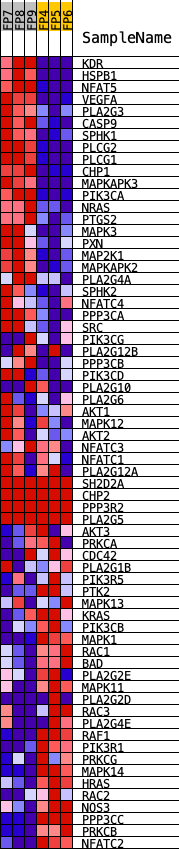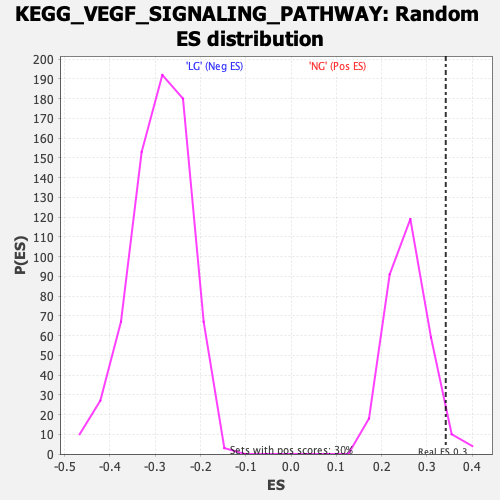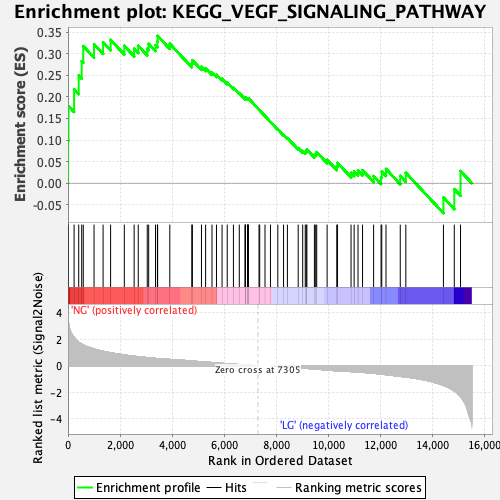
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NG_versus_LG.phenotype.cls#NG_versus_LG_repos |
| Phenotype | phenotype.cls#NG_versus_LG_repos |
| Upregulated in class | NG |
| GeneSet | KEGG_VEGF_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.3414762 |
| Normalized Enrichment Score (NES) | 1.3251008 |
| Nominal p-value | 0.03986711 |
| FDR q-value | 0.30607215 |
| FWER p-Value | 0.992 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDR | Kdr | 4 | 4.063 | 0.0995 | Yes | ||
| 2 | HSPB1 | Hspb1 | 20 | 3.266 | 0.1788 | Yes | ||
| 3 | NFAT5 | Nfat5 | 234 | 2.145 | 0.2177 | Yes | ||
| 4 | VEGFA | Vegfa | 412 | 1.773 | 0.2498 | Yes | ||
| 5 | PLA2G3 | Pla2g3 | 527 | 1.644 | 0.2828 | Yes | ||
| 6 | CASP9 | Casp9 | 587 | 1.576 | 0.3176 | Yes | ||
| 7 | SPHK1 | Sphk1 | 1001 | 1.267 | 0.3220 | Yes | ||
| 8 | PLCG2 | Plcg2 | 1349 | 1.099 | 0.3266 | Yes | ||
| 9 | PLCG1 | Plcg1 | 1638 | 0.987 | 0.3322 | Yes | ||
| 10 | CHP1 | Chp1 | 2165 | 0.825 | 0.3184 | Yes | ||
| 11 | MAPKAPK3 | Mapkapk3 | 2543 | 0.725 | 0.3118 | Yes | ||
| 12 | PIK3CA | Pik3ca | 2701 | 0.689 | 0.3186 | Yes | ||
| 13 | NRAS | Nras | 3047 | 0.622 | 0.3116 | Yes | ||
| 14 | PTGS2 | Ptgs2 | 3102 | 0.610 | 0.3230 | Yes | ||
| 15 | MAPK3 | Mapk3 | 3365 | 0.559 | 0.3198 | Yes | ||
| 16 | PXN | Pxn | 3438 | 0.546 | 0.3286 | Yes | ||
| 17 | MAP2K1 | Map2k1 | 3446 | 0.545 | 0.3415 | Yes | ||
| 18 | MAPKAPK2 | Mapkapk2 | 3914 | 0.484 | 0.3231 | No | ||
| 19 | PLA2G4A | Pla2g4a | 4759 | 0.366 | 0.2775 | No | ||
| 20 | SPHK2 | Sphk2 | 4782 | 0.361 | 0.2850 | No | ||
| 21 | NFATC4 | Nfatc4 | 5131 | 0.307 | 0.2700 | No | ||
| 22 | PPP3CA | Ppp3ca | 5291 | 0.283 | 0.2667 | No | ||
| 23 | SRC | Src | 5536 | 0.248 | 0.2570 | No | ||
| 24 | PIK3CG | Pik3cg | 5708 | 0.224 | 0.2514 | No | ||
| 25 | PLA2G12B | Pla2g12b | 5923 | 0.189 | 0.2422 | No | ||
| 26 | PPP3CB | Ppp3cb | 6122 | 0.159 | 0.2333 | No | ||
| 27 | PIK3CD | Pik3cd | 6360 | 0.128 | 0.2211 | No | ||
| 28 | PLA2G10 | Pla2g10 | 6585 | 0.093 | 0.2089 | No | ||
| 29 | PLA2G6 | Pla2g6 | 6809 | 0.067 | 0.1961 | No | ||
| 30 | AKT1 | Akt1 | 6815 | 0.066 | 0.1974 | No | ||
| 31 | MAPK12 | Mapk12 | 6820 | 0.065 | 0.1988 | No | ||
| 32 | AKT2 | Akt2 | 6903 | 0.054 | 0.1948 | No | ||
| 33 | NFATC3 | Nfatc3 | 6904 | 0.054 | 0.1961 | No | ||
| 34 | NFATC1 | Nfatc1 | 6933 | 0.051 | 0.1956 | No | ||
| 35 | PLA2G12A | Pla2g12a | 6934 | 0.051 | 0.1968 | No | ||
| 36 | SH2D2A | Sh2d2a | 7344 | 0.000 | 0.1703 | No | ||
| 37 | CHP2 | Chp2 | 7370 | 0.000 | 0.1687 | No | ||
| 38 | PPP3R2 | Ppp3r2 | 7574 | 0.000 | 0.1556 | No | ||
| 39 | PLA2G5 | Pla2g5 | 7784 | 0.000 | 0.1421 | No | ||
| 40 | AKT3 | Akt3 | 8066 | -0.032 | 0.1247 | No | ||
| 41 | PRKCA | Prkca | 8287 | -0.065 | 0.1121 | No | ||
| 42 | CDC42 | Cdc42 | 8435 | -0.086 | 0.1047 | No | ||
| 43 | PLA2G1B | Pla2g1b | 8850 | -0.143 | 0.0814 | No | ||
| 44 | PIK3R5 | Pik3r5 | 9023 | -0.172 | 0.0745 | No | ||
| 45 | PTK2 | Ptk2 | 9121 | -0.184 | 0.0727 | No | ||
| 46 | MAPK13 | Mapk13 | 9159 | -0.190 | 0.0750 | No | ||
| 47 | KRAS | Kras | 9184 | -0.193 | 0.0782 | No | ||
| 48 | PIK3CB | Pik3cb | 9473 | -0.238 | 0.0654 | No | ||
| 49 | MAPK1 | Mapk1 | 9523 | -0.246 | 0.0683 | No | ||
| 50 | RAC1 | Rac1 | 9562 | -0.252 | 0.0721 | No | ||
| 51 | BAD | Bad | 9962 | -0.319 | 0.0541 | No | ||
| 52 | PLA2G2E | Pla2g2e | 10333 | -0.376 | 0.0394 | No | ||
| 53 | MAPK11 | Mapk11 | 10363 | -0.382 | 0.0469 | No | ||
| 54 | PLA2G2D | Pla2g2d | 10882 | -0.432 | 0.0240 | No | ||
| 55 | RAC3 | Rac3 | 11001 | -0.441 | 0.0272 | No | ||
| 56 | PLA2G4E | Pla2g4e | 11150 | -0.470 | 0.0292 | No | ||
| 57 | RAF1 | Raf1 | 11324 | -0.492 | 0.0301 | No | ||
| 58 | PIK3R1 | Pik3r1 | 11750 | -0.565 | 0.0165 | No | ||
| 59 | PRKCG | Prkcg | 12036 | -0.622 | 0.0133 | No | ||
| 60 | MAPK14 | Mapk14 | 12063 | -0.629 | 0.0271 | No | ||
| 61 | HRAS | Hras | 12226 | -0.666 | 0.0329 | No | ||
| 62 | RAC2 | Rac2 | 12774 | -0.795 | 0.0171 | No | ||
| 63 | NOS3 | Nos3 | 12988 | -0.841 | 0.0240 | No | ||
| 64 | PPP3CC | Ppp3cc | 14434 | -1.476 | -0.0332 | No | ||
| 65 | PRKCB | Prkcb | 14851 | -1.889 | -0.0138 | No | ||
| 66 | NFATC2 | Nfatc2 | 15091 | -2.333 | 0.0281 | No |