Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | preliminary.phenotype.cls#NGP_versus_HG.phenotype.cls#NGP_versus_HG_repos |
| Phenotype | phenotype.cls#NGP_versus_HG_repos |
| Upregulated in class | NGP |
| GeneSet | KEGG_EPITHELIAL_CELL_SIGNALING_IN_HELICOBACTER_PYLORI_INFECTION |
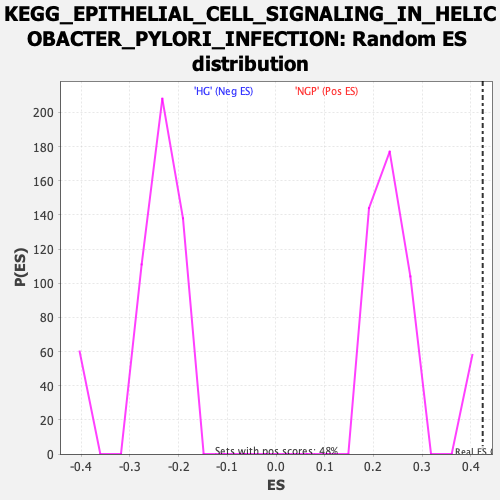
| Enrichment Score (ES) | 0.42459857 |
| Normalized Enrichment Score (NES) | 1.6684622 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.39487472 |
| FWER p-Value | 0.156 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP6V0B | Atp6v0b | 516 | 2.828 | 0.0007 | Yes | ||
| 2 | ATP6V1A | Atp6v1a | 679 | 2.574 | 0.0213 | Yes | ||
| 3 | IKBKG | Ikbkg | 706 | 2.530 | 0.0501 | Yes | ||
| 4 | PLCG2 | Plcg2 | 735 | 2.492 | 0.0783 | Yes | ||
| 5 | F11R | F11r | 750 | 2.476 | 0.1073 | Yes | ||
| 6 | ATP6V0D1 | Atp6v0d1 | 781 | 2.439 | 0.1347 | Yes | ||
| 7 | GIT1 | Git1 | 917 | 2.296 | 0.1537 | Yes | ||
| 8 | ATP6V1D | Atp6v1d | 948 | 2.260 | 0.1790 | Yes | ||
| 9 | ATP6V1B2 | Atp6v1b2 | 1065 | 2.129 | 0.1971 | Yes | ||
| 10 | ATP6V1C1 | Atp6v1c1 | 1132 | 2.064 | 0.2177 | Yes | ||
| 11 | ATP6V1F | Atp6v1f | 1147 | 2.051 | 0.2416 | Yes | ||
| 12 | JAM3 | Jam3 | 1183 | 2.008 | 0.2635 | Yes | ||
| 13 | LYN | Lyn | 1345 | 1.850 | 0.2754 | Yes | ||
| 14 | ATP6AP1 | Atp6ap1 | 1377 | 1.821 | 0.2954 | Yes | ||
| 15 | TCIRG1 | Tcirg1 | 1479 | 1.737 | 0.3098 | Yes | ||
| 16 | ATP6V1G1 | Atp6v1g1 | 1533 | 1.696 | 0.3268 | Yes | ||
| 17 | CXCR2 | Cxcr2 | 1558 | 1.675 | 0.3454 | Yes | ||
| 18 | MAPK12 | Mapk12 | 1560 | 1.674 | 0.3655 | Yes | ||
| 19 | PLCG1 | Plcg1 | 1648 | 1.613 | 0.3794 | Yes | ||
| 20 | ATP6V0C | Atp6v0c | 1653 | 1.608 | 0.3985 | Yes | ||
| 21 | ATP6V1H | Atp6v1h | 1675 | 1.596 | 0.4164 | Yes | ||
| 22 | ATP6V1E1 | Atp6v1e1 | 1829 | 1.504 | 0.4246 | Yes | ||
| 23 | ATP6V0E1 | Atp6v0e | 2994 | 0.981 | 0.3612 | No | ||
| 24 | ATP6V0E2 | Atp6v0e2 | 3180 | 0.924 | 0.3603 | No | ||
| 25 | IKBKB | Ikbkb | 3470 | 0.850 | 0.3519 | No | ||
| 26 | MAPK11 | Mapk11 | 4400 | 0.621 | 0.2993 | No | ||
| 27 | MAPK8 | Mapk8 | 4803 | 0.518 | 0.2796 | No | ||
| 28 | MAPK10 | Mapk10 | 5168 | 0.449 | 0.2614 | No | ||
| 29 | CHUK | Chuk | 5683 | 0.379 | 0.2328 | No | ||
| 30 | ATP6V0A2 | Atp6v0a2 | 6233 | 0.291 | 0.2008 | No | ||
| 31 | RELA | Rela | 6511 | 0.233 | 0.1857 | No | ||
| 32 | CSK | Csk | 6540 | 0.226 | 0.1866 | No | ||
| 33 | ADAM10 | Adam10 | 6851 | 0.165 | 0.1685 | No | ||
| 34 | NFKBIA | Nfkbia | 6940 | 0.148 | 0.1646 | No | ||
| 35 | ATP6V1G2 | Atp6v1g2 | 7538 | 0.039 | 0.1265 | No | ||
| 36 | MAP2K4 | Map2k4 | 8722 | -0.131 | 0.0516 | No | ||
| 37 | SRC | Src | 8880 | -0.156 | 0.0433 | No | ||
| 38 | ATP6V0A1 | Atp6v0a1 | 8992 | -0.175 | 0.0382 | No | ||
| 39 | MAPK9 | Mapk9 | 10579 | -0.451 | -0.0589 | No | ||
| 40 | ADAM17 | Adam17 | 10614 | -0.459 | -0.0555 | No | ||
| 41 | NOD1 | Nod1 | 11034 | -0.540 | -0.0761 | No | ||
| 42 | CDC42 | Cdc42 | 11074 | -0.550 | -0.0720 | No | ||
| 43 | MAPK14 | Mapk14 | 11756 | -0.714 | -0.1074 | No | ||
| 44 | TJP1 | Tjp1 | 11958 | -0.773 | -0.1111 | No | ||
| 45 | ATP6V1C2 | Atp6v1c2 | 12423 | -0.908 | -0.1302 | No | ||
| 46 | RAC1 | Rac1 | 12614 | -0.976 | -0.1307 | No | ||
| 47 | PTPN11 | Ptpn11 | 12664 | -0.992 | -0.1219 | No | ||
| 48 | MAPK13 | Mapk13 | 12835 | -1.057 | -0.1202 | No | ||
| 49 | ATP6V1B1 | Atp6v1b1 | 12852 | -1.064 | -0.1084 | No | ||
| 50 | CASP3 | Casp3 | 12911 | -1.085 | -0.0990 | No | ||
| 51 | PAK1 | Pak1 | 13152 | -1.182 | -0.1003 | No | ||
| 52 | ATP6V0A4 | Atp6v0a4 | 13323 | -1.264 | -0.0961 | No | ||
| 53 | HBEGF | Hbegf | 13569 | -1.400 | -0.0950 | No | ||
| 54 | MET | Met | 13710 | -1.474 | -0.0863 | No | ||
| 55 | JAM2 | Jam2 | 14045 | -1.723 | -0.0871 | No | ||
| 56 | NFKB1 | Nfkb1 | 14275 | -1.949 | -0.0785 | No | ||
| 57 | EGFR | Egfr | 14536 | -2.270 | -0.0679 | No | ||
| 58 | MAP3K14 | Map3k14 | 14636 | -2.430 | -0.0450 | No | ||
| 59 | JUN | Jun | 14704 | -2.538 | -0.0187 | No | ||
| 60 | CCL5 | Ccl5 | 14720 | -2.558 | 0.0111 | No | ||
| 61 | ATP6V0D2 | Atp6v0d2 | 15082 | -3.390 | 0.0286 | No |