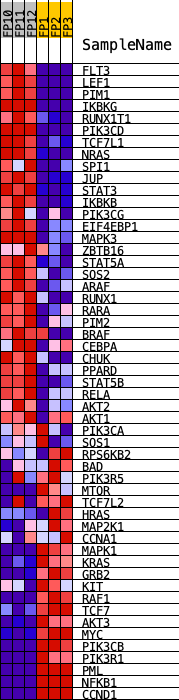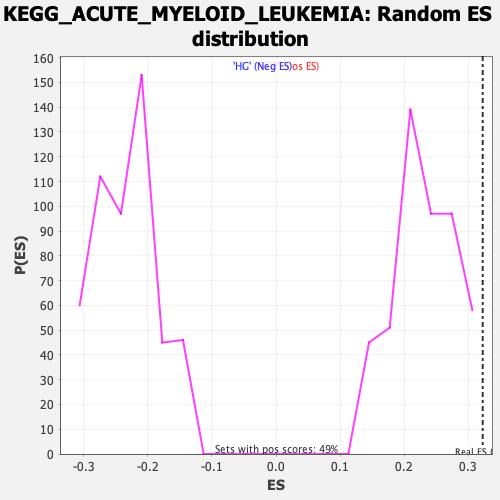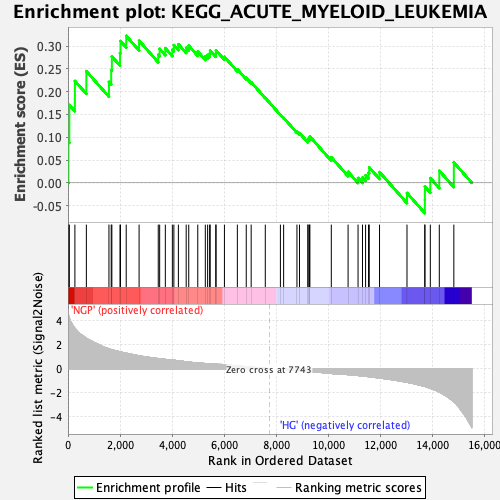
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_ACUTE_MYELOID_LEUKEMIA |
| Enrichment Score (ES) | 0.32256338 |
| Normalized Enrichment Score (NES) | 1.3930627 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2735 |
| FWER p-Value | 0.856 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FLT3 | Flt3 | 17 | 4.618 | 0.0898 | Yes | ||
| 2 | LEF1 | Lef1 | 52 | 4.231 | 0.1709 | Yes | ||
| 3 | PIM1 | Pim1 | 265 | 3.373 | 0.2236 | Yes | ||
| 4 | IKBKG | Ikbkg | 706 | 2.530 | 0.2450 | Yes | ||
| 5 | RUNX1T1 | Runx1t1 | 1574 | 1.663 | 0.2216 | Yes | ||
| 6 | PIK3CD | Pik3cd | 1661 | 1.603 | 0.2476 | Yes | ||
| 7 | TCF7L1 | Tcf7l1 | 1686 | 1.589 | 0.2774 | Yes | ||
| 8 | NRAS | Nras | 1999 | 1.405 | 0.2849 | Yes | ||
| 9 | SPI1 | Spi1 | 2018 | 1.398 | 0.3112 | Yes | ||
| 10 | JUP | Jup | 2238 | 1.295 | 0.3226 | Yes | ||
| 11 | STAT3 | Stat3 | 2733 | 1.085 | 0.3120 | No | ||
| 12 | IKBKB | Ikbkb | 3470 | 0.850 | 0.2811 | No | ||
| 13 | PIK3CG | Pik3cg | 3524 | 0.831 | 0.2941 | No | ||
| 14 | EIF4EBP1 | Eif4ebp1 | 3742 | 0.777 | 0.2954 | No | ||
| 15 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.2922 | No | ||
| 16 | ZBTB16 | Zbtb16 | 4070 | 0.708 | 0.3024 | No | ||
| 17 | STAT5A | Stat5a | 4244 | 0.660 | 0.3042 | No | ||
| 18 | SOS2 | Sos2 | 4544 | 0.582 | 0.2963 | No | ||
| 19 | ARAF | Araf | 4642 | 0.557 | 0.3010 | No | ||
| 20 | RUNX1 | Runx1 | 4989 | 0.482 | 0.2882 | No | ||
| 21 | RARA | Rara | 5279 | 0.432 | 0.2780 | No | ||
| 22 | PIM2 | Pim2 | 5368 | 0.425 | 0.2807 | No | ||
| 23 | BRAF | Braf | 5445 | 0.408 | 0.2838 | No | ||
| 24 | CEBPA | Cebpa | 5466 | 0.405 | 0.2904 | No | ||
| 25 | CHUK | Chuk | 5683 | 0.379 | 0.2839 | No | ||
| 26 | PPARD | Ppard | 5691 | 0.378 | 0.2909 | No | ||
| 27 | STAT5B | Stat5b | 6014 | 0.334 | 0.2767 | No | ||
| 28 | RELA | Rela | 6511 | 0.233 | 0.2492 | No | ||
| 29 | AKT2 | Akt2 | 6853 | 0.164 | 0.2304 | No | ||
| 30 | AKT1 | Akt1 | 7042 | 0.129 | 0.2208 | No | ||
| 31 | PIK3CA | Pik3ca | 7585 | 0.030 | 0.1864 | No | ||
| 32 | SOS1 | Sos1 | 8166 | -0.030 | 0.1495 | No | ||
| 33 | RPS6KB2 | Rps6kb2 | 8290 | -0.055 | 0.1426 | No | ||
| 34 | BAD | Bad | 8802 | -0.146 | 0.1125 | No | ||
| 35 | PIK3R5 | Pik3r5 | 8900 | -0.160 | 0.1094 | No | ||
| 36 | MTOR | Mtor | 9221 | -0.219 | 0.0930 | No | ||
| 37 | TCF7L2 | Tcf7l2 | 9232 | -0.220 | 0.0967 | No | ||
| 38 | HRAS | Hras | 9287 | -0.231 | 0.0977 | No | ||
| 39 | MAP2K1 | Map2k1 | 9298 | -0.232 | 0.1017 | No | ||
| 40 | CCNA1 | Ccna1 | 10123 | -0.394 | 0.0561 | No | ||
| 41 | MAPK1 | Mapk1 | 10768 | -0.489 | 0.0242 | No | ||
| 42 | KRAS | Kras | 11151 | -0.567 | 0.0106 | No | ||
| 43 | GRB2 | Grb2 | 11322 | -0.609 | 0.0116 | No | ||
| 44 | KIT | Kit | 11441 | -0.637 | 0.0165 | No | ||
| 45 | RAF1 | Raf1 | 11555 | -0.662 | 0.0223 | No | ||
| 46 | TCF7 | Tcf7 | 11581 | -0.668 | 0.0338 | No | ||
| 47 | AKT3 | Akt3 | 11977 | -0.777 | 0.0235 | No | ||
| 48 | MYC | Myc | 13033 | -1.135 | -0.0223 | No | ||
| 49 | PIK3CB | Pik3cb | 13719 | -1.479 | -0.0375 | No | ||
| 50 | PIK3R1 | Pik3r1 | 13721 | -1.480 | -0.0084 | No | ||
| 51 | PML | Pml | 13930 | -1.638 | 0.0104 | No | ||
| 52 | NFKB1 | Nfkb1 | 14275 | -1.949 | 0.0265 | No | ||
| 53 | CCND1 | Ccnd1 | 14834 | -2.752 | 0.0447 | No |