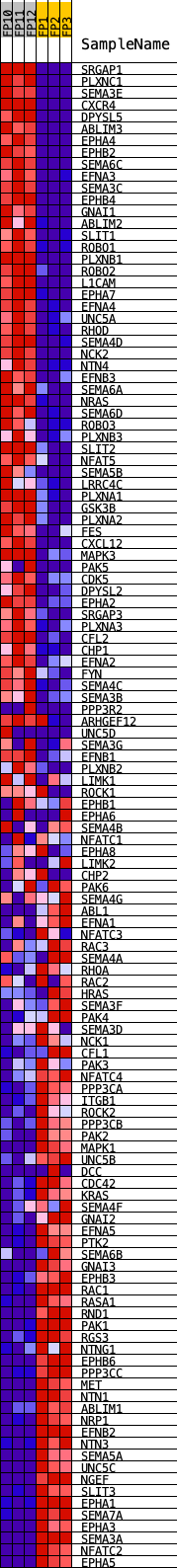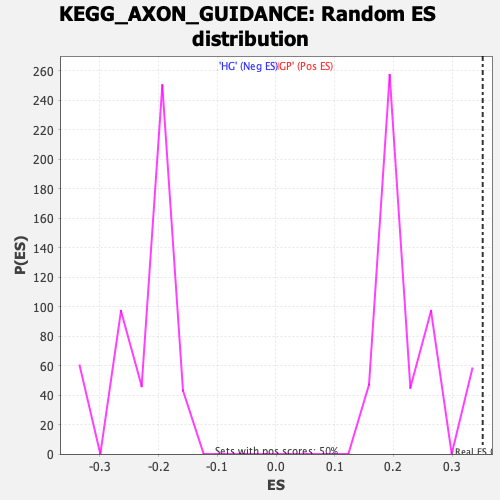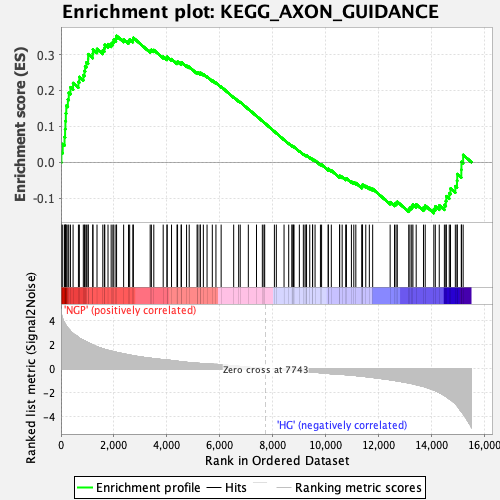
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_AXON_GUIDANCE |
| Enrichment Score (ES) | 0.35204583 |
| Normalized Enrichment Score (NES) | 1.6010805 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.29111183 |
| FWER p-Value | 0.376 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SRGAP1 | Srgap1 | 15 | 4.654 | 0.0281 | Yes | ||
| 2 | PLXNC1 | Plxnc1 | 65 | 4.169 | 0.0509 | Yes | ||
| 3 | SEMA3E | Sema3e | 134 | 3.830 | 0.0704 | Yes | ||
| 4 | CXCR4 | Cxcr4 | 155 | 3.749 | 0.0924 | Yes | ||
| 5 | DPYSL5 | Dpysl5 | 167 | 3.712 | 0.1149 | Yes | ||
| 6 | ABLIM3 | Ablim3 | 183 | 3.625 | 0.1365 | Yes | ||
| 7 | EPHA4 | Epha4 | 200 | 3.559 | 0.1577 | Yes | ||
| 8 | EPHB2 | Ephb2 | 260 | 3.388 | 0.1750 | Yes | ||
| 9 | SEMA6C | Sema6c | 294 | 3.306 | 0.1935 | Yes | ||
| 10 | EFNA3 | Efna3 | 357 | 3.116 | 0.2089 | Yes | ||
| 11 | SEMA3C | Sema3c | 457 | 2.928 | 0.2207 | Yes | ||
| 12 | EPHB4 | Ephb4 | 659 | 2.607 | 0.2239 | Yes | ||
| 13 | GNAI1 | Gnai1 | 694 | 2.552 | 0.2376 | Yes | ||
| 14 | ABLIM2 | Ablim2 | 844 | 2.370 | 0.2427 | Yes | ||
| 15 | SLIT1 | Slit1 | 887 | 2.326 | 0.2545 | Yes | ||
| 16 | ROBO1 | Robo1 | 912 | 2.301 | 0.2673 | Yes | ||
| 17 | PLXNB1 | Plxnb1 | 960 | 2.245 | 0.2783 | Yes | ||
| 18 | ROBO2 | Robo2 | 1025 | 2.169 | 0.2876 | Yes | ||
| 19 | L1CAM | L1cam | 1027 | 2.168 | 0.3011 | Yes | ||
| 20 | EPHA7 | Epha7 | 1201 | 1.986 | 0.3022 | Yes | ||
| 21 | EFNA4 | Efna4 | 1208 | 1.977 | 0.3142 | Yes | ||
| 22 | UNC5A | Unc5a | 1362 | 1.835 | 0.3157 | Yes | ||
| 23 | RHOD | Rhod | 1581 | 1.657 | 0.3119 | Yes | ||
| 24 | SEMA4D | Sema4d | 1641 | 1.615 | 0.3181 | Yes | ||
| 25 | NCK2 | Nck2 | 1656 | 1.606 | 0.3272 | Yes | ||
| 26 | NTN4 | Ntn4 | 1778 | 1.531 | 0.3289 | Yes | ||
| 27 | EFNB3 | Efnb3 | 1897 | 1.464 | 0.3304 | Yes | ||
| 28 | SEMA6A | Sema6a | 1962 | 1.432 | 0.3352 | Yes | ||
| 29 | NRAS | Nras | 1999 | 1.405 | 0.3416 | Yes | ||
| 30 | SEMA6D | Sema6d | 2078 | 1.370 | 0.3451 | Yes | ||
| 31 | ROBO3 | Robo3 | 2102 | 1.359 | 0.3520 | Yes | ||
| 32 | PLXNB3 | Plxnb3 | 2369 | 1.239 | 0.3425 | No | ||
| 33 | SLIT2 | Slit2 | 2554 | 1.152 | 0.3377 | No | ||
| 34 | NFAT5 | Nfat5 | 2597 | 1.137 | 0.3421 | No | ||
| 35 | SEMA5B | Sema5b | 2718 | 1.091 | 0.3411 | No | ||
| 36 | LRRC4C | Lrrc4c | 2737 | 1.083 | 0.3467 | No | ||
| 37 | PLXNA1 | Plxna1 | 3373 | 0.868 | 0.3109 | No | ||
| 38 | GSK3B | Gsk3b | 3418 | 0.860 | 0.3134 | No | ||
| 39 | PLXNA2 | Plxna2 | 3510 | 0.836 | 0.3127 | No | ||
| 40 | FES | Fes | 3865 | 0.745 | 0.2943 | No | ||
| 41 | CXCL12 | Cxcl12 | 4011 | 0.720 | 0.2894 | No | ||
| 42 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.2939 | No | ||
| 43 | PAK5 | Pak7 | 4180 | 0.678 | 0.2873 | No | ||
| 44 | CDK5 | Cdk5 | 4388 | 0.624 | 0.2777 | No | ||
| 45 | DPYSL2 | Dpysl2 | 4408 | 0.618 | 0.2804 | No | ||
| 46 | EPHA2 | Epha2 | 4545 | 0.582 | 0.2752 | No | ||
| 47 | SRGAP3 | Srgap3 | 4553 | 0.581 | 0.2783 | No | ||
| 48 | PLXNA3 | Plxna3 | 4749 | 0.530 | 0.2690 | No | ||
| 49 | CFL2 | Cfl2 | 4849 | 0.507 | 0.2657 | No | ||
| 50 | CHP1 | Chp1 | 5147 | 0.454 | 0.2492 | No | ||
| 51 | EFNA2 | Efna2 | 5178 | 0.447 | 0.2501 | No | ||
| 52 | FYN | Fyn | 5264 | 0.436 | 0.2473 | No | ||
| 53 | SEMA4C | Sema4c | 5268 | 0.435 | 0.2498 | No | ||
| 54 | SEMA3B | Sema3b | 5384 | 0.421 | 0.2450 | No | ||
| 55 | PPP3R2 | Ppp3r2 | 5524 | 0.401 | 0.2384 | No | ||
| 56 | ARHGEF12 | Arhgef12 | 5723 | 0.368 | 0.2279 | No | ||
| 57 | UNC5D | Unc5d | 5860 | 0.356 | 0.2213 | No | ||
| 58 | SEMA3G | Sema3g | 6057 | 0.326 | 0.2106 | No | ||
| 59 | EFNB1 | Efnb1 | 6531 | 0.229 | 0.1813 | No | ||
| 60 | PLXNB2 | Plxnb2 | 6720 | 0.195 | 0.1703 | No | ||
| 61 | LIMK1 | Limk1 | 6779 | 0.181 | 0.1676 | No | ||
| 62 | ROCK1 | Rock1 | 7085 | 0.119 | 0.1486 | No | ||
| 63 | EPHB1 | Ephb1 | 7394 | 0.062 | 0.1290 | No | ||
| 64 | EPHA6 | Epha6 | 7615 | 0.025 | 0.1148 | No | ||
| 65 | SEMA4B | Sema4b | 7676 | 0.012 | 0.1110 | No | ||
| 66 | NFATC1 | Nfatc1 | 7704 | 0.008 | 0.1093 | No | ||
| 67 | EPHA8 | Epha8 | 8071 | -0.011 | 0.0856 | No | ||
| 68 | LIMK2 | Limk2 | 8144 | -0.026 | 0.0811 | No | ||
| 69 | CHP2 | Chp2 | 8436 | -0.082 | 0.0627 | No | ||
| 70 | PAK6 | Pak6 | 8612 | -0.113 | 0.0521 | No | ||
| 71 | SEMA4G | Sema4g | 8738 | -0.135 | 0.0448 | No | ||
| 72 | ABL1 | Abl1 | 8739 | -0.135 | 0.0456 | No | ||
| 73 | EFNA1 | Efna1 | 8793 | -0.142 | 0.0431 | No | ||
| 74 | NFATC3 | Nfatc3 | 8813 | -0.148 | 0.0428 | No | ||
| 75 | RAC3 | Rac3 | 9015 | -0.180 | 0.0308 | No | ||
| 76 | SEMA4A | Sema4a | 9166 | -0.208 | 0.0224 | No | ||
| 77 | RHOA | Rhoa | 9229 | -0.220 | 0.0197 | No | ||
| 78 | RAC2 | Rac2 | 9271 | -0.227 | 0.0185 | No | ||
| 79 | HRAS | Hras | 9287 | -0.231 | 0.0189 | No | ||
| 80 | SEMA3F | Sema3f | 9408 | -0.254 | 0.0127 | No | ||
| 81 | PAK4 | Pak4 | 9515 | -0.272 | 0.0076 | No | ||
| 82 | SEMA3D | Sema3d | 9516 | -0.272 | 0.0092 | No | ||
| 83 | NCK1 | Nck1 | 9609 | -0.291 | 0.0051 | No | ||
| 84 | CFL1 | Cfl1 | 9812 | -0.332 | -0.0060 | No | ||
| 85 | PAK3 | Pak3 | 9856 | -0.340 | -0.0066 | No | ||
| 86 | NFATC4 | Nfatc4 | 10107 | -0.391 | -0.0204 | No | ||
| 87 | PPP3CA | Ppp3ca | 10109 | -0.391 | -0.0181 | No | ||
| 88 | ITGB1 | Itgb1 | 10224 | -0.414 | -0.0229 | No | ||
| 89 | ROCK2 | Rock2 | 10537 | -0.442 | -0.0404 | No | ||
| 90 | PPP3CB | Ppp3cb | 10538 | -0.443 | -0.0376 | No | ||
| 91 | PAK2 | Pak2 | 10634 | -0.463 | -0.0409 | No | ||
| 92 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.0465 | No | ||
| 93 | UNC5B | Unc5b | 10796 | -0.495 | -0.0451 | No | ||
| 94 | DCC | Dcc | 10984 | -0.530 | -0.0540 | No | ||
| 95 | CDC42 | Cdc42 | 11074 | -0.550 | -0.0563 | No | ||
| 96 | KRAS | Kras | 11151 | -0.567 | -0.0577 | No | ||
| 97 | SEMA4F | Sema4f | 11379 | -0.624 | -0.0686 | No | ||
| 98 | GNAI2 | Gnai2 | 11388 | -0.627 | -0.0652 | No | ||
| 99 | EFNA5 | Efna5 | 11405 | -0.629 | -0.0623 | No | ||
| 100 | PTK2 | Ptk2 | 11527 | -0.655 | -0.0661 | No | ||
| 101 | SEMA6B | Sema6b | 11662 | -0.688 | -0.0705 | No | ||
| 102 | GNAI3 | Gnai3 | 11788 | -0.724 | -0.0741 | No | ||
| 103 | EPHB3 | Ephb3 | 12449 | -0.918 | -0.1112 | No | ||
| 104 | RAC1 | Rac1 | 12614 | -0.976 | -0.1158 | No | ||
| 105 | RASA1 | Rasa1 | 12671 | -0.994 | -0.1132 | No | ||
| 106 | RND1 | Rnd1 | 12719 | -1.014 | -0.1100 | No | ||
| 107 | PAK1 | Pak1 | 13152 | -1.182 | -0.1306 | No | ||
| 108 | RGS3 | Rgs3 | 13198 | -1.206 | -0.1260 | No | ||
| 109 | NTNG1 | Ntng1 | 13266 | -1.236 | -0.1227 | No | ||
| 110 | EPHB6 | Ephb6 | 13310 | -1.254 | -0.1176 | No | ||
| 111 | PPP3CC | Ppp3cc | 13434 | -1.328 | -0.1173 | No | ||
| 112 | MET | Met | 13710 | -1.474 | -0.1260 | No | ||
| 113 | NTN1 | Ntn1 | 13775 | -1.524 | -0.1207 | No | ||
| 114 | ABLIM1 | Ablim1 | 14101 | -1.773 | -0.1307 | No | ||
| 115 | NRP1 | Nrp1 | 14157 | -1.822 | -0.1229 | No | ||
| 116 | EFNB2 | Efnb2 | 14306 | -1.985 | -0.1202 | No | ||
| 117 | NTN3 | Ntn3 | 14500 | -2.220 | -0.1188 | No | ||
| 118 | SEMA5A | Sema5a | 14555 | -2.304 | -0.1080 | No | ||
| 119 | UNC5C | Unc5c | 14574 | -2.329 | -0.0946 | No | ||
| 120 | NGEF | Ngef | 14685 | -2.507 | -0.0861 | No | ||
| 121 | SLIT3 | Slit3 | 14732 | -2.577 | -0.0730 | No | ||
| 122 | EPHA1 | Epha1 | 14914 | -2.904 | -0.0667 | No | ||
| 123 | SEMA7A | Sema7a | 14978 | -3.070 | -0.0516 | No | ||
| 124 | EPHA3 | Epha3 | 14987 | -3.090 | -0.0329 | No | ||
| 125 | SEMA3A | Sema3a | 15141 | -3.539 | -0.0207 | No | ||
| 126 | NFATC2 | Nfatc2 | 15146 | -3.562 | 0.0012 | No | ||
| 127 | EPHA5 | Epha5 | 15210 | -3.737 | 0.0205 | No |