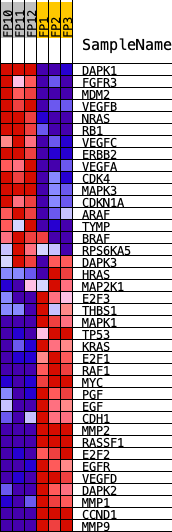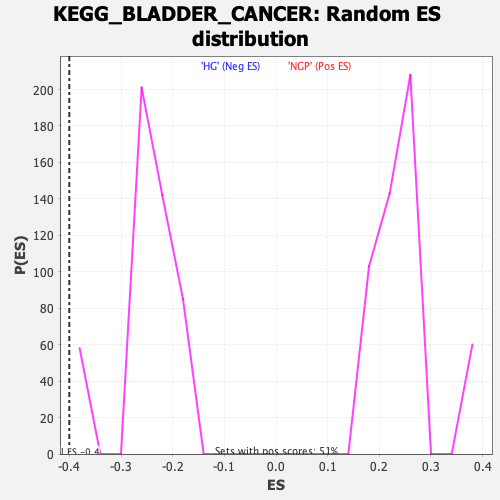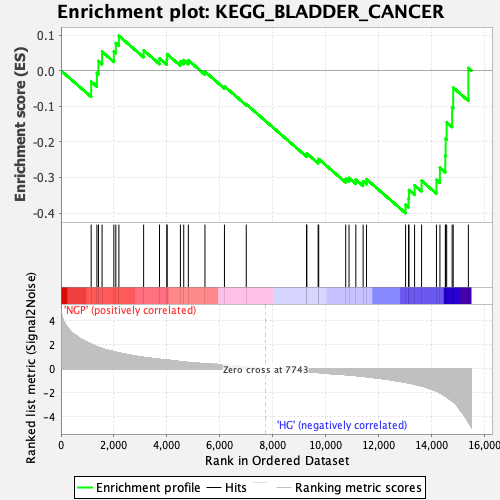
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_BLADDER_CANCER |
| Enrichment Score (ES) | -0.4003769 |
| Normalized Enrichment Score (NES) | -1.5728766 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.12991883 |
| FWER p-Value | 0.413 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DAPK1 | Dapk1 | 1141 | 2.054 | -0.0308 | No | ||
| 2 | FGFR3 | Fgfr3 | 1358 | 1.838 | -0.0064 | No | ||
| 3 | MDM2 | Mdm2 | 1417 | 1.780 | 0.0269 | No | ||
| 4 | VEGFB | Vegfb | 1553 | 1.680 | 0.0532 | No | ||
| 5 | NRAS | Nras | 1999 | 1.405 | 0.0538 | No | ||
| 6 | RB1 | Rb1 | 2074 | 1.372 | 0.0777 | No | ||
| 7 | VEGFC | Vegfc | 2190 | 1.316 | 0.0977 | No | ||
| 8 | ERBB2 | Erbb2 | 3125 | 0.943 | 0.0570 | No | ||
| 9 | VEGFA | Vegfa | 3726 | 0.782 | 0.0346 | No | ||
| 10 | CDK4 | Cdk4 | 4004 | 0.722 | 0.0318 | No | ||
| 11 | MAPK3 | Mapk3 | 4012 | 0.720 | 0.0463 | No | ||
| 12 | CDKN1A | Cdkn1a | 4516 | 0.587 | 0.0261 | No | ||
| 13 | ARAF | Araf | 4642 | 0.557 | 0.0296 | No | ||
| 14 | TYMP | Tymp | 4816 | 0.516 | 0.0292 | No | ||
| 15 | BRAF | Braf | 5445 | 0.408 | -0.0028 | No | ||
| 16 | RPS6KA5 | Rps6ka5 | 6182 | 0.301 | -0.0440 | No | ||
| 17 | DAPK3 | Dapk3 | 7007 | 0.135 | -0.0944 | No | ||
| 18 | HRAS | Hras | 9287 | -0.231 | -0.2368 | No | ||
| 19 | MAP2K1 | Map2k1 | 9298 | -0.232 | -0.2326 | No | ||
| 20 | E2F3 | E2f3 | 9725 | -0.313 | -0.2536 | No | ||
| 21 | THBS1 | Thbs1 | 9747 | -0.318 | -0.2483 | No | ||
| 22 | MAPK1 | Mapk1 | 10768 | -0.489 | -0.3040 | No | ||
| 23 | TP53 | Trp53 | 10894 | -0.512 | -0.3014 | No | ||
| 24 | KRAS | Kras | 11151 | -0.567 | -0.3061 | No | ||
| 25 | E2F1 | E2f1 | 11427 | -0.633 | -0.3106 | No | ||
| 26 | RAF1 | Raf1 | 11555 | -0.662 | -0.3050 | No | ||
| 27 | MYC | Myc | 13033 | -1.135 | -0.3767 | Yes | ||
| 28 | PGF | Pgf | 13145 | -1.179 | -0.3593 | Yes | ||
| 29 | EGF | Egf | 13161 | -1.188 | -0.3355 | Yes | ||
| 30 | CDH1 | Cdh1 | 13373 | -1.294 | -0.3221 | Yes | ||
| 31 | MMP2 | Mmp2 | 13638 | -1.437 | -0.3092 | Yes | ||
| 32 | RASSF1 | Rassf1 | 14200 | -1.866 | -0.3065 | Yes | ||
| 33 | E2F2 | E2f2 | 14332 | -2.016 | -0.2729 | Yes | ||
| 34 | EGFR | Egfr | 14536 | -2.270 | -0.2387 | Yes | ||
| 35 | VEGFD | Vegfd | 14548 | -2.290 | -0.1916 | Yes | ||
| 36 | DAPK2 | Dapk2 | 14587 | -2.342 | -0.1452 | Yes | ||
| 37 | MMP1 | Mmp1a | 14793 | -2.677 | -0.1026 | Yes | ||
| 38 | CCND1 | Ccnd1 | 14834 | -2.752 | -0.0478 | Yes | ||
| 39 | MMP9 | Mmp9 | 15408 | -4.429 | 0.0076 | Yes |