | Syf2 | 4086 | 0.450 | -0.1820 | No | ||
| 9 | SLU7 | Slu7 | 4678 | 0.379 | -0.2153 | No | ||
| 10 | HSPA2 | Hspa2 | 4685 | 0.377 | -0.2106 | No | ||
| 11 | BCAS2 | Bcas2 | 4712 | 0.374 | -0.2074 | No | ||
| 12 | SF3B3 | Sf3b3 | 4908 | 0.340 | -0.2155 | No | ||
| 13 | SNRNP200 | Snrnp200 | 5077 | 0.316 | -0.2222 | No | ||
| 14 | U2SURP | U2surp | 5206 | 0.294 | -0.2266 | No | ||
| 15 | PUF60 | Puf60 | 5409 | 0.267 | -0.2361 | No | ||
| 16 | CRNKL1 | Crnkl1 | 5553 | 0.246 | -0.2421 | No | ||
| 17 | DHX16 | Dhx16 | 5695 | 0.226 | -0.2483 | No | ||
| 18 | PCBP1 | Pcbp1 | 5710 | 0.224 | -0.2462 | No | ||
| 19 | SF3B2 | Sf3b2 | 5884 | 0.195 | -0.2548 | No | ||
| 20 | DDX42 | Ddx42 | 5911 | 0.191 | -0.2540 | No | ||
| 21 | PRPF8 | Prpf8 | 6179 | 0.152 | -0.2693 | No | ||
| 22 | SNRNP70 | Snrnp70 | 6199 | 0.150 | -0.2685 | No | ||
| 23 | PRPF6 | Prpf6 | 6371 | 0.126 | -0.2779 | No | ||
| 24 | DHX38 | Dhx38 | 6451 | 0.114 | -0.2815 | No | ||
| 25 | TXNL4A | Txnl4a | 6733 | 0.078 | -0.2987 | No | ||
| 26 | HNRNPK | Hnrnpk | 6849 | 0.059 | -0.3054 | No | ||
| 27 | XAB2 | Xab2 | 7057 | 0.034 | -0.3184 | No | ||
| 28 | SMNDC1 | Smndc1 | 7226 | 0.012 | -0.3291 | No | ||
| 29 | PLRG1 | Plrg1 | 7964 | -0.016 | -0.3767 | No | ||
| 30 | DDX23 | Ddx23 | 8103 | -0.038 | -0.3852 | No | ||
| 31 | ACIN1 | Acin1 | 8122 | -0.042 | -0.3858 | No | ||
| 32 | AQR | Aqr | 8611 | -0.109 | -0.4160 | No | ||
| 33 | DDX39B | Ddx39b | 8645 | -0.114 | -0.4166 | No | ||
| 34 | PQBP1 | Pqbp1 | 8649 | -0.114 | -0.4153 | No | ||
| 35 | CHERP | Cherp | 8858 | -0.145 | -0.4268 | No | ||
| 36 | BUD31 | Bud31 | 9019 | -0.172 | -0.4349 | No | ||
| 37 | ALYREF | 4931428L18Rik | 9037 | -0.175 | -0.4337 | No | ||
| 38 | HNRNPA3 | Gm17190 | 9142 | -0.187 | -0.4380 | No | ||
| 39 | CCDC12 | Ccdc12 | 9500 | -0.242 | -0.4579 | No | ||
| 40 | U2AF2 | U2af2 | 9535 | -0.248 | -0.4568 | No | ||
| 41 | DDX5 | Ddx5 | 9538 | -0.249 | -0.4536 | No | ||
| 42 | SNW1 | Snw1 | 9624 | -0.262 | -0.4557 | No | ||
| 43 | CWC15 | Cwc15 | 9667 | -0.269 | -0.4548 | No | ||
| 44 | RBM22 | Rbm22 | 9703 | -0.274 | -0.4534 | No | ||
| 45 | CTNNBL1 | Ctnnbl1 | 9864 | -0.300 | -0.4598 | No | ||
| 46 | CDC40 | Cdc40 | 10027 | -0.330 | -0.4659 | No | ||
| 47 | RBM25 | Rbm25 | 10242 | -0.363 | -0.4750 | No | ||
| 48 | CDC5L | Cdc5l | 10322 | -0.374 | -0.4751 | No | ||
| 49 | SF3A1 | Sf3a1 | 10383 | -0.386 | -0.4739 | No | ||
| 50 | SF3A2 | Sf3a2 | 10405 | -0.390 | -0.4701 | No | ||
| 51 | SNRPB | Snrpb | 10415 | -0.392 | -0.4654 | No | ||
| 52 | PRPF40A | Prpf40a | 10447 | -0.397 | -0.4622 | No | ||
| 53 | SRSF4 | Srsf4 | 10795 | -0.423 | -0.4791 | No | ||
| 54 | SNRPC | Snrpc | 11081 | -0.457 | -0.4915 | No | ||
| 55 | RBM8A | Rbm8a | 11090 | -0.460 | -0.4859 | No | ||
| 56 | SF3B4 | Sf3b4 | 11215 | -0.478 | -0.4875 | No | ||
| 57 | SART1 | Sart1 | 11436 | -0.508 | -0.4950 | Yes | ||
| 58 | SF3B5 | Sf3b5 | 11440 | -0.509 | -0.4885 | Yes | ||
| 59 | NCBP1 | Ncbp1 | 11523 | -0.521 | -0.4869 | Yes | ||
| 60 | RBM17 | Rbm17 | 11618 | -0.538 | -0.4858 | Yes | ||
| 61 | ZMAT2 | Zmat2 | 11670 | -0.549 | -0.4818 | Yes | ||
| 62 | TRA2A | Tra2a | 11696 | -0.555 | -0.4760 | Yes | ||
| 63 | DHX15 | Dhx15 | 11778 | -0.568 | -0.4737 | Yes | ||
| 64 | SNRNP27 | Snrnp27 | 11869 | -0.585 | -0.4718 | Yes | ||
| 65 | PRPF3 | Prpf3 | 11874 | -0.588 | -0.4642 | Yes | ||
| 66 | DDX46 | Ddx46 | 11922 | -0.595 | -0.4593 | Yes | ||
| 67 | HNRNPM | Hnrnpm | 11967 | -0.605 | -0.4541 | Yes | ||
| 68 | SRSF6 | Srsf6 | 12066 | -0.629 | -0.4521 | Yes | ||
| 69 | LSM8 | Lsm8 | 12217 | -0.664 | -0.4530 | Yes | ||
| 70 | SF3B6 | Sf3b6 | 12220 | -0.665 | -0.4443 | Yes | ||
| 71 | PRPF19 | Prpf19 | 12234 | -0.668 | -0.4362 | Yes | ||
| 72 | SNRPD3 | Snrpd3 | 12260 | -0.675 | -0.4289 | Yes | ||
| 73 | HNRNPU | Hnrnpu | 12289 | -0.681 | -0.4216 | Yes | ||
| 74 | SRSF9 | Srsf9 | 12322 | -0.687 | -0.4146 | Yes | ||
| 75 | PPIE | Ppie | 12426 | -0.713 | -0.4118 | Yes | ||
| 76 | MAGOH | Magoh | 12457 | -0.718 | -0.4041 | Yes | ||
| 77 | SNRPD2 | Snrpd2 | 12592 | -0.751 | -0.4029 | Yes | ||
| 78 | LSM4 | Lsm4 | 12616 | -0.760 | -0.3942 | Yes | ||
| 79 | TCERG1 | Tcerg1 | 12670 | -0.774 | -0.3874 | Yes | ||
| 80 | EFTUD2 | Eftud2 | 12716 | -0.784 | -0.3798 | Yes | ||
| 81 | SNRPG | Snrpg | 12800 | -0.802 | -0.3746 | Yes | ||
| 82 | SRSF10 | Srsf10 | 12883 | -0.823 | -0.3689 | Yes | ||
| 83 | LSM7 | Lsm7 | 12885 | -0.823 | -0.3580 | Yes | ||
| 84 | SNRPB2 | Snrpb2 | 12923 | -0.828 | -0.3494 | Yes | ||
| 85 | SF3A3 | Sf3a3 | 13253 | -0.914 | -0.3586 | Yes | ||
| 86 | SNU13 | Snu13 | 13339 | -0.943 | -0.3516 | Yes | ||
| 87 | NCBP2 | Ncbp2 | 13427 | -0.972 | -0.3443 | Yes | ||
| 88 | SRSF2 | Srsf2 | 13519 | -1.004 | -0.3368 | Yes | ||
| 89 | PRPF38A | Prpf38a | 13568 | -1.018 | -0.3264 | Yes | ||
| 90 | PHF5A | Phf5a | 13591 | -1.027 | -0.3141 | Yes | ||
| 91 | USP39 | Usp39 | 13682 | -1.060 | -0.3059 | Yes | ||
| 92 | EIF4A3 | Eif4a3 | 13687 | -1.062 | -0.2920 | Yes | ||
| 93 | SNRPA | Snrpa | 13700 | -1.069 | -0.2785 | Yes | ||
| 94 | THOC3 | Thoc3 | 13797 | -1.103 | -0.2701 | Yes | ||
| 95 | SNRPF | Snrpf | 13810 | -1.109 | -0.2561 | Yes | ||
| 96 | HNRNPA1 | Hnrnpa1 | 13833 | -1.118 | -0.2427 | Yes | ||
| 97 | LSM6 | Lsm6 | 13880 | -1.138 | -0.2305 | Yes | ||
| 98 | PRPF4 | Prpf4 | 13916 | -1.159 | -0.2173 | Yes | ||
| 99 | SNRPD1 | Snrpd1 | 14055 | -1.231 | -0.2099 | Yes | ||
| 100 | SRSF3 | Srsf3 | 14117 | -1.277 | -0.1969 | Yes | ||
| 101 | LSM2 | Lsm2 | 14186 | -1.310 | -0.1839 | Yes | ||
| 102 | TRA2B | Tra2b | 14272 | -1.361 | -0.1713 | Yes | ||
| 103 | RBMX | Rbmx | 14283 | -1.368 | -0.1537 | Yes | ||
| 104 | PPIH | Gm7879 | 14368 | -1.432 | -0.1401 | Yes | ||
| 105 | PRPF31 | Prpf31 | 14414 | -1.464 | -0.1235 | Yes | ||
| 106 | SRSF7 | Srsf7 | 14449 | -1.485 | -0.1060 | Yes | ||
| 107 | HSPA1L | Hspa1l | 14505 | -1.535 | -0.0891 | Yes | ||
| 108 | LSM5 | Lsm5 | 14532 | -1.557 | -0.0701 | Yes | ||
| 109 | SNRNP40 | Snrnp40 | 14546 | -1.566 | -0.0501 | Yes | ||
| 110 | MAGOHB | Magohb | 14844 | -1.880 | -0.0443 | Yes | ||
| 111 | LSM3 | Lsm3 | 14909 | -1.980 | -0.0221 | Yes | ||
| 112 | PPIL1 | Ppil1 | 14919 | -2.007 | 0.0040 | Yes | ||
| 113 | SNRPA1 | Snrpa1 | 15208 | -2.649 | 0.0206 | Yes |