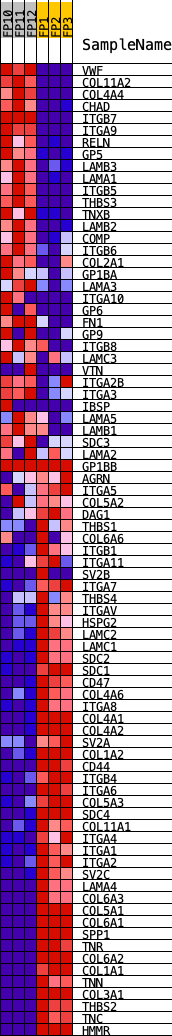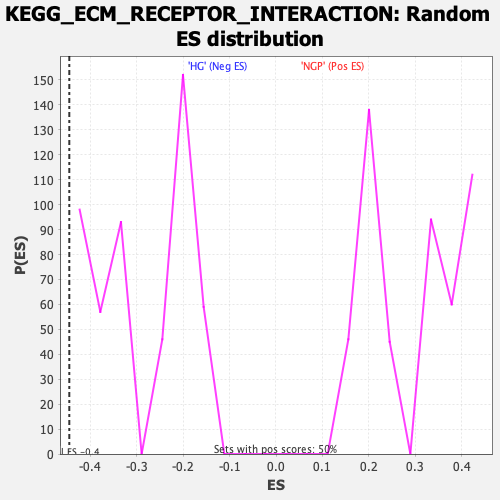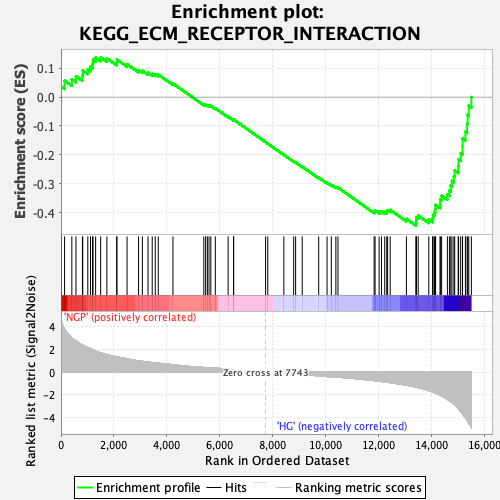
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | HG |
| GeneSet | KEGG_ECM_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | -0.44521514 |
| Normalized Enrichment Score (NES) | -1.5086037 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15305476 |
| FWER p-Value | 0.668 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VWF | Vwf | 3 | 4.885 | 0.0365 | No | ||
| 2 | COL11A2 | Col11a2 | 135 | 3.829 | 0.0568 | No | ||
| 3 | COL4A4 | Col4a4 | 410 | 3.008 | 0.0616 | No | ||
| 4 | CHAD | Chad | 566 | 2.774 | 0.0724 | No | ||
| 5 | ITGB7 | Itgb7 | 810 | 2.407 | 0.0748 | No | ||
| 6 | ITGA9 | Itga9 | 822 | 2.396 | 0.0921 | No | ||
| 7 | RELN | Reln | 1014 | 2.179 | 0.0961 | No | ||
| 8 | GP5 | Gp5 | 1116 | 2.082 | 0.1052 | No | ||
| 9 | LAMB3 | Lamb3 | 1193 | 1.992 | 0.1152 | No | ||
| 10 | LAMA1 | Lama1 | 1210 | 1.976 | 0.1290 | No | ||
| 11 | ITGB5 | Itgb5 | 1310 | 1.874 | 0.1367 | No | ||
| 12 | THBS3 | Thbs3 | 1500 | 1.724 | 0.1374 | No | ||
| 13 | TNXB | Tnxb | 1735 | 1.558 | 0.1340 | No | ||
| 14 | LAMB2 | Lamb2 | 2107 | 1.356 | 0.1202 | No | ||
| 15 | COMP | Comp | 2116 | 1.353 | 0.1298 | No | ||
| 16 | ITGB6 | Itgb6 | 2499 | 1.178 | 0.1139 | No | ||
| 17 | COL2A1 | Col2a1 | 2933 | 1.004 | 0.0934 | No | ||
| 18 | GP1BA | Gp1ba | 3077 | 0.959 | 0.0914 | No | ||
| 19 | LAMA3 | Lama3 | 3292 | 0.895 | 0.0842 | No | ||
| 20 | ITGA10 | Gm42957 | 3450 | 0.853 | 0.0805 | No | ||
| 21 | GP6 | Gp6 | 3565 | 0.820 | 0.0793 | No | ||
| 22 | FN1 | Fn1 | 3678 | 0.794 | 0.0780 | No | ||
| 23 | GP9 | Gp9 | 4234 | 0.662 | 0.0470 | No | ||
| 24 | ITGB8 | Itgb8 | 5402 | 0.417 | -0.0254 | No | ||
| 25 | LAMC3 | Lamc3 | 5471 | 0.404 | -0.0268 | No | ||
| 26 | VTN | Vtn | 5537 | 0.401 | -0.0280 | No | ||
| 27 | ITGA2B | Itga2b | 5599 | 0.396 | -0.0290 | No | ||
| 28 | ITGA3 | Itga3 | 5667 | 0.382 | -0.0304 | No | ||
| 29 | IBSP | Ibsp | 5837 | 0.356 | -0.0387 | No | ||
| 30 | LAMA5 | Lama5 | 6321 | 0.271 | -0.0679 | No | ||
| 31 | LAMB1 | Lamb1 | 6524 | 0.230 | -0.0793 | No | ||
| 32 | SDC3 | Sdc3 | 6532 | 0.229 | -0.0780 | No | ||
| 33 | LAMA2 | Lama2 | 7738 | 0.001 | -0.1560 | No | ||
| 34 | GP1BB | Gp1bb | 7821 | 0.000 | -0.1613 | No | ||
| 35 | AGRN | Agrn | 8429 | -0.082 | -0.2000 | No | ||
| 36 | ITGA5 | Itga5 | 8800 | -0.145 | -0.2229 | No | ||
| 37 | COL5A2 | Col5a2 | 8874 | -0.155 | -0.2264 | No | ||
| 38 | DAG1 | Dag1 | 9124 | -0.200 | -0.2411 | No | ||
| 39 | THBS1 | Thbs1 | 9747 | -0.318 | -0.2789 | No | ||
| 40 | COL6A6 | Col6a6 | 10063 | -0.382 | -0.2965 | No | ||
| 41 | ITGB1 | Itgb1 | 10224 | -0.414 | -0.3037 | No | ||
| 42 | ITGA11 | Itga11 | 10396 | -0.426 | -0.3116 | No | ||
| 43 | SV2B | Sv2b | 10476 | -0.440 | -0.3134 | No | ||
| 44 | ITGA7 | Itga7 | 11843 | -0.740 | -0.3963 | No | ||
| 45 | THBS4 | Thbs4 | 11879 | -0.752 | -0.3929 | No | ||
| 46 | ITGAV | Itgav | 12032 | -0.796 | -0.3968 | No | ||
| 47 | HSPG2 | Hspg2 | 12121 | -0.817 | -0.3963 | No | ||
| 48 | LAMC2 | Lamc2 | 12244 | -0.851 | -0.3978 | No | ||
| 49 | LAMC1 | Lamc1 | 12320 | -0.874 | -0.3961 | No | ||
| 50 | SDC2 | Sdc2 | 12354 | -0.886 | -0.3916 | No | ||
| 51 | SDC1 | Sdc1 | 12451 | -0.919 | -0.3909 | No | ||
| 52 | CD47 | Cd47 | 13064 | -1.147 | -0.4219 | No | ||
| 53 | COL4A6 | Col4a6 | 13425 | -1.324 | -0.4353 | Yes | ||
| 54 | ITGA8 | Itga8 | 13437 | -1.329 | -0.4260 | Yes | ||
| 55 | COL4A1 | Col4a1 | 13442 | -1.331 | -0.4163 | Yes | ||
| 56 | COL4A2 | Col4a2 | 13510 | -1.361 | -0.4104 | Yes | ||
| 57 | SV2A | Sv2a | 13910 | -1.620 | -0.4240 | Yes | ||
| 58 | COL1A2 | Col1a2 | 14054 | -1.731 | -0.4203 | Yes | ||
| 59 | CD44 | Cd44 | 14072 | -1.745 | -0.4083 | Yes | ||
| 60 | ITGB4 | Itgb4 | 14125 | -1.795 | -0.3982 | Yes | ||
| 61 | ITGA6 | Itga6 | 14158 | -1.823 | -0.3865 | Yes | ||
| 62 | COL5A3 | Col5a3 | 14168 | -1.835 | -0.3733 | Yes | ||
| 63 | SDC4 | Sdc4 | 14341 | -2.025 | -0.3693 | Yes | ||
| 64 | COL11A1 | Col11a1 | 14344 | -2.028 | -0.3542 | Yes | ||
| 65 | ITGA4 | Itga4 | 14396 | -2.091 | -0.3417 | Yes | ||
| 66 | ITGA1 | Itga1 | 14614 | -2.383 | -0.3379 | Yes | ||
| 67 | ITGA2 | Itga2 | 14691 | -2.524 | -0.3238 | Yes | ||
| 68 | SV2C | Sv2c | 14743 | -2.591 | -0.3077 | Yes | ||
| 69 | LAMA4 | Lama4 | 14786 | -2.655 | -0.2904 | Yes | ||
| 70 | COL6A3 | Col6a3 | 14860 | -2.803 | -0.2741 | Yes | ||
| 71 | COL5A1 | Col5a1 | 14890 | -2.862 | -0.2545 | Yes | ||
| 72 | COL6A1 | Col6a1 | 15022 | -3.185 | -0.2390 | Yes | ||
| 73 | SPP1 | Spp1 | 15038 | -3.234 | -0.2157 | Yes | ||
| 74 | TNR | Tnr | 15117 | -3.492 | -0.1945 | Yes | ||
| 75 | COL6A2 | Col6a2 | 15186 | -3.673 | -0.1713 | Yes | ||
| 76 | COL1A1 | Col1a1 | 15187 | -3.673 | -0.1437 | Yes | ||
| 77 | TNN | Tnn | 15295 | -4.041 | -0.1203 | Yes | ||
| 78 | COL3A1 | Col3a1 | 15359 | -4.249 | -0.0925 | Yes | ||
| 79 | THBS2 | Thbs2 | 15382 | -4.337 | -0.0613 | Yes | ||
| 80 | TNC | Tnc | 15423 | -4.499 | -0.0301 | Yes | ||
| 81 | HMMR | Hmmr | 15521 | -4.876 | 0.0003 | Yes |